Difference between revisions of "Part:BBa K515005"
Jianfei Song (Talk | contribs) |
|||
| (21 intermediate revisions by 7 users not shown) | |||
| Line 4: | Line 4: | ||
a translational unit of superfolder GFP | a translational unit of superfolder GFP | ||
| − | + | ===Characterization for Universal Golden Gate Cloning (Alma College)=== | |
| − | === | + | |
| − | + | We used this BioBrick to characterize a new approach to Golden Gate assembly - one inspired by a paper by [https://www.ncbi.nlm.nih.gov/pubmed/25524097 Bryksin et al]. In this approach, universal PCR primers add BsaI sites and overhang sequences to any BioBrick, thanks to a specialized mismatch priming. This eliminates the need to order separate primers for every BioBrick you want to assembly using Golden Gate. We designed forward primers with overhangs sequences AATT, AAGG, ACTC, AGGT, and reverse primers with overhangs AAGG, ACTC, AGGT, and TGCA. For more information, refer to the diagram to the right or visit the contribution page on our Wiki. | |
| − | + | ||
| − | + | ||
| − | + | Using this (and other) BioBricks and the above primers, we carried out PCR reactions to determine if this approach was feasible. The reactions were generally setup as follows: 0.5uL of purified plasmid DNA for BioBrick BBa_K515005 was mixed with 9.5uL ddH20, 1.25uL of each primer (@ a 10uM concentration), and finally with 12.5uL of a 2X Mix for [https://www.neb.com/products/m0492-q5-high-fidelity-2x-master-mix Q5 DNA Polymerase]. Reaction conditions were setup as suggested by NEB for Q5 polymerase, typically with an annealing temperature between 51 and 55C. In some trials, 9uL of ddH20 and 0.5ul of DMSO were added. | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | A depiction of the primer binding, and a representative gel, is below. | |
| − | + | ||
| − | + | ||
| − | + | [[File:T--Alma--universalprimers2019_2.png|400px]] | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | PCR products from some reaction combinations were selected and sequenced from the middle towards the end - this way, we can confirm if the overhang and BsaI site were added correctly. VF2 and VR primers were used as controls in the event that some primer pairs gave no or incorrect products. | |
| − | + | ||
| + | As you can see in the table below, we had mixed results - some of these PCRs required optimization, and were sensitive to annealing temperature and the speed at which the temperature shifted (evident by comparing the reaction in an old thermocycler versus a new mini8 PCR machine). However, we were able to show that this approach can be viable and does indeed lead to products that should work in a Golden Gate assembly reaction. | ||
| − | + | [[File:T--Alma--universalprimers2019_3.png|400px]] | |
| − | + | Legend: green indicates a successfully confirmed reaction (with the bold cell representing the result corresponding to the representative chromatogram above), light green indicates a sequencing analysis with partial success/confirmation/matching, orange-pink (EtBr color) indicates a PCR reaction that gave the correct product, and purple indicates a reaction that did not work. Empty cells represent untested combinations. *These reactions did not work initially, but were successful at least once (required further optimization). | |
| − | + | ||
| + | == '''Team iBowu-China 2021 Contribution'''== | ||
| + | <div> | ||
| + | <b>Group: iBowu-China 2021 </b> | ||
| + | <br> | ||
| + | <b>Author: Rachel Chen</b> | ||
| + | <br> | ||
| + | <b>Introduction:</b> | ||
| − | <p> | + | <p> |
| − | + | We used this part as a positive comparison for control group in the measurement of green fluorescence. Carried on a pSB1C3 plasmid, this part can effectively produce green fluorescence, together with a constitutive promoter. Our contribution includes | |
| + | # We transformed both DH5a and BL21(DE3) and produced green fluorescence without any addition of induction reagents such as IPTG. | ||
| + | # In both cases, the light intensity is much stronger than the background by about 100 fold. | ||
| − | < | + | <br> |
| − | + | ||
| − | + | ||
| + | <b>Protocol:</b> | ||
| + | The plasmid was obtained from BNDS-China 2021 team. It was then expressed both in E. coli DH5a and E. coli BL21(DE3) with Cm resistance. The bacteria culture was incubated without addition of any iPTG in both cases overnight at 37 degree Celsius. GFP was measured with a microplate reader and OD600 was taken for normalization of the concentration. A control group was also measured for background light intensity where the E. coli BL21(DE3) without this plasmid was cultured. A picture was taken to show the effect of green fluorescence protein expression. | ||
| − | < | + | <br> |
| − | < | + | <b>Results:</b> |
| − | + | <br> | |
| − | + | [[File: T--iBowu-China--2021gfp1.png |800px|thumb|center| Figure 1. Green fluorescence protein expressed by <i>E. coli</i> DH5a and BL21(DE3) under visible light and their measured intensity normalized by their concentration (OD600). The third column was the control group where the E. coli BL21(DE3) without this plasmid was used. ]] | |
| − | < | + | <br> |
| − | + | The results show this part can produce green fluorescence protein highly effectively. In BL21(DE3), the expression is stronger than in DH5a. The intensity of the GFP light in BL21(DE3) is about 2x the intensity in DH5a. In both cases, the light intensity is much stronger than the background by about 100 fold. | |
| − | + | ||
| − | < | + | <br> |
| − | < | + | <b>Summary:</b> |
| + | This part can be used for effective expression of green fluorescence protein. It is an ideal choice for control group, and it can also be engineered to express other sequences. | ||
| + | </p> | ||
| + | <br> | ||
Latest revision as of 11:36, 11 October 2022
superfolder GFP (sfGFP)
a translational unit of superfolder GFP
Characterization for Universal Golden Gate Cloning (Alma College)
We used this BioBrick to characterize a new approach to Golden Gate assembly - one inspired by a paper by Bryksin et al. In this approach, universal PCR primers add BsaI sites and overhang sequences to any BioBrick, thanks to a specialized mismatch priming. This eliminates the need to order separate primers for every BioBrick you want to assembly using Golden Gate. We designed forward primers with overhangs sequences AATT, AAGG, ACTC, AGGT, and reverse primers with overhangs AAGG, ACTC, AGGT, and TGCA. For more information, refer to the diagram to the right or visit the contribution page on our Wiki.
Using this (and other) BioBricks and the above primers, we carried out PCR reactions to determine if this approach was feasible. The reactions were generally setup as follows: 0.5uL of purified plasmid DNA for BioBrick BBa_K515005 was mixed with 9.5uL ddH20, 1.25uL of each primer (@ a 10uM concentration), and finally with 12.5uL of a 2X Mix for Q5 DNA Polymerase. Reaction conditions were setup as suggested by NEB for Q5 polymerase, typically with an annealing temperature between 51 and 55C. In some trials, 9uL of ddH20 and 0.5ul of DMSO were added.
A depiction of the primer binding, and a representative gel, is below.
PCR products from some reaction combinations were selected and sequenced from the middle towards the end - this way, we can confirm if the overhang and BsaI site were added correctly. VF2 and VR primers were used as controls in the event that some primer pairs gave no or incorrect products.
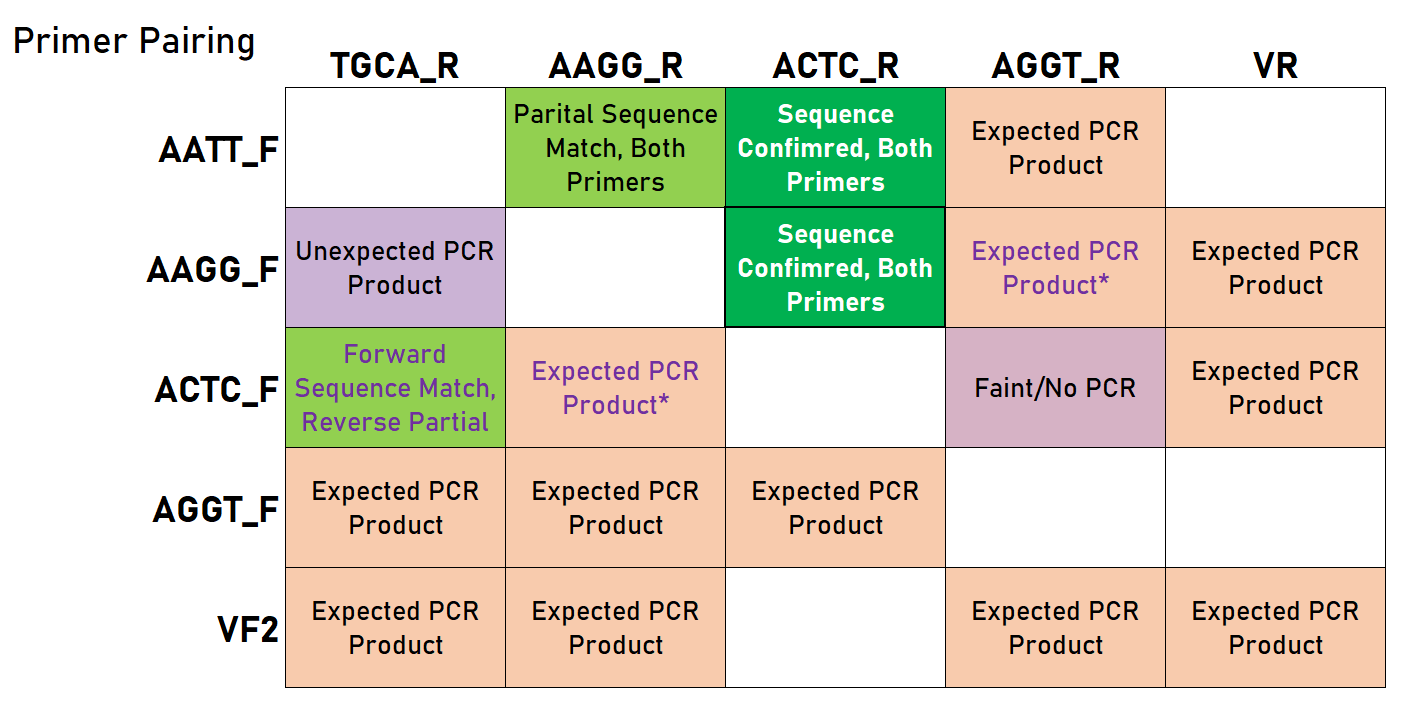
As you can see in the table below, we had mixed results - some of these PCRs required optimization, and were sensitive to annealing temperature and the speed at which the temperature shifted (evident by comparing the reaction in an old thermocycler versus a new mini8 PCR machine). However, we were able to show that this approach can be viable and does indeed lead to products that should work in a Golden Gate assembly reaction.
Legend: green indicates a successfully confirmed reaction (with the bold cell representing the result corresponding to the representative chromatogram above), light green indicates a sequencing analysis with partial success/confirmation/matching, orange-pink (EtBr color) indicates a PCR reaction that gave the correct product, and purple indicates a reaction that did not work. Empty cells represent untested combinations. *These reactions did not work initially, but were successful at least once (required further optimization).
Team iBowu-China 2021 Contribution
Group: iBowu-China 2021
Author: Rachel Chen
Introduction:
We used this part as a positive comparison for control group in the measurement of green fluorescence. Carried on a pSB1C3 plasmid, this part can effectively produce green fluorescence, together with a constitutive promoter. Our contribution includes
- We transformed both DH5a and BL21(DE3) and produced green fluorescence without any addition of induction reagents such as IPTG.
- In both cases, the light intensity is much stronger than the background by about 100 fold.
Protocol: The plasmid was obtained from BNDS-China 2021 team. It was then expressed both in E. coli DH5a and E. coli BL21(DE3) with Cm resistance. The bacteria culture was incubated without addition of any iPTG in both cases overnight at 37 degree Celsius. GFP was measured with a microplate reader and OD600 was taken for normalization of the concentration. A control group was also measured for background light intensity where the E. coli BL21(DE3) without this plasmid was cultured. A picture was taken to show the effect of green fluorescence protein expression.
Results:
The results show this part can produce green fluorescence protein highly effectively. In BL21(DE3), the expression is stronger than in DH5a. The intensity of the GFP light in BL21(DE3) is about 2x the intensity in DH5a. In both cases, the light intensity is much stronger than the background by about 100 fold.
Summary:
This part can be used for effective expression of green fluorescence protein. It is an ideal choice for control group, and it can also be engineered to express other sequences.