Difference between revisions of "Part:BBa K1949060"
| (14 intermediate revisions by the same user not shown) | |||
| Line 2: | Line 2: | ||
<partinfo>BBa_K1949060 short</partinfo> | <partinfo>BBa_K1949060 short</partinfo> | ||
| − | <span style="margin-left: 10px;">We simulated our final genetic circuits and found that the circuits did not work, because Prhl strength was too weak. (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). We therefore considered using the improved Prhl (<partinfo>BBa_K1529300</partinfo>,<partinfo>BBa_K1529310</partinfo>) established by | + | <span style="margin-left: 10px;">We simulated our final genetic circuits and found that the circuits did not work, because Prhl (<partinfo>BBa_I14017</partinfo>) strength was too weak. (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). We therefore considered using the improved Prhl (<partinfo>BBa_K1529300</partinfo>,<partinfo>BBa_K1529310</partinfo>) established by iGEM 2014 Tokyo_Tech team, but we noticed that they were inappropriate for two reasons (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). Then, we decided to improve Prhl and obtain our original improved Prhl (included in <partinfo>BBa_K1949060</partinfo>) that suited our goal. |
| − | ===Characterize and | + | ===Characterize and Improve=== |
<span style="margin-left: 10px;">Our purpose is to create strong Prhl for our final genetic circuits. | <span style="margin-left: 10px;">Our purpose is to create strong Prhl for our final genetic circuits. | ||
| Line 10: | Line 10: | ||
This experiment consists of the three parts below. | This experiment consists of the three parts below. | ||
| − | I. Improved Prhl by iGEM 2014 Tokyo_Tech and | + | I. Improved Prhl by iGEM 2014 Tokyo_Tech team and characterization of <i>rhlR</i> |
II. Improvement of the wild type Prhl | II. Improvement of the wild type Prhl | ||
| − | III. Comparison of the improved Prhl by iGEM 2014 Tokyo_Tech to our original improved Prhl | + | III. Comparison of the improved Prhl by iGEM 2014 Tokyo_Tech team to our original improved Prhl |
The past improved Prhl did not suit for our final circuits and we could construct the improved Prhl appropriate to our final circuits. | The past improved Prhl did not suit for our final circuits and we could construct the improved Prhl appropriate to our final circuits. | ||
| − | ====I. Improved Prhl by iGEM 2014 Tokyo_Tech and | + | ====I. Improved Prhl by iGEM 2014 Tokyo_Tech team and characterization of <i>RhlR</i>==== |
<span style="margin-left: 10px;">We found that Prhl(RL) (<partinfo>BBa_K1529300</partinfo>) activity was weak and the expression level depended on LVA tag (Fig.1); LVA-tagged proteins are prone to be degraded by cellular proteases. Prhl(LR) (<partinfo>BBa_K1529310</partinfo>) activity was strong and unexpectedly reacted with C12 (crosstalk) (Fig.3). | <span style="margin-left: 10px;">We found that Prhl(RL) (<partinfo>BBa_K1529300</partinfo>) activity was weak and the expression level depended on LVA tag (Fig.1); LVA-tagged proteins are prone to be degraded by cellular proteases. Prhl(LR) (<partinfo>BBa_K1529310</partinfo>) activity was strong and unexpectedly reacted with C12 (crosstalk) (Fig.3). | ||
| Line 27: | Line 27: | ||
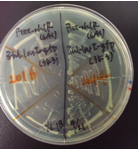
<span style="margin-left: 10px;">The colonies of transformants with a rhlR (<partinfo>BBa_C0171</partinfo>) plasmid looked rough and the growth rate was low(Fig.2-left), while the colonies of transformants with a rhlR-LVA (<partinfo>BBa_C0071</partinfo>) plasmid looked smooth and the growth rate was normal(Fig.2-right). However, the reason for this result is unclear. | <span style="margin-left: 10px;">The colonies of transformants with a rhlR (<partinfo>BBa_C0171</partinfo>) plasmid looked rough and the growth rate was low(Fig.2-left), while the colonies of transformants with a rhlR-LVA (<partinfo>BBa_C0071</partinfo>) plasmid looked smooth and the growth rate was normal(Fig.2-right). However, the reason for this result is unclear. | ||
| − | [[Image:prhl2.png|thumb|center|400px|Fig.2 The colonies of transformants with a rhlR (left) or a rhlR-LVA right]]<br> | + | [[Image:prhl2.png|thumb|center|400px|Fig.2 The colonies of transformants with a rhlR (left) or a rhlR-LVA (right)]]<br> |
| − | + | ||
====II. Improvement the wild type Prhl==== | ====II. Improvement the wild type Prhl==== | ||
| Line 36: | Line 35: | ||
[[Image:prhl4.png|thumb|center|400px|Fig. 3 RFU of GFP / turbidity of imoroved Prhl mutants]]<br> | [[Image:prhl4.png|thumb|center|400px|Fig. 3 RFU of GFP / turbidity of imoroved Prhl mutants]]<br> | ||
| − | The sequences of these | + | The sequences of these two chosen mutants are shown below. The small characters indicate the scar sequence and a single point mutation is colored with red. |
| − | ====III. Comparison the improved Prhl by iGEM 2014 Tokyo_Tech to our original improved Prhl==== | + | Native Prhl (<partinfo>BBa_R0071</partinfo>) |
| + | |||
| + | TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTAAAAAGTGTTC | ||
| + | |||
| + | Prhl(NM) (<partinfo>BBa_K1949060</partinfo>) | ||
| + | |||
| + | TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTA<font color=#ff0000><b>t</b></font>AAAGTGTTC | ||
| + | |||
| + | |||
| + | ====III. Comparison of the improved Prhl by iGEM 2014 Tokyo_Tech team to our original improved Prhl==== | ||
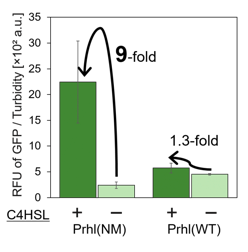
<span style="margin-left: 10px;">Prhl(NM) was chosen from the many Prhl mutants, and by comparing Prhl(NM) to Prhl(LR), we obtained the result below (Fig.4). The reaction activity of Prhl(NM) to C4 was stronger than that of Prhl(LR), and Prhl(NM) did not react with C12 at all. | <span style="margin-left: 10px;">Prhl(NM) was chosen from the many Prhl mutants, and by comparing Prhl(NM) to Prhl(LR), we obtained the result below (Fig.4). The reaction activity of Prhl(NM) to C4 was stronger than that of Prhl(LR), and Prhl(NM) did not react with C12 at all. | ||
| − | [[Image: | + | [[Image:prhl5.png|thumb|center|400px|Fig.4 Comparison of Prhl(NM) to Prhl(LR)]]<br> |
Latest revision as of 06:25, 25 October 2016
Prhl(NM)-rbs-gfp
We simulated our final genetic circuits and found that the circuits did not work, because Prhl (BBa_I14017) strength was too weak. (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). We therefore considered using the improved Prhl (BBa_K1529300,BBa_K1529310) established by iGEM 2014 Tokyo_Tech team, but we noticed that they were inappropriate for two reasons (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). Then, we decided to improve Prhl and obtain our original improved Prhl (included in BBa_K1949060) that suited our goal.
Characterize and Improve
Our purpose is to create strong Prhl for our final genetic circuits.
This experiment consists of the three parts below.
I. Improved Prhl by iGEM 2014 Tokyo_Tech team and characterization of rhlR
II. Improvement of the wild type Prhl
III. Comparison of the improved Prhl by iGEM 2014 Tokyo_Tech team to our original improved Prhl
The past improved Prhl did not suit for our final circuits and we could construct the improved Prhl appropriate to our final circuits.
I. Improved Prhl by iGEM 2014 Tokyo_Tech team and characterization of RhlR
We found that Prhl(RL) (BBa_K1529300) activity was weak and the expression level depended on LVA tag (Fig.1); LVA-tagged proteins are prone to be degraded by cellular proteases. Prhl(LR) (BBa_K1529310) activity was strong and unexpectedly reacted with C12 (crosstalk) (Fig.3).
The colonies of transformants with a rhlR (BBa_C0171) plasmid looked rough and the growth rate was low(Fig.2-left), while the colonies of transformants with a rhlR-LVA (BBa_C0071) plasmid looked smooth and the growth rate was normal(Fig.2-right). However, the reason for this result is unclear.
II. Improvement the wild type Prhl
By introducing a single point mutation into wild type Prhl (BBa_R0071) by PCR, we obtained 198 Prhl mutants and chose the two Prhl mutants of which promoter activity was stronger than wild type Prhl(Fig.3).
The sequences of these two chosen mutants are shown below. The small characters indicate the scar sequence and a single point mutation is colored with red.
Native Prhl (BBa_R0071)
TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTAAAAAGTGTTC
Prhl(NM) (BBa_K1949060)
TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTAtAAAGTGTTC
III. Comparison of the improved Prhl by iGEM 2014 Tokyo_Tech team to our original improved Prhl
Prhl(NM) was chosen from the many Prhl mutants, and by comparing Prhl(NM) to Prhl(LR), we obtained the result below (Fig.4). The reaction activity of Prhl(NM) to C4 was stronger than that of Prhl(LR), and Prhl(NM) did not react with C12 at all.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 723