Difference between revisions of "Part:BBa K1100014"
(→Manipulating and Experimental Data (Quantitative Measurement)) |
(→Our design of AleaderT) |
||
| (16 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K1100014 short</partinfo> | <partinfo>BBa_K1100014 short</partinfo> | ||
| + | |||
| + | '''''The first artificial triphase riboswitch.''''' | ||
ALeaderT is a novel riboswitch transformed from the leader RNA of the aminoglycoside resistance genes. It has a low minimal level and a high dynamic range, with a low noise in the system and minimal difference under different regulation. | ALeaderT is a novel riboswitch transformed from the leader RNA of the aminoglycoside resistance genes. It has a low minimal level and a high dynamic range, with a low noise in the system and minimal difference under different regulation. | ||
| Line 23: | Line 25: | ||
1.Riboswitch Background. | 1.Riboswitch Background. | ||
| − | Riboswitches are regulatory RNAs that regulate gene expression by binding small molecules. Recently, a novel riboswitch was reported which is present in the leader RNA of the aminoglycoside resistance genes that encode the aminoglycoside acetyl transferase (AAC) and aminoglycoside adenyl transferase (AAD) enzymes. When the aminoglycosides bind to the leader RNA, a conformational change will be induced, leading to the expression of the aminoglycoside resistance genes (Jia X et al, 2013.). In addition, the leader RNA encompass an integrase site (attl1), which is overlapped by a short peptide encoded by an ORF (ORF11) ( | + | Riboswitches are regulatory RNAs that regulate gene expression by binding small molecules. Recently, a novel riboswitch was reported which is present in the leader RNA of the aminoglycoside resistance genes that encode the aminoglycoside acetyl transferase (AAC) and aminoglycoside adenyl transferase (AAD) enzymes. When the aminoglycosides bind to the leader RNA, a conformational change will be induced, leading to the expression of the aminoglycoside resistance genes (Jia X et al, 2013.). In addition, the leader RNA encompass an integrase site (attl1), which is overlapped by a short peptide encoded by an ORF (ORF11) (Berçot B, et al, 2002). Therefore, Aleader is a complicated riboswitch containing ORF11, cistron, aminoglycoside-binding domain and att1 recombination site. Due to the complicacy of Aleader, the secondary structure of it could not be predicted by most software such as Mfold and NUPACK (Jia X et al, 2013.). |
[[File:AleaderT1.jpg|600px|thumb|center| '''Figure1'''. Proposed Model for the Induction of Aminoglycoside Resistance: Aminoglycoside binding to the leader RNA induces a change in the leader RNA structure]] | [[File:AleaderT1.jpg|600px|thumb|center| '''Figure1'''. Proposed Model for the Induction of Aminoglycoside Resistance: Aminoglycoside binding to the leader RNA induces a change in the leader RNA structure]] | ||
| Line 35: | Line 37: | ||
Take the advantages of Csy4, by physically separating linked genetic elements with cleavage sites, we can achieve assembly of promoters, ribosome binding sites, cis-regulatory elements, or even multigene operons with predictable and standardized functions in bacteria. In other words, it has nice compatibility that enables Csy4 to cooperate with other effective elements to produce new functions ( Lei Qi et al, 2012). | Take the advantages of Csy4, by physically separating linked genetic elements with cleavage sites, we can achieve assembly of promoters, ribosome binding sites, cis-regulatory elements, or even multigene operons with predictable and standardized functions in bacteria. In other words, it has nice compatibility that enables Csy4 to cooperate with other effective elements to produce new functions ( Lei Qi et al, 2012). | ||
| − | |||
| − | |||
== Our design of AleaderT == | == Our design of AleaderT == | ||
| Line 54: | Line 54: | ||
''k2'':distribution constant between different conformations ]] | ''k2'':distribution constant between different conformations ]] | ||
That shows the leakage expression of triphase riboswitch significantly decrease compared to the biphase riboswitch. | That shows the leakage expression of triphase riboswitch significantly decrease compared to the biphase riboswitch. | ||
| + | |||
| + | Want to know about our triphase model? Click [http://2013.igem.org/Team:Fudan/triphase_riboswitch.html '''Here'''] | ||
Also, via adding a csy4 cleavage loci before AleaderT, we effectively weaken the interference brought by the 5'UTR and make our part more standardized(Lei Qi et al, 2012). | Also, via adding a csy4 cleavage loci before AleaderT, we effectively weaken the interference brought by the 5'UTR and make our part more standardized(Lei Qi et al, 2012). | ||
| Line 61: | Line 63: | ||
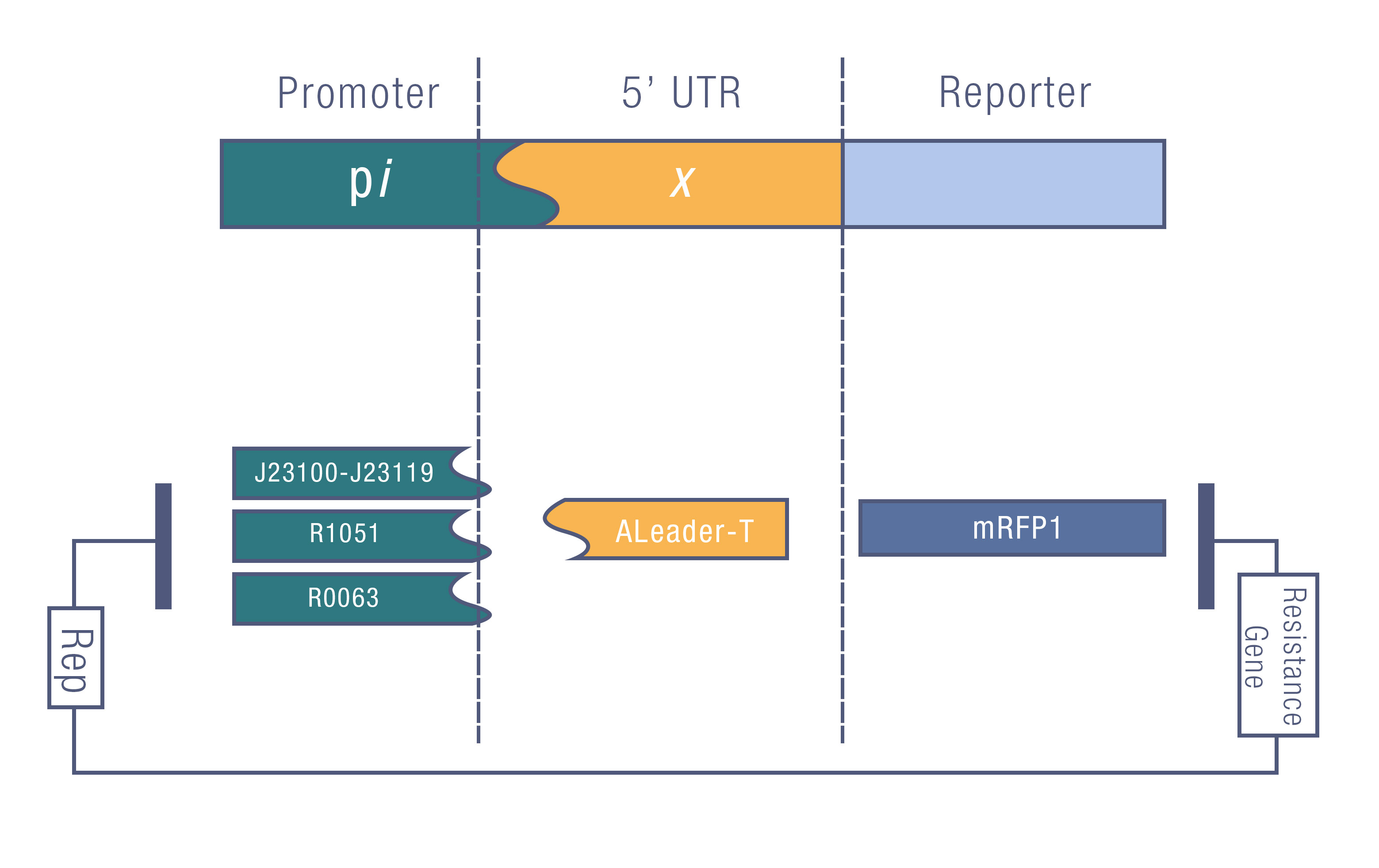
[[File:Aleader5.jpg|800px|thumb|center| '''Figure 6'''. The Promoter Library for Quantitative Estimation of ALeaderT.]] | [[File:Aleader5.jpg|800px|thumb|center| '''Figure 6'''. The Promoter Library for Quantitative Estimation of ALeaderT.]] | ||
| − | We tested the ALeaderT’s function on translation control with the reporter mRFP1 and a promoter library | + | We tested the ALeaderT’s function on translation control with the reporter mRFP1 and a promoter library measured by fluorescent. |
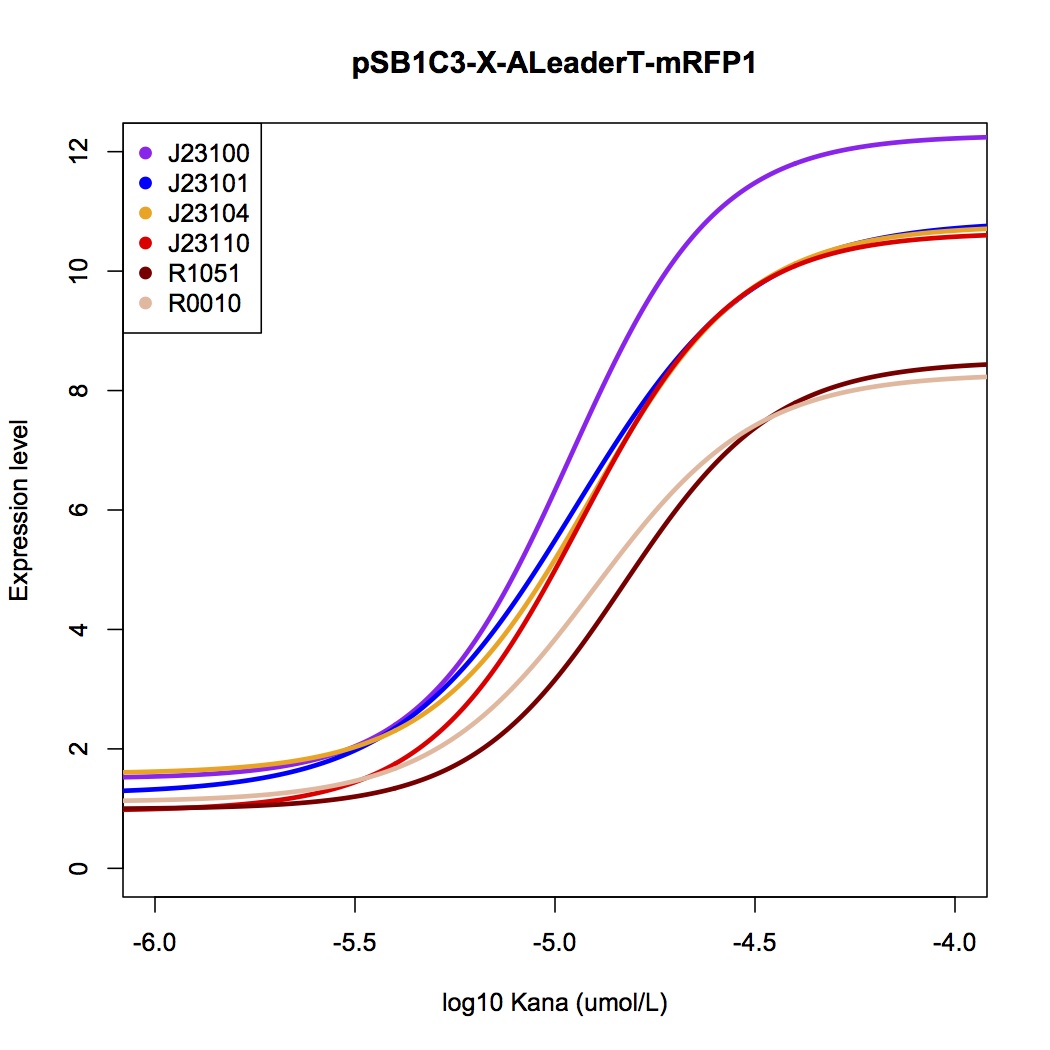
[[File:AleaderT7.jpg|600px|thumb|center| '''Figure 7'''. The dose-response curve of ALeaderT with a promoter library, measured by mRFP fluorescence test.]] | [[File:AleaderT7.jpg|600px|thumb|center| '''Figure 7'''. The dose-response curve of ALeaderT with a promoter library, measured by mRFP fluorescence test.]] | ||
| Line 67: | Line 69: | ||
[[File:AleaderT8.jpg|600px|thumb|center|'''Figure 8'''. ALeaderT has lower basal level than ALeader. The highest levels of them are not significantly different, though ALeaderT have a little higher highest level. Moreover, the robustness of ALeaderT is better than the wild type ALeader. T_Highest: highest level of ALeaderT, T_Basal: basal level of ALeaderT, A_Highest: highest level of ALeader(wt), A_Basal: basal level of ALeader(wt)]] | [[File:AleaderT8.jpg|600px|thumb|center|'''Figure 8'''. ALeaderT has lower basal level than ALeader. The highest levels of them are not significantly different, though ALeaderT have a little higher highest level. Moreover, the robustness of ALeaderT is better than the wild type ALeader. T_Highest: highest level of ALeaderT, T_Basal: basal level of ALeaderT, A_Highest: highest level of ALeader(wt), A_Basal: basal level of ALeader(wt)]] | ||
| + | |||
| + | [[File:AleaderT9.jpg|600px|thumb|center|'''Figure 9'''. The AleaderT shows a higher dynamic range than the wild type ALeader.]] | ||
| + | |||
| + | To test if ALeaderT successfully integrate multi-functions into one riboswitch, we further verified whether the ALeaderT could work as a terminator with a tricistron design (ALeaderT-BCD operon). Similar to the previous iGEM termination efficiency test, we assembled a translational unit of fluorescence protein with ALeaderT to quantify the mRNA transcription, and to eliminate the influence from the SD2 initiated translation, we designed an in frame stop codon. We found that ALeaderT functions as a terminator, inhibited by aminoglycoside antibiotics ('''Figure 10'''), consistent with the prediction. | ||
| + | |||
| + | [[File:AleaderT10.jpg|600px|thumb|center|'''Figure 10'''. The conformation B of ALeaderT is a functional terminator with the maximal termination efficiency (TE) over 80. Aminoglycoside antibiotics repress the termination efficiency. ]] | ||
| + | Since the translation control from ALeaderT is relative to both of the conformation B transcriptional regulation and conformation C SD3 initiation, we can found that the dynamic range of translation caused by ALeaderT is higher than those of the ALeaderT aterminator, and wild type ALeader initiation ('''Figure 11''') | ||
| + | |||
| + | [[File:AleaderT11.jpg|600px|thumb|center|'''Figure 11'''. The functions of members in the ALeader family with promoter J23100, psB1C3 and the Csy4 loci insulator: The ALeaderT induced translation has a higher dynamic range, a higher highest level and a lower basal level, than those of ALeaderT-BCD (TCD test) and ALeader(wt).]] | ||
| + | |||
| + | == Reference == | ||
| + | [1] Jia X, Zhang J, Sun W, et al. Riboswitch Control of Aminoglycoside Antibiotic Resistance[J]. Cell, 2013, 152(1): 68-81. | ||
| + | |||
| + | [2] Hanau‐Berçot B, Podglajen I, Casin I, et al. An intrinsic control element for translational initiation in class 1 integrons[J]. Molecular microbiology, 2002, 44(1): 119-130. | ||
| + | |||
| + | [3] Qi L, Haurwitz R E, Shao W, et al. RNA processing enables predictable programming of gene expression[J]. Nature biotechnology, 2012, 30(10): 1002-1006. | ||
Latest revision as of 01:55, 28 September 2013
ALeaderT (with insulator)
The first artificial triphase riboswitch.
ALeaderT is a novel riboswitch transformed from the leader RNA of the aminoglycoside resistance genes. It has a low minimal level and a high dynamic range, with a low noise in the system and minimal difference under different regulation.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Background
1.Riboswitch Background.
Riboswitches are regulatory RNAs that regulate gene expression by binding small molecules. Recently, a novel riboswitch was reported which is present in the leader RNA of the aminoglycoside resistance genes that encode the aminoglycoside acetyl transferase (AAC) and aminoglycoside adenyl transferase (AAD) enzymes. When the aminoglycosides bind to the leader RNA, a conformational change will be induced, leading to the expression of the aminoglycoside resistance genes (Jia X et al, 2013.). In addition, the leader RNA encompass an integrase site (attl1), which is overlapped by a short peptide encoded by an ORF (ORF11) (Berçot B, et al, 2002). Therefore, Aleader is a complicated riboswitch containing ORF11, cistron, aminoglycoside-binding domain and att1 recombination site. Due to the complicacy of Aleader, the secondary structure of it could not be predicted by most software such as Mfold and NUPACK (Jia X et al, 2013.).
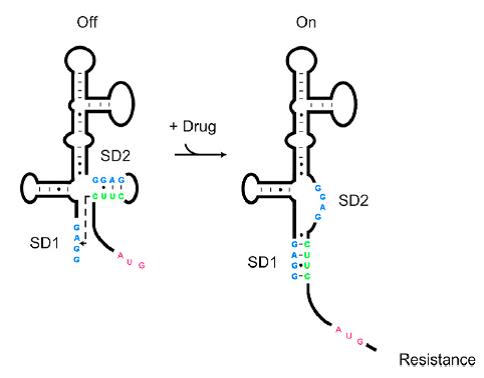
As shown in Figure 1, the 75nt RNA sequence of Aleader contains two SD sequences (ribosome binding sites) and an anti-SD sequence (CUUC) which can complementarily pair with either of the SD sequences. In the absence of aminoglycosides, anti-SD pairs with SD2. The binding of ribosomes to SD1 triggers the translation of a small peptide which stops at the stop codon ahead of SD2, and therefore inhibits the translation of the desired gene after SD2. When aminoglycosides (kanamycin for example) exists, it will induce a structural change of Aleader. The anti-SD sequence base-pairs with SD1, consequently unmask SD2 for ribosomal binding, which results in the translation of the following gene.
2.Csy4 Background.
Csy4 is a member of CRISPR pathway discovered in Pseudomonas aeruginosa. It is a single endoRNase that recognizes and cleaves a 28-nucleotide repetitive sequence and produces stable transcripts with a 5’hydroxylgroup, which can eliminate unwanted interactions between 5’UTRs and translational elements such as RBSs to standardize the expression of the elements. With the insertion of cleavage sites in mRNA, Csy4 can decouple transcript stability and remove structural interactions between multiple genes and associated regulatory elements on the same transcript so that we can get more stable expression and predictable results( Lei Qi et al, 2012).
Take the advantages of Csy4, by physically separating linked genetic elements with cleavage sites, we can achieve assembly of promoters, ribosome binding sites, cis-regulatory elements, or even multigene operons with predictable and standardized functions in bacteria. In other words, it has nice compatibility that enables Csy4 to cooperate with other effective elements to produce new functions ( Lei Qi et al, 2012).
Our design of AleaderT
Many riboswitches have lower dynamic range and higher leakage rate compared with protein regulators. To reduce the basal level and enhance the dynamic range, while having no effect on the protein product, we designed a novel riboswitch called ALeaderT, following the principle of three-phase switch design.
We added a GGGS linker (dock III), a common neutral peptide linker, after the SD2 and ATG. Impressively, by RNA 2D-prediction, we found that dock III will capture antiSD2 sequence to form a stable stem loop, with characteristics of terminator, though neutral as an amino peptide. We suppose that the ALeaderT will have three different steady-state conformations, A “Translation off Termination off”, B “Translation on Termination off”, and C “Translation on Termination on”(Figure 3).
The A conformation is the ligand-free conformation that SD2 and antiSD form the stem loop, resulting in no translation nor termination. The B conformation is the ligand-binding conformation that SD2 is released and SD1 binds the antiSD, leading to the translation. And in C conformation, the SD2 is also released, making it possible to bind the ribosome, but the antiSD and dock III form a stable stem loop to reduce the transcription. Through mathematical modeling, we found that the leakage expression would significantly decrease, while the dynamic range increased several-fold. The working range is usually changed, as well.
1.Dynamic Range(shown with ratio):
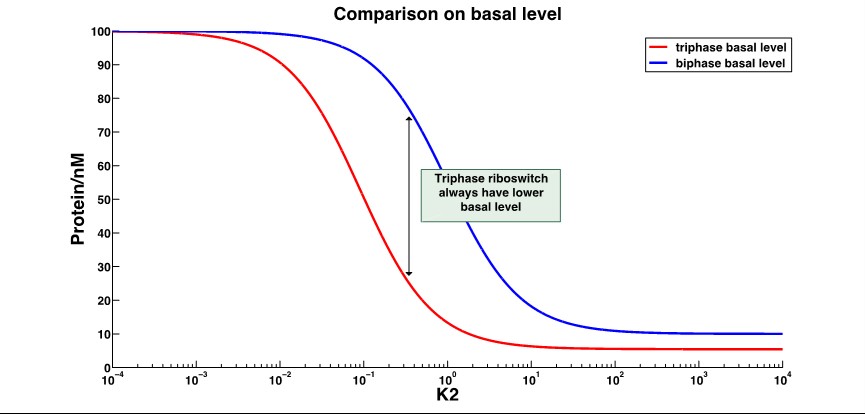
2.Basal level expression(with ON behavior riboswitch):
That shows the leakage expression of triphase riboswitch significantly decrease compared to the biphase riboswitch.
Want to know about our triphase model? Click [http://2013.igem.org/Team:Fudan/triphase_riboswitch.html Here]
Also, via adding a csy4 cleavage loci before AleaderT, we effectively weaken the interference brought by the 5'UTR and make our part more standardized(Lei Qi et al, 2012).
Manipulating and Experimental Data (Quantitative Measurement)
We design a series of experiments to test the function of ALeaderT with the reporter mRFP1 and a promoter library(Figure 6). It showed a very low minimal level and high dynamic range, consistent with the previous prediction. Moreover, it changed the hill function working curve into a square-like curve. Surprisingly, the ALeaderT also showed virtues of marvelous low noise in the system and minimal difference under different regulation. Therefore it will have its talent on the precisely regulation of RNA and protein expression.
We tested the ALeaderT’s function on translation control with the reporter mRFP1 and a promoter library measured by fluorescent.
It showed a very low basal level and high dynamic range, consistent with the modeling prediction(Figure 8,9). Marvelously, the ALeaderT is also much more standardized and robust than the wild type ALeader(Figure 8). In other words, with different promoters, AleaderT’s function varies insignificantly. Therefore it is talented on the precisely regulation of RNA and protein expression.

To test if ALeaderT successfully integrate multi-functions into one riboswitch, we further verified whether the ALeaderT could work as a terminator with a tricistron design (ALeaderT-BCD operon). Similar to the previous iGEM termination efficiency test, we assembled a translational unit of fluorescence protein with ALeaderT to quantify the mRNA transcription, and to eliminate the influence from the SD2 initiated translation, we designed an in frame stop codon. We found that ALeaderT functions as a terminator, inhibited by aminoglycoside antibiotics (Figure 10), consistent with the prediction.
Since the translation control from ALeaderT is relative to both of the conformation B transcriptional regulation and conformation C SD3 initiation, we can found that the dynamic range of translation caused by ALeaderT is higher than those of the ALeaderT aterminator, and wild type ALeader initiation (Figure 11)
Reference
[1] Jia X, Zhang J, Sun W, et al. Riboswitch Control of Aminoglycoside Antibiotic Resistance[J]. Cell, 2013, 152(1): 68-81.
[2] Hanau‐Berçot B, Podglajen I, Casin I, et al. An intrinsic control element for translational initiation in class 1 integrons[J]. Molecular microbiology, 2002, 44(1): 119-130.
[3] Qi L, Haurwitz R E, Shao W, et al. RNA processing enables predictable programming of gene expression[J]. Nature biotechnology, 2012, 30(10): 1002-1006.