Difference between revisions of "Part:BBa K1997012"
Zhuchushu13 (Talk | contribs) (→Special Design) |
|||
| (8 intermediate revisions by 2 users not shown) | |||
| Line 5: | Line 5: | ||
This part is an integrated tool for protein-protein interaction research using split-HRP system as reporter. the "Zif268" subpart can be easily replaced using Golden Gate technique with BsaI | This part is an integrated tool for protein-protein interaction research using split-HRP system as reporter. the "Zif268" subpart can be easily replaced using Golden Gate technique with BsaI | ||
| − | |||
===Usage and Biology=== | ===Usage and Biology=== | ||
| + | Intercellular protein–protein interactions (PPIs) enable communication between cells in diverse biological processes, including cell proliferation, immune responses, infection, and synaptic transmission, but they are challenging to visualize because existing techniques have insufficient sensitivity and/or specificity.<sup>1–3 </sup> | ||
| + | |||
| + | However, a split horseradish peroxidase (sHRP) has been reported as a sensitive and specific tool for the detection of intercellular PPIs. | ||
| + | |||
| + | The two sHRP fragments, engineered through screening of 17 cut sites in HRP followed by directed evolution, reconstitute into an active form when driven together by an intercellular PPI, producing bright fluorescence or contrast for electron microscopy. Fusing the sHRP fragments to the proteins neurexin (NRX) and neuroligin (NLG), which bind each other across the synaptic cleft<sup>4 </sup>, enabled sensitive visualization of synapses between specific sets of neurons, including two classes of synapses in the mouse visual system. sHRP should be widely applicable to studying mechanisms of communication between a variety of cell types. | ||
| + | |||
| + | ===Special Design=== | ||
| + | |||
| + | As a member of the collection PPI tool kit, special designs were taken for to optimize the applicability and adaptive of such parts. Specifically, a novel designed substitution system, through which, two proteins could be fused with their corresponding split-HRP fragment at the same time using Golden-Gate Assembly, was introduced to dramatically simplify the cloning process). | ||
| + | |||
| + | [[File:sHRPfig1.jpg|500px|]] | ||
| + | |||
| + | [[File:NUDT-012-2.jpg|500px|]] | ||
| + | |||
| + | Figure 1. Schematic representation of the workflow of the substitution system | ||
| + | |||
| + | Coding sequence of proteins to be studied can be assembled with a RBS in between, a PCR procedure adding a 5’-ATAGGGGAGACC-3’ flank to the sense strand and a 3’-TCCAGAGTCAAA-5’ flank to the anti-sense would make it a proper substrate for the BsaI nuclease digest. Once digested, the product could be ligated together with the BsaI treated BBa_K1997012 to form a brand new device expressing the proteins of sHRP-N-Protein1, Protein2-sHRP-C and. The interaction between Protein1 and protein 2 could then be determined through the value of OD<sub>450 </sub> | ||
| + | |||
| + | ===Sequence and Features=== | ||
<!-- --> | <!-- --> | ||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
<partinfo>BBa_K1997012 SequenceAndFeatures</partinfo> | <partinfo>BBa_K1997012 SequenceAndFeatures</partinfo> | ||
| + | ==Experimental Validation== | ||
| − | + | This part is validated through four ways: enzyme cutting, PCR, Sequence, and functional testing | |
| − | ===Functional | + | |
| − | < | + | ===Sequencing=== |
| − | + | ||
| + | This part is sequenced as correct after construction. | ||
| + | |||
| + | ===PCR=== | ||
| + | |||
| + | '''Methods''' | ||
| + | |||
| + | The PCR is performed with Premix EX Taq by Takara. | ||
| + | |||
| + | F-Prime: 5’- GAATTCGCGGCCGCTTCTAGAATGC-3’ | ||
| + | |||
| + | R-Prime: 5’- GGACTAGTATTATTGTTTGTCTGCC-3’ | ||
| + | |||
| + | The PCR protocol is selected based on the Users Manuel. | ||
| + | The Electrophoresis was performed on a 1% Agarose glu. | ||
| + | The result of the agarose electrophoresis was shown on the picture below. | ||
| + | |||
| + | [[File:NUDT-012-1.jpg|300px|]] | ||
| + | |||
| + | ===Enzyme digestion test === | ||
| + | |||
| + | '''Methods''' | ||
| + | |||
| + | After the assembly ,the plasmid was transferred into the Competent ''E. coli'' DH5α). After culturing overnight in LB,we minipreped the plasmid for cutting. | ||
| + | The preparation of the plasmid was performed with TIANprep Mini Plasmid Kit from ''TIANGEN''. The cutting procedure was performed with EcoRI and SpeI restriction endonuclease bought from ''TAKARA''. | ||
| + | |||
| + | The plasmid was cutted in a 20μL system at 37 ℃ for 2 hours. | ||
| + | The Electrophoresis was performed on a 1% Agarose glu. | ||
| + | |||
| + | The result of the agarose electrophoresis was shown on the picture above. | ||
| + | |||
| + | ===Functional Test=== | ||
| + | |||
| + | This part was tested together with K1997013 using K1997013 as control. | ||
| + | |||
| + | To proof the basic function of our split-HRP reporting system, two devices, containing split-HRP fragments and a complete or split zinc finger protein, were built under control of a lac operon controlled T7 promoter. The complete zinc finger protein was to stimulate a PPI positive situation, while the split one was to stimulate a PPI negative situation. For such assay, E.coli carrying respective plasmid was cultured overnight under IPTG induction. Cells were then collected and lysed by high-pressure homogenizer. Once lysed and ultra-filtrated (to remove small molecules), TMB substrate solution (with heme supplementation) were added into the cell lysate for the measurement of HRP activity. The plate was then incubated under 37°C for 5min before adding the Stop solution. | ||
| + | |||
| + | Value of OD<sub>450 </sub> obtained after adding the Stop Solution showed significant variation between the PPI positive group and the PPI negative group. Thus then validated the function of this part. | ||
| + | |||
| + | [[File:T--NUDT_CHINA--partsfig4.jpg|700px|]] | ||
| + | |||
| + | Figure 2. Evaluation of the Signal-Noise Ratio of split HRP system (A) Schematic representation of the evaluation protocol. The complete Zif-268 protein was introduced to simulate the condition where strong interaction among two proteins occur, whereas the split-zif protein was used to simulate the condition where no interaction exists. (B) Experiment showing the OD<sub>450 </sub> value under two different conditions. Relative OD<sub>450 </sub> value was calculated with normalization of the OD<sub>450 </sub> value. This experiment was run in three parallel reactions, and the data represent results obtained from at least three independent experiments. **p<0.01. | ||
| + | |||
| + | To further demonstrate the substitution system, we replaced the Zif268 region in this part into a FRB-RBS-FKBP fragment. The further experimental validation can be seen on BBa_K1997011. | ||
| + | |||
| + | ===References=== | ||
| + | |||
| + | 1. Kim, S.A., Tai, C.-Y., Mok, L.-P., Mosser, E.A. & Schuman, E.M. Calcium-dependent | ||
| + | dynamics of cadherin interactions at cell-cell junctions. Proc. Natl. Acad. Sci. USA | ||
| + | 108, 9857–9862 (2011). | ||
| + | |||
| + | 2. Feinberg, E.H. et al. GFP Reconstitution Across Synaptic Partners (GRASP) defnes | ||
| + | cell contacts and synapses in living nervous systems. Neuron 57, 353–363 (2008). | ||
| + | |||
| + | 3. Liu, D.S., Loh, K.H., Lam, S.S., White, K.A. & Ting, A.Y. Imaging trans-cellular neurexinneuroligin interactions by enzymatic probe ligation. PLoS One 8, e52823 (2013). | ||
| + | |||
| + | 4. Craig, A.M. & Kang, Y. Neurexin-neuroligin signaling in synapse development. Curr. | ||
| + | Opin. Neurobiol. 17, 43–52 (2007). | ||
Latest revision as of 01:18, 21 October 2016
P+R->sHRP-N->Zif268-> sHRP-C->TER
This part is an integrated tool for protein-protein interaction research using split-HRP system as reporter. the "Zif268" subpart can be easily replaced using Golden Gate technique with BsaI
Usage and Biology
Intercellular protein–protein interactions (PPIs) enable communication between cells in diverse biological processes, including cell proliferation, immune responses, infection, and synaptic transmission, but they are challenging to visualize because existing techniques have insufficient sensitivity and/or specificity.1–3
However, a split horseradish peroxidase (sHRP) has been reported as a sensitive and specific tool for the detection of intercellular PPIs.
The two sHRP fragments, engineered through screening of 17 cut sites in HRP followed by directed evolution, reconstitute into an active form when driven together by an intercellular PPI, producing bright fluorescence or contrast for electron microscopy. Fusing the sHRP fragments to the proteins neurexin (NRX) and neuroligin (NLG), which bind each other across the synaptic cleft4 , enabled sensitive visualization of synapses between specific sets of neurons, including two classes of synapses in the mouse visual system. sHRP should be widely applicable to studying mechanisms of communication between a variety of cell types.
Special Design
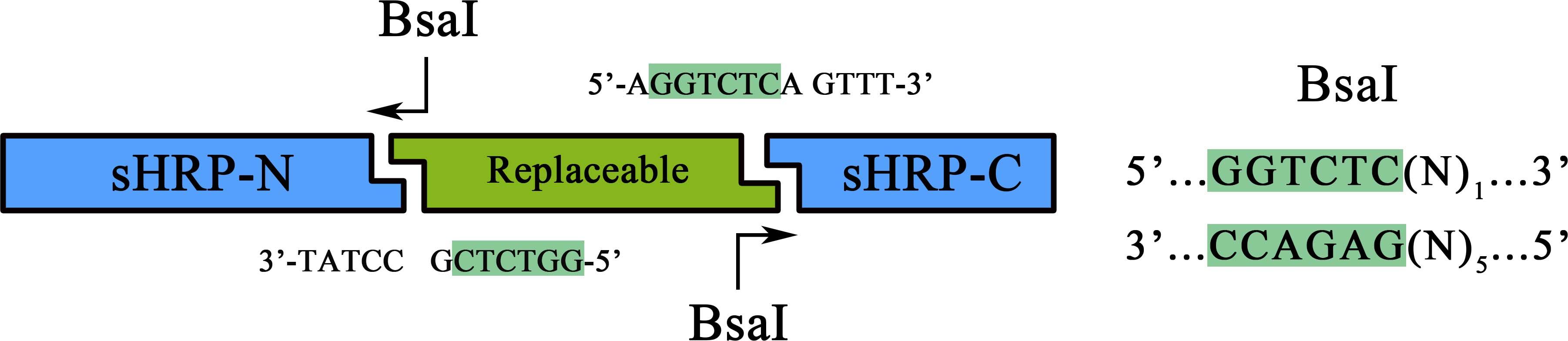
As a member of the collection PPI tool kit, special designs were taken for to optimize the applicability and adaptive of such parts. Specifically, a novel designed substitution system, through which, two proteins could be fused with their corresponding split-HRP fragment at the same time using Golden-Gate Assembly, was introduced to dramatically simplify the cloning process).
Figure 1. Schematic representation of the workflow of the substitution system
Coding sequence of proteins to be studied can be assembled with a RBS in between, a PCR procedure adding a 5’-ATAGGGGAGACC-3’ flank to the sense strand and a 3’-TCCAGAGTCAAA-5’ flank to the anti-sense would make it a proper substrate for the BsaI nuclease digest. Once digested, the product could be ligated together with the BsaI treated BBa_K1997012 to form a brand new device expressing the proteins of sHRP-N-Protein1, Protein2-sHRP-C and. The interaction between Protein1 and protein 2 could then be determined through the value of OD450
Sequence and Features
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 1171
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1174
Illegal BsaI.rc site found at 907
Illegal SapI.rc site found at 1288
Experimental Validation
This part is validated through four ways: enzyme cutting, PCR, Sequence, and functional testing
Sequencing
This part is sequenced as correct after construction.
PCR
Methods
The PCR is performed with Premix EX Taq by Takara.
F-Prime: 5’- GAATTCGCGGCCGCTTCTAGAATGC-3’
R-Prime: 5’- GGACTAGTATTATTGTTTGTCTGCC-3’
The PCR protocol is selected based on the Users Manuel. The Electrophoresis was performed on a 1% Agarose glu. The result of the agarose electrophoresis was shown on the picture below.
Enzyme digestion test
Methods
After the assembly ,the plasmid was transferred into the Competent E. coli DH5α). After culturing overnight in LB,we minipreped the plasmid for cutting. The preparation of the plasmid was performed with TIANprep Mini Plasmid Kit from TIANGEN. The cutting procedure was performed with EcoRI and SpeI restriction endonuclease bought from TAKARA.
The plasmid was cutted in a 20μL system at 37 ℃ for 2 hours. The Electrophoresis was performed on a 1% Agarose glu.
The result of the agarose electrophoresis was shown on the picture above.
Functional Test
This part was tested together with K1997013 using K1997013 as control.
To proof the basic function of our split-HRP reporting system, two devices, containing split-HRP fragments and a complete or split zinc finger protein, were built under control of a lac operon controlled T7 promoter. The complete zinc finger protein was to stimulate a PPI positive situation, while the split one was to stimulate a PPI negative situation. For such assay, E.coli carrying respective plasmid was cultured overnight under IPTG induction. Cells were then collected and lysed by high-pressure homogenizer. Once lysed and ultra-filtrated (to remove small molecules), TMB substrate solution (with heme supplementation) were added into the cell lysate for the measurement of HRP activity. The plate was then incubated under 37°C for 5min before adding the Stop solution.
Value of OD450 obtained after adding the Stop Solution showed significant variation between the PPI positive group and the PPI negative group. Thus then validated the function of this part.
Figure 2. Evaluation of the Signal-Noise Ratio of split HRP system (A) Schematic representation of the evaluation protocol. The complete Zif-268 protein was introduced to simulate the condition where strong interaction among two proteins occur, whereas the split-zif protein was used to simulate the condition where no interaction exists. (B) Experiment showing the OD450 value under two different conditions. Relative OD450 value was calculated with normalization of the OD450 value. This experiment was run in three parallel reactions, and the data represent results obtained from at least three independent experiments. **p<0.01.
To further demonstrate the substitution system, we replaced the Zif268 region in this part into a FRB-RBS-FKBP fragment. The further experimental validation can be seen on BBa_K1997011.
References
1. Kim, S.A., Tai, C.-Y., Mok, L.-P., Mosser, E.A. & Schuman, E.M. Calcium-dependent dynamics of cadherin interactions at cell-cell junctions. Proc. Natl. Acad. Sci. USA 108, 9857–9862 (2011).
2. Feinberg, E.H. et al. GFP Reconstitution Across Synaptic Partners (GRASP) defnes cell contacts and synapses in living nervous systems. Neuron 57, 353–363 (2008).
3. Liu, D.S., Loh, K.H., Lam, S.S., White, K.A. & Ting, A.Y. Imaging trans-cellular neurexinneuroligin interactions by enzymatic probe ligation. PLoS One 8, e52823 (2013).
4. Craig, A.M. & Kang, Y. Neurexin-neuroligin signaling in synapse development. Curr. Opin. Neurobiol. 17, 43–52 (2007).