Difference between revisions of "Part:BBa I742154"
| Line 47: | Line 47: | ||
<partinfo>BBa_I742154 parameters</partinfo> | <partinfo>BBa_I742154 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| + | |||
| + | |||
| + | <h2>Contribution by Team 2024 Foshan-GreatBay</h2> | ||
| + | |||
| + | <h3>Summary</h3> | ||
| + | |||
| + | To increase the yield of β-carotene in yeast cells, we constructed a new composite part BBa_K5419008 (pXI-2-PaCrtY) with crtY (BBa_I742154) gene fragment. This composited part was used together with other composite parts, BBa_K5419000 (pX-2-PaCrtE), BBa_K5419003 (pX-3-PagCrtB), and BBa_K5419005 (pXII-5-BtCrtI), for the construction of yeast strains with high β-carotene production.</p> | ||
| + | |||
| + | <h3>Construction Design</h3> | ||
| + | <p> | ||
| + | We constructed the plasmid by placing the gene under the regulation of a strong constitutive GAP promoter and a CYC terminator, respectively. Integration sequence was added upstream and downstream of the expression cassette to integrate the target gene into the genome of <i>S. cerevisiae</i> using the CRISPR/Cas9 system (Figure 1). | ||
| + | </p> | ||
| + | |||
| + | <!-- Figure 1 --> | ||
| + | <div style="text-align:center;"> | ||
| + | "https://static.igem.wiki/teams/5419/bba-k5419008/figure-1.jpg" | ||
| + | <div class="caption">Figure 1 Design diagrams of pXI-2-PaCrtY.</div> | ||
| + | </div> | ||
| + | <h3>Experimental Approach</h3> | ||
| + | <h4>Construction of integration plasmids</h4> | ||
| + | <p> | ||
| + | Firstly, we obtained the target gene expression frames (GAP promotor-gene-CYC terminator) by PCR amplification, and agarose gel electrophoresis results showed that we succeeded in obtaining these fragments. Next, we double-digested the target fragment and the vector (containing the <i>S. cerevisiae</i> XI-2 integration site genes) and obtained the plasmid by enzymatic ligation. Finally, we transformed the enzyme-ligation product into <i>E. coli</i> DH5α competent cells, and the colony PCR and sequencing results showed that we successfully constructed the plasmid (Figure 2). | ||
| + | </p> | ||
| + | |||
| + | <!-- Figure 2 --> | ||
| + | <div style="text-align:center;"> | ||
| + | "https://static.igem.wiki/teams/5419/bba-k5419008/figure-2.jpg" | ||
| + | <div class="caption">Figure 2 The construction results of the pXI-2-PaCrtY plasmid.</div> | ||
| + | </div> | ||
Latest revision as of 10:34, 2 October 2024
crtY (lycopene cyclase) coding sequence
Toulouse_INSA-UPS 2020contributed to the characterisation of this part by adding a new documentation learned form literature on the expression and stability of CrtI.
(--antonmykhailiuk 19:10, 08 October 2020 (UTC+2))
Coding sequence for crtY (lycopene cyclase) from Pantoea ananatis (formerly Erwinia uredovora) DSMZ 30080 (ATCC 19321). Accession: D90087. Part of the carotenoid biosynthesis pathway.
Contribution from other teams
Toulouse_INSA-UPS 2020's contribution
Characterisation
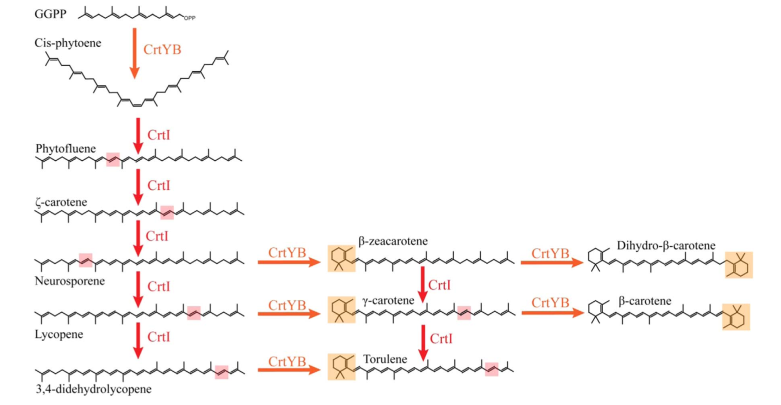
Since the CrtI (phytoene desaturase) is a part of the biosynthesis pathway of carotenoids (fig. 1), it is often co-expressed with the other enzymes of the pathway: such as CrtB or CrtY. Rabeharindranto et al. analyzed the expression of single domain CrtI, CrtY, and CrtB (fig.2). CrtB is unambiguously detected as an intense bands in all strains B, B/I or Y/B/I. On the other hand, CrtI can be observed at the expected size (67kDa) only when coexpressed with CrtY domain (strain Y/B/I). The explanation could be that CrtI protein needs to be co-expressed together with CrtY to have a normal production/stability. A faint band with 50kDa migration pattern could be observed in B/I strain which could further support the idea of low production or stability of CrtI in absence of CrtY.
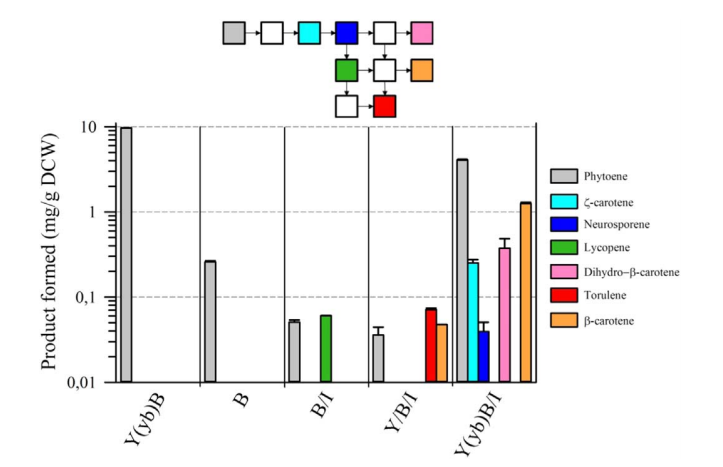
The second point concerns the presence of a natural fusion of CrtY (lycopene cyclase) and CrtB(phytoene synthase). Although, in eucaryotes, CrtB enzyme is predicted to be cytosolic and CrtY enzyme is predicted to be transmembrane [2], surprisingly, there is a natural fusion between CrtY and CrtB which gives CrtYB enzyme[3]. There is a strong opinion on the importance of this natural fusion on the activity of phytoene synthase [4,5]. Rabeharindranto et al. confirmed the impact of CrtY and CrtB splitting on the phytoene production as it has significantly decreased (40 times) in the B strain compared to the Y(yb)B strain (fig. 3).
References for Toulouse_INSA-UPS 2020's contribution
- [1]Rabeharindranto, H., Castaño-Cerezo, S., Lautier, T., Garcia-Alles, L. F., Treitz, C., Tholey, A., & Truan, G. (2019). Enzyme-fusion strategies for redirecting and improving carotenoid synthesis in S. cerevisiae. Metabolic Engineering Communications, 8, e00086. https://doi.org/10.1016/j.mec.2019.e00086
- [2]Schaub, P., Yu, Q., Gemmecker, S., Poussin-Courmontagne, P., Mailliot, J., McEwen, A.G., et al., 2012. On the structure and function of the phytoene desaturase CRTI from Pantoea ananatis, a membrane-peripheral and FAD-dependent oxidase/isomerase. PLoS One 7, e39550. http://dx.doi.org/10.1371/journal.pone.0039550
- [3]Verdoes, J.C., Krubasik, P., Sandmann, G., Van Ooyen, A.J.J., 1999. Isolation and functional characterisation of a novel type of carotenoid biosynthetic gene from Xanthophyllomyces dendrorhous. Mol. Gen. Genet. MGG 262, 453–461.
- [4]Niklitschek, M., Alcaíno, J., Barahona, S., Sepúlveda, D., Lozano, C., Carmona, M., et al., 2008. Genomic organization of the structural genes controlling the astaxanthin biosynthesis pathway of Xanthophyllomyces dendrorhous. Biol. Res. 41, 93–108. http://dx.doi.org/10.4067/S0716-97602008000100011.
- [5]Xie, W., Lv, X., Ye, L., Zhou, P., Yu, H., 2015a. Construction of lycopene-overproducing
Saccharomyces cerevisiae by combining directed evolution and metabolic engineering. Metab. Eng. 30, 69–78. http://dx.doi.org/10.1016/j.ymben.2015.04.009.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Contribution by Team 2024 Foshan-GreatBay
Summary
To increase the yield of β-carotene in yeast cells, we constructed a new composite part BBa_K5419008 (pXI-2-PaCrtY) with crtY (BBa_I742154) gene fragment. This composited part was used together with other composite parts, BBa_K5419000 (pX-2-PaCrtE), BBa_K5419003 (pX-3-PagCrtB), and BBa_K5419005 (pXII-5-BtCrtI), for the construction of yeast strains with high β-carotene production.Construction Design
We constructed the plasmid by placing the gene under the regulation of a strong constitutive GAP promoter and a CYC terminator, respectively. Integration sequence was added upstream and downstream of the expression cassette to integrate the target gene into the genome of S. cerevisiae using the CRISPR/Cas9 system (Figure 1).
""
Experimental Approach
Construction of integration plasmids
Firstly, we obtained the target gene expression frames (GAP promotor-gene-CYC terminator) by PCR amplification, and agarose gel electrophoresis results showed that we succeeded in obtaining these fragments. Next, we double-digested the target fragment and the vector (containing the S. cerevisiae XI-2 integration site genes) and obtained the plasmid by enzymatic ligation. Finally, we transformed the enzyme-ligation product into E. coli DH5α competent cells, and the colony PCR and sequencing results showed that we successfully constructed the plasmid (Figure 2).
""

