Difference between revisions of "Part:BBa K1028003"
(→Scanning Electron Microscopy Imaging (SEM)) |
|||
| (12 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K1028003 short</partinfo> | <partinfo>BBa_K1028003 short</partinfo> | ||
| − | + | ==Introduction== | |
<html> | <html> | ||
<img src="http://i276.photobucket.com/albums/kk29/JosephBeetBeet/NEWBS3Image.png" style="width:600px;margin-right:160px" ></a> | <img src="http://i276.photobucket.com/albums/kk29/JosephBeetBeet/NEWBS3Image.png" style="width:600px;margin-right:160px" ></a> | ||
| Line 8: | Line 8: | ||
</p></html> | </p></html> | ||
| − | + | This part is a multi-system BioBrick designed by the Leeds 2013 iGEM team to bind specifically to silica macro structures via the use of the INP.Si4 complex ([https://parts.igem.org/Part:BBa_K1028001 Part BBa_K1028001]) with high affinity and specificity. In unison with the binding GFP production in maintained under the control of the CpxR promoter ([https://parts.igem.org/wiki/index.php?title=Part:BBa_K1028002 Part BBa_K1028002]) which should only be highly activated upon binding with the silica macro structures. | |
| + | |||
| + | This system, affectionately coined to MicroBeagle, is an iGEM first in that it acts as a detection system through physical binding of the entire cell to its target substrate, which activates the reporter gene. The system was designed with modularity in mind, silica binding was used a surrogate for pathogen detection with a view to using the MicroBeagle as a water purification testing device. However the Si4 binding domain can easily be exchanged for any other short-medium length binding sequence making this part, and general ideology for detection, a very useful starting point for future iGEM teams looking to create systems for detection of macro structures in solution. | ||
| + | |||
| + | ==Experimental Data Concerning the Binding Domain== | ||
| + | |||
| + | ====Fluorescent Activated Cell Sorting (FACS) ==== | ||
| + | |||
| + | <html> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/thumb/b/be/Leeds_FACS_Overall_Information_Pic.png/800px-Leeds_FACS_Overall_Information_Pic.png" style="width:800px;margin-right:160px" ></a> | ||
| + | |||
| + | <p style="text-align:center"><b>Fig.2:</b> INP.Si4, labelled here as Device 2 which also contains GFP coding region under the control of the CpxR promoter, characterized using FACS. This data shows successful binding of a small portion of our expression cells to the silica beads used, this result is discussed in more depth below. | ||
| + | </p></html> | ||
| + | |||
| + | Device 2 contains our INP+Si4 construct, along with the Cpx and GFP genes. It was characterized using a flow cytometry method called Fluorescence-activated cell sorting (FACS) in solution with and without beads. The FACS images contain Side scattering and Forward scattering information, relating to relative size and density of the cells. | ||
| + | |||
| + | In the assay containing Device 2 with beads, there was an increase in the percentage of Population 3 (coloured in blue), from 0.1 to 2.2. This is suggestive of representing our Device 2 cells bound to silica beads. In contrast, the assay, which contained only Device 2 cells, had a very insignificant percentage of Population 3 (0.1%). | ||
| + | |||
| + | This suggests that our Device may successfully be binding to the silica beads utilizing the INP+Si4 construct, however not in all cases. This could either be due to not having enough time for the Device to binding to the silica, or that our Si4 binding domain is not as effective as it should be. | ||
| + | |||
| + | ====Scanning Electron Microscopy Imaging (SEM)==== | ||
| + | |||
| + | |||
| + | <html> | ||
| + | <img src="http://i276.photobucket.com/albums/kk29/JosephBeetBeet/SEMImage2FromOleg-1mMTeOS.png" style="width:900px;margin-right:160px" ></a> | ||
| + | |||
| + | <p style="text-align:left"><b>Fig.3</b> SEM image showing expression cells containing Device 2 in solution with 1mM TeOS. | ||
| + | </p></html> | ||
| + | |||
| + | The image above contains images that were produced by the Scanning Electron Microscope (SEM) at the University of Leeds. Device 2 was incubated in solution of varying concentrations of an alkylsilane compound, which was solubilized using the sol-gel method from Tetraethoxysilane (TEOS). Samples were incubated at room temperature, as well as some were done at 45ºC. For the control samples, cells without the Device 2 construct gene were used, and did not react with the alkylsilane compound. | ||
| + | |||
| + | The alkylsilane compound forming distinct spheres can be witnessed in one of the images. EDX chemical composition data is also included with each image, which allows assessment of the elements that are present in each sample. The high carbon output is partially due to the carbon adhesive used to isolate the samples for SEM. The alkylsilane compound would envelop the cells either fully or partially and this can be gathered from the images shown. The images of control samples exemplify the alkylsilane compound not extensively bound to the cells. | ||
| + | |||
| + | ==Experimental Data Regarding the Reporter System== | ||
| + | <html> | ||
| + | <img src="http://i276.photobucket.com/albums/kk29/JosephBeetBeet/BS1CharacterisationBars.png" style="width:700px;margin-left:160px" ></a> | ||
| + | |||
| + | <p style="text-align:center"><b>Fig.3:</b> The effects of a range of both physical and chemical stresses on the plasma membrane and their effect on activation of the CpxR promoter | ||
| + | </p></html> | ||
| + | <br> | ||
| + | Individually many of the experiments produced erratic and seemingly random data which is listed independently below. However plotting all average readings together shows that there is a dramatic increase in response to the stress induced by the silica beads comparative to all the chemically induced membrane stresses we used. | ||
| + | |||
| + | This indicates that it would be possible to detect the presence of silica beads in solution, in principle, using our binding and detection system, certainly when silica beads are present at relatively high concentrations. However further work needs to be done to assess whether this high response remains across a range of silica bead concentrations in solution. | ||
| + | |||
| + | In light of these results, some of these experiments were repeated for more accurate results. | ||
| + | How the Cpx pathway reacts to temperature is already well characterised in the literature and by the [http://2010.igem.org/Team:Calgary/Parts/Characterization Calgary 2010 iGem team.] | ||
| + | |||
| + | The Ethanol gradient, Triton X-100 gradient and pH gradient were all repeated. | ||
| + | In the second experiment, a robotic pipette was used to directly fill the 96 well plate to reduce human error. | ||
| + | Excluding wells with imposed pH gradient, all wells were kept at pH 7 using Tris.HCl buffer, the pH gradient was also achieved with a Tris.HCl buffer. | ||
| + | |||
| + | The 96 well plates were then incubated while shaking while shaking for 2 hours at 37 degrees to allow time for GFP to mature. A longer incubation time was introduced to give more conclusive results. | ||
| + | We also performed preliminary investigation on the effects of ethanol and temperature extremes on the CpxR promoter but the results were inconclusive warranting further studies, possibly by the next years Leeds iGEM team. | ||
| + | <br> | ||
| + | <br> | ||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
===Usage and Biology=== | ===Usage and Biology=== | ||
Latest revision as of 18:23, 4 October 2013
Silica Detection System
Introduction
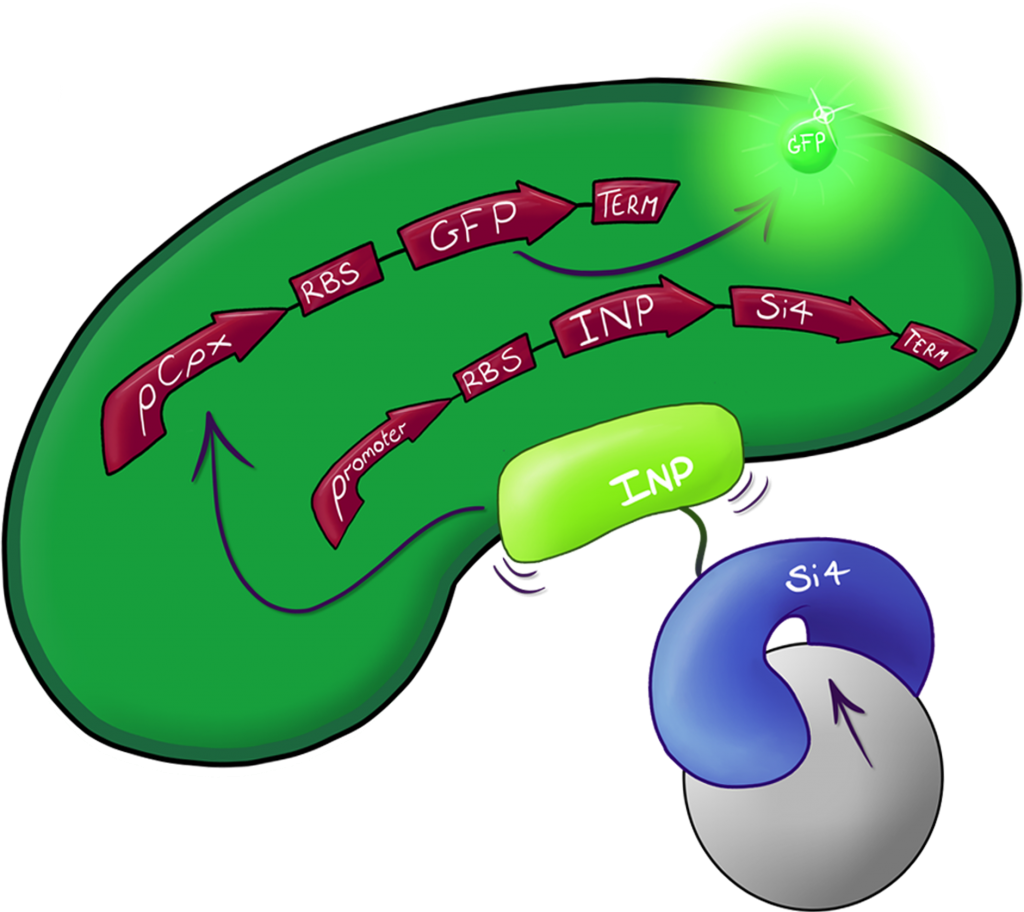
Fig.2: The Culmination of the Leeds 2013 iGEM project, a system capable of both binding and detecting silica particles in solution.
This part is a multi-system BioBrick designed by the Leeds 2013 iGEM team to bind specifically to silica macro structures via the use of the INP.Si4 complex (Part BBa_K1028001) with high affinity and specificity. In unison with the binding GFP production in maintained under the control of the CpxR promoter (Part BBa_K1028002) which should only be highly activated upon binding with the silica macro structures.
This system, affectionately coined to MicroBeagle, is an iGEM first in that it acts as a detection system through physical binding of the entire cell to its target substrate, which activates the reporter gene. The system was designed with modularity in mind, silica binding was used a surrogate for pathogen detection with a view to using the MicroBeagle as a water purification testing device. However the Si4 binding domain can easily be exchanged for any other short-medium length binding sequence making this part, and general ideology for detection, a very useful starting point for future iGEM teams looking to create systems for detection of macro structures in solution.
Experimental Data Concerning the Binding Domain
Fluorescent Activated Cell Sorting (FACS)

Fig.2: INP.Si4, labelled here as Device 2 which also contains GFP coding region under the control of the CpxR promoter, characterized using FACS. This data shows successful binding of a small portion of our expression cells to the silica beads used, this result is discussed in more depth below.
Device 2 contains our INP+Si4 construct, along with the Cpx and GFP genes. It was characterized using a flow cytometry method called Fluorescence-activated cell sorting (FACS) in solution with and without beads. The FACS images contain Side scattering and Forward scattering information, relating to relative size and density of the cells.
In the assay containing Device 2 with beads, there was an increase in the percentage of Population 3 (coloured in blue), from 0.1 to 2.2. This is suggestive of representing our Device 2 cells bound to silica beads. In contrast, the assay, which contained only Device 2 cells, had a very insignificant percentage of Population 3 (0.1%).
This suggests that our Device may successfully be binding to the silica beads utilizing the INP+Si4 construct, however not in all cases. This could either be due to not having enough time for the Device to binding to the silica, or that our Si4 binding domain is not as effective as it should be.
Scanning Electron Microscopy Imaging (SEM)
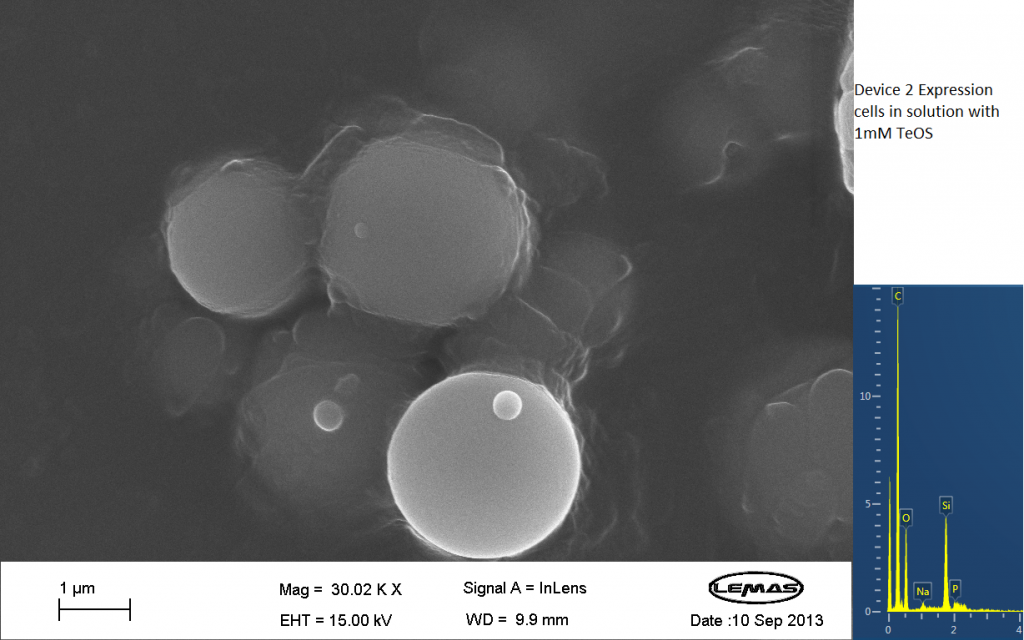

Fig.3 SEM image showing expression cells containing Device 2 in solution with 1mM TeOS.
The image above contains images that were produced by the Scanning Electron Microscope (SEM) at the University of Leeds. Device 2 was incubated in solution of varying concentrations of an alkylsilane compound, which was solubilized using the sol-gel method from Tetraethoxysilane (TEOS). Samples were incubated at room temperature, as well as some were done at 45ºC. For the control samples, cells without the Device 2 construct gene were used, and did not react with the alkylsilane compound.
The alkylsilane compound forming distinct spheres can be witnessed in one of the images. EDX chemical composition data is also included with each image, which allows assessment of the elements that are present in each sample. The high carbon output is partially due to the carbon adhesive used to isolate the samples for SEM. The alkylsilane compound would envelop the cells either fully or partially and this can be gathered from the images shown. The images of control samples exemplify the alkylsilane compound not extensively bound to the cells.
Experimental Data Regarding the Reporter System
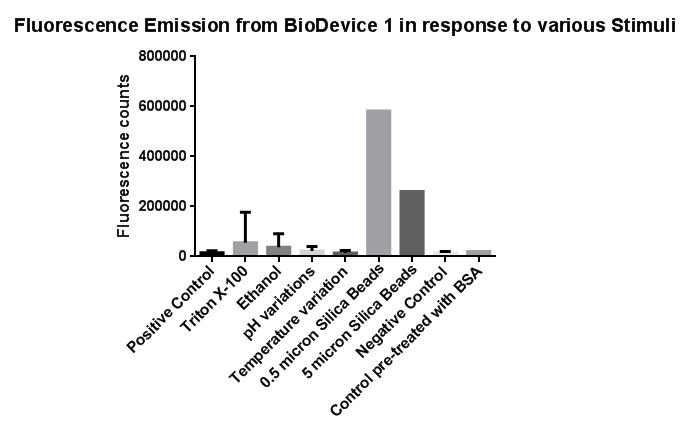
Fig.3: The effects of a range of both physical and chemical stresses on the plasma membrane and their effect on activation of the CpxR promoter
Individually many of the experiments produced erratic and seemingly random data which is listed independently below. However plotting all average readings together shows that there is a dramatic increase in response to the stress induced by the silica beads comparative to all the chemically induced membrane stresses we used.
This indicates that it would be possible to detect the presence of silica beads in solution, in principle, using our binding and detection system, certainly when silica beads are present at relatively high concentrations. However further work needs to be done to assess whether this high response remains across a range of silica bead concentrations in solution.
In light of these results, some of these experiments were repeated for more accurate results. How the Cpx pathway reacts to temperature is already well characterised in the literature and by the [http://2010.igem.org/Team:Calgary/Parts/Characterization Calgary 2010 iGem team.]
The Ethanol gradient, Triton X-100 gradient and pH gradient were all repeated. In the second experiment, a robotic pipette was used to directly fill the 96 well plate to reduce human error. Excluding wells with imposed pH gradient, all wells were kept at pH 7 using Tris.HCl buffer, the pH gradient was also achieved with a Tris.HCl buffer.
The 96 well plates were then incubated while shaking while shaking for 2 hours at 37 degrees to allow time for GFP to mature. A longer incubation time was introduced to give more conclusive results.
We also performed preliminary investigation on the effects of ethanol and temperature extremes on the CpxR promoter but the results were inconclusive warranting further studies, possibly by the next years Leeds iGEM team.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 62
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 472
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 1917
