Difference between revisions of "Part:BBa K1321014"
| (8 intermediate revisions by 2 users not shown) | |||
| Line 8: | Line 8: | ||
At present the cloning for these constructs is still in progress to correct an illegal EcoRI site which was identified in the parts with this CBD. This can be achieved with a silent mutation via site-directed mutagenesis and we aim to send these parts to the registry once this is complete. | At present the cloning for these constructs is still in progress to correct an illegal EcoRI site which was identified in the parts with this CBD. This can be achieved with a silent mutation via site-directed mutagenesis and we aim to send these parts to the registry once this is complete. | ||
| + | In our first assay performed to determine the relative strengths of various CBDs’ binding to bacterial cellulose – represented by the percentage fluorescence left from CBDs fused to [https://parts.igem.org/Part:BBa_K1321337 sfGFP (RFC25)] bound to bacterial cellulose discs, when subjected to various washes (protocol [http://2014.igem.org/Team:Imperial/Protocols here]) – it was determined that the CBDcipA-sfGFP fusion had the greatest binding ability in comparison to the four other CBDs fused to sfGFP after three washes with 5% BSA. | ||
| + | |||
| + | In the same assay, results suggested that CBDcipA that on average overall it had the second best ability to bind bacterial cellulose, as shown by the washes with PBS, 70% EtOH and dH2O. | ||
| + | |||
| + | [[File:IC14_-_dH2Obplot1.png|700px|centre|]] | ||
| + | |||
| + | [[File:IC14-EtOHbplot1.png|700px|centre|]] | ||
| + | |||
| + | [[File:IC14-PBSbplot1.png|700px|centre|]] | ||
| + | |||
| + | [[File:IC14-BSAbplot1.png|700px|centre|]] | ||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
===Usage and Biology=== | ===Usage and Biology=== | ||
| + | |||
<!-- --> | <!-- --> | ||
Latest revision as of 04:56, 2 November 2014
CBDCipA with N and C-terminal linker
This CBD is from the Cellulosomal -scaffolding protein A (cipA) of Clostridium thermocellum including the endogenous linker sequences at the N and C-terminus (UniProt ID Q06851 link http://www.uniprot.org/uniprot/Q06851). It is in RFC25 format to allow for easy use in protein fusions.
The CBD is part of the CBM3 family (link to cazy http://www.cazy.org/CBM3.html). This CBD has been used in many application: in fusions with cell adhesion peptides to enhance the properties of cellulose as a cell-growth matrix (Andrade et al 2010a, Andrade et al 2010b), fused to enzymes to remove contaminants from water (Kauffmann et al 2000) and fused to an antimicrobial peptide (Ramos, Domingues & Gama 2010).
At present the cloning for these constructs is still in progress to correct an illegal EcoRI site which was identified in the parts with this CBD. This can be achieved with a silent mutation via site-directed mutagenesis and we aim to send these parts to the registry once this is complete.
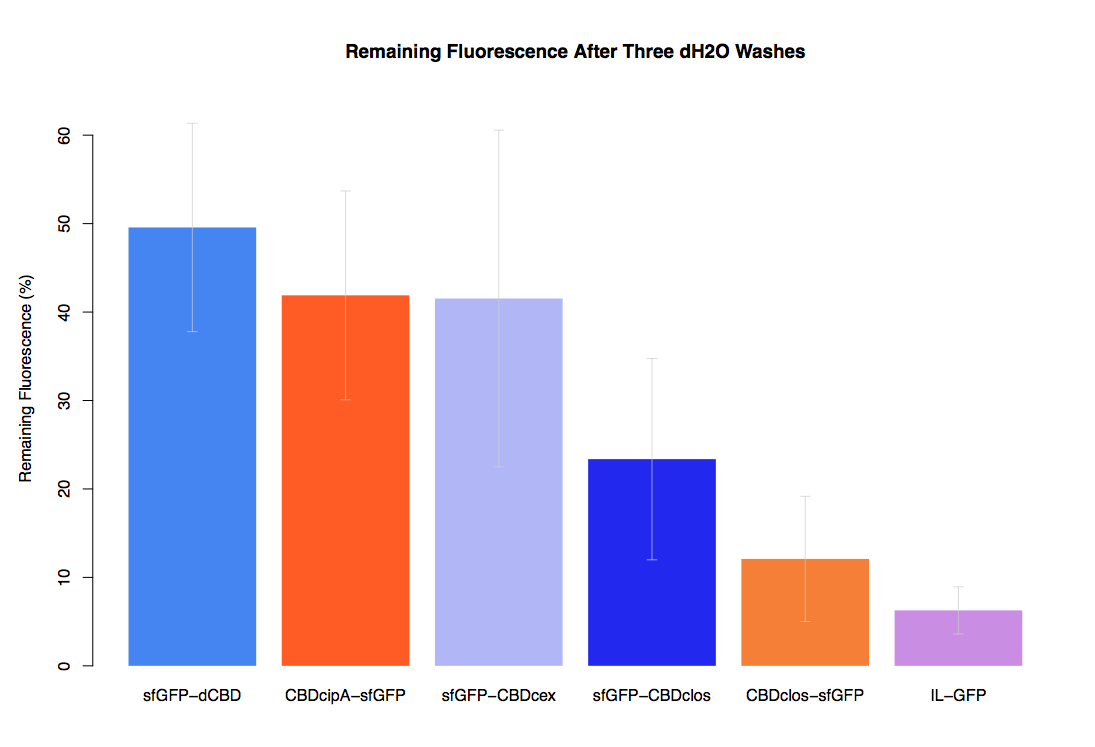
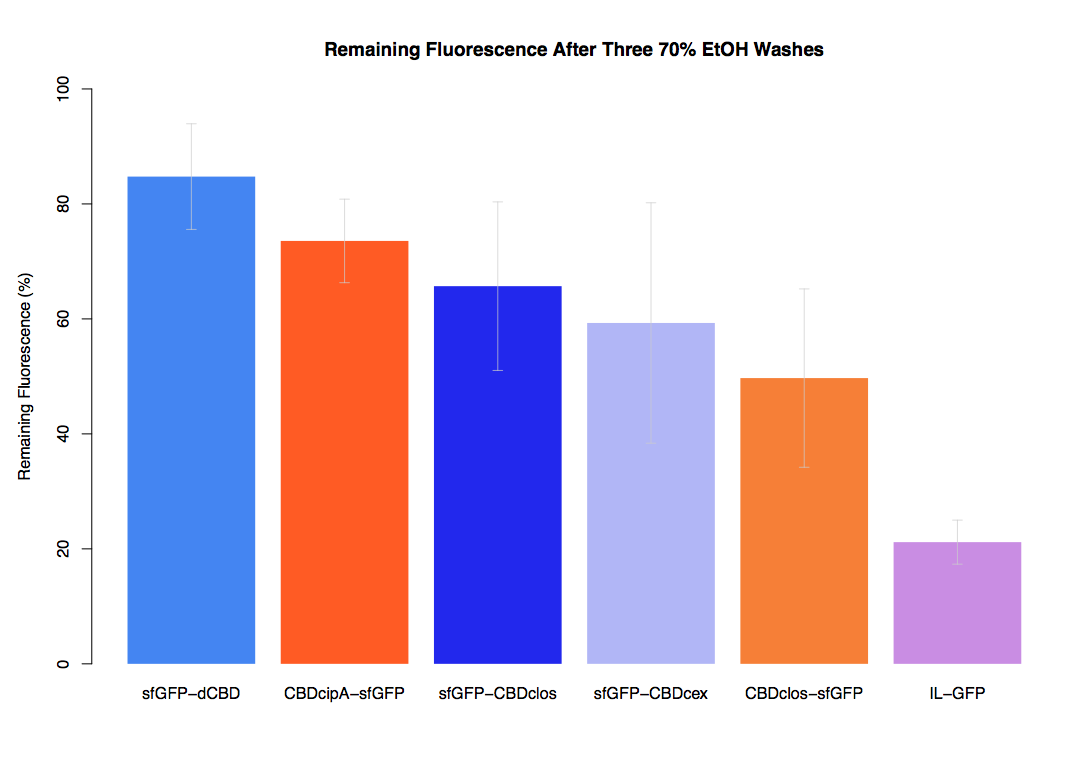
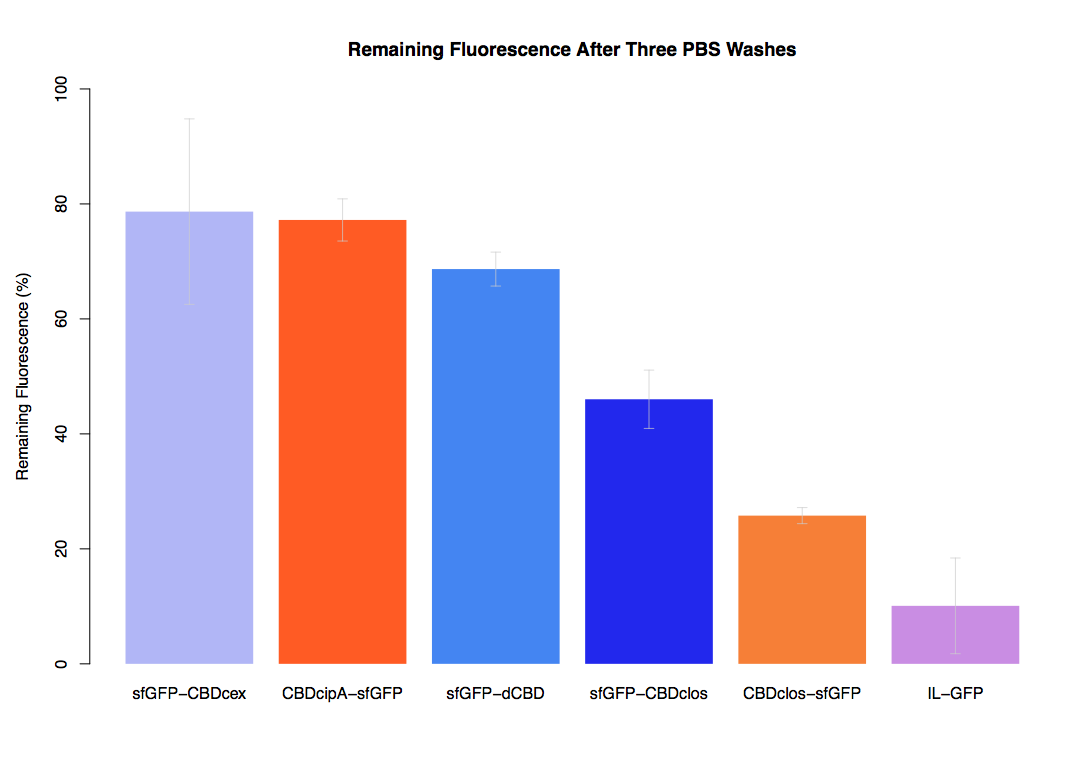
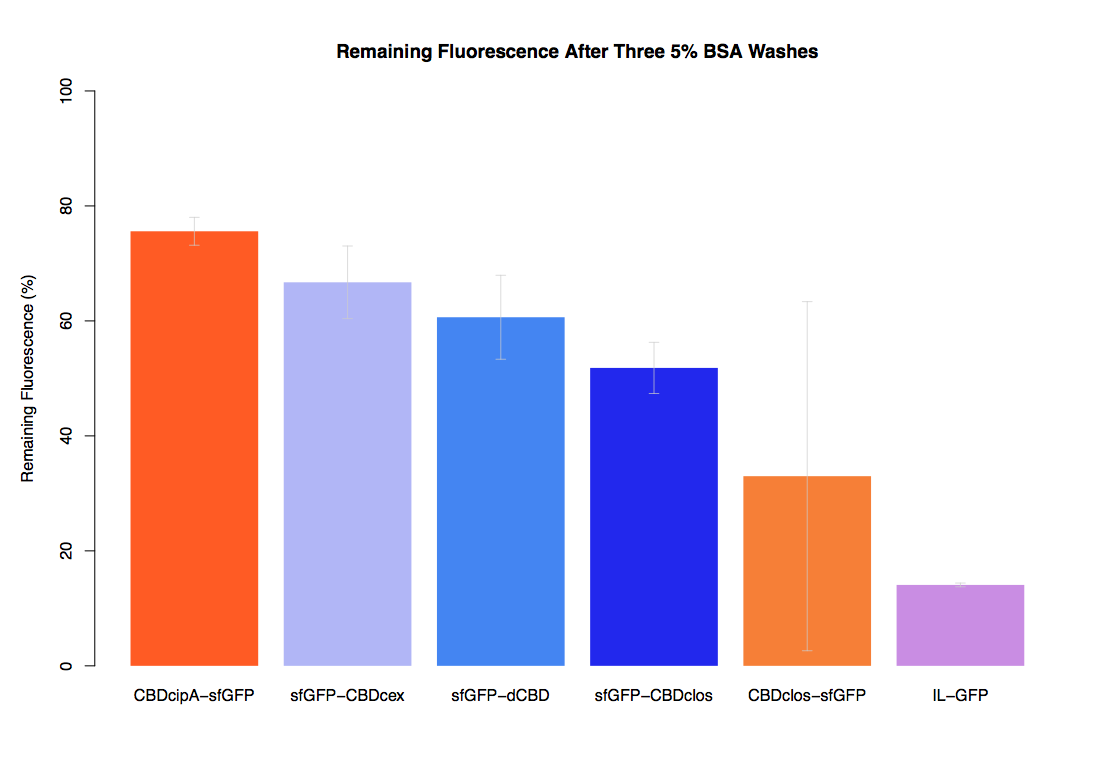
In our first assay performed to determine the relative strengths of various CBDs’ binding to bacterial cellulose – represented by the percentage fluorescence left from CBDs fused to sfGFP (RFC25) bound to bacterial cellulose discs, when subjected to various washes (protocol [http://2014.igem.org/Team:Imperial/Protocols here]) – it was determined that the CBDcipA-sfGFP fusion had the greatest binding ability in comparison to the four other CBDs fused to sfGFP after three washes with 5% BSA.
In the same assay, results suggested that CBDcipA that on average overall it had the second best ability to bind bacterial cellulose, as shown by the washes with PBS, 70% EtOH and dH2O.
Sequence and Features
- 10INCOMPATIBLE WITH RFC[10]Illegal EcoRI site found at 148
- 12INCOMPATIBLE WITH RFC[12]Illegal EcoRI site found at 148
- 21INCOMPATIBLE WITH RFC[21]Illegal EcoRI site found at 148
- 23INCOMPATIBLE WITH RFC[23]Illegal EcoRI site found at 148
- 25INCOMPATIBLE WITH RFC[25]Illegal EcoRI site found at 148
- 1000COMPATIBLE WITH RFC[1000]
References
Andrade, F.K., Moreira, S.M.G., Domingues, L. & Gama, F.M.P. (2010) Improving the affinity of fibroblasts for bacterial cellulose using carbohydrate-binding modules fused to RGD. Journal of biomedical materials research. Part A. [Online] 92 (1), 9–17. Available from: doi:10.1002/jbm.a.32284.
Andrade, F.K., Costa, R., Domingues, L., Soares, R., et al. (2010) Improving bacterial cellulose for blood vessel replacement: Functionalization with a chimeric protein containing a cellulose-binding module and an adhesion peptide. Acta biomaterialia. [Online] 6 (10), 4034–4041. Available from: doi:10.1016/j.actbio.2010.04.023.
Kauffmann, C., Shoseyov, O., Shpigel, E., Bayer, E.A., et al. (2000) Novel Methodology for Enzymatic Removal of Atrazine from Water by CBD-Fusion Protein Immobilized on Cellulose. Environmental Science & Technology. [Online] 34 (7), 1292–1296. Available from: doi:10.1021/es990754h.
Ramos, R., Domingues, L. & Gama, M. (2010) Escherichia coli expression and purification of LL37 fused to a family III carbohydrate-binding module from Clostridium thermocellum. Protein expression and purification. [Online] 71 (1), 1–7. Available from: doi:10.1016/j.pep.2009.10.016.