Difference between revisions of "Part:BBa R0040:Experience"
(→User Reviews) |
(→2023 University of Warwick iGEM (Team BioMonix)) |
||
| (35 intermediate revisions by 8 users not shown) | |||
| Line 2: | Line 2: | ||
This experience page is provided so that any user may enter their experience using this part.<BR>Please enter | This experience page is provided so that any user may enter their experience using this part.<BR>Please enter | ||
how you used this part and how it worked out. | how you used this part and how it worked out. | ||
| + | |||
| + | ===Applications of BBa_R0040=== | ||
| + | |||
| + | |||
| + | ===2023 University of Warwick iGEM (Team BioMonix)=== | ||
| + | |||
| + | We utilised the pTet sequence as a part of a larger construct, in the pJBEI-6410 backbone. | ||
| + | |||
| + | <html> | ||
| + | <head> | ||
| + | <body> | ||
| + | <div style="display: flex; flex-direction: column; align-items: center; text-align: center;"> | ||
| + | <figure> | ||
| + | <img src="https://static.igem.wiki/teams/4600/wiki/cleanshot-2023-10-12-at-02-56-45.png" alt="Team" style="width: 300px;" class="center"> | ||
| + | <figcaption>Figure 1: Restriction digest of the base pJBEI-6410 and pJBEI-6410* (which includes pTet) using Apal and EcoRI. Used a 1kb plus DNA ladder.</figcaption> | ||
| + | </figure> | ||
| + | |||
| + | 6410 plasmid had shown a band at 1007 bp while 6410* show a band at 1061 bp , showing that there is insertion of pTet sequence into the plasmid, as that is a difference of 54 bases and is around the area that we expected pTet to be inputted into our plasmid. | ||
| + | </body> | ||
| + | </head> | ||
| + | </html> | ||
| + | |||
| + | === 2017 Georgia State iGEM Team === | ||
| + | |||
| + | Georgia State Growth Medium Study | ||
| + | |||
| + | |||
| + | [[File:T--Georgia State--BBa R0040_Georgia_State.png|600px|thumb|center|''Mean Fluorescence Intensity measured by flow cytometry of GFP expressed from 6-hour cultures of BBa R0040 in multiple growth mediums.'' | ||
| + | ]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | '''Group:''' 2016 BIT-China iGEM team | ||
| + | |||
| + | '''Summary:''' Characterizing Part:BBa_R0040 through measuring red fluorescence intensity in E.coli TOP10 and comparing the promoter strength with mutants. The mutation is designed in -35 region of the Ptet promoter. | ||
| + | |||
| + | '''Documentation''' | ||
| + | |||
| + | Purpose: We used Ptet promoter to expression the killer gene in our project. In order to decrease the expression level of toxins, we mutated the -35 region to get Ptet promoters with lower strength. | ||
| + | Results: | ||
| + | Measuring the RFP intensity using enzyme labeled instrument. | ||
| + | [[File:BIT-CHINA-PARTS-MUTATION-1.jpg|600px|thumb|center|Fig1. Compared with the original promoter, the promoter strength of mutants Ptet-1, Ptet-2, Ptet-3, Ptet-25 is sharply decreased.]] | ||
| + | Sequencing the plasmids | ||
| + | [[File:BIT-CHINA-PARTS-MUTATION-2.jpg|600px|thumb|center]] | ||
| + | [[File:BIT-CHINA-PARTS-MUTATION-3.jpg|600px|thumb|center]] | ||
| + | [[File:BIT-CHINA-PARTS-MUTATION-4.jpg|600px|thumb|center]] | ||
| + | [[File:BIT-CHINA-PARTS-MUTATION-5.jpg|600px|thumb|center]] | ||
| + | |||
| + | |||
| + | '''Methods:''' | ||
| + | |||
| + | Random primers are synthesized by GENEWIZ. | ||
| + | |||
| + | Template is Ptet-B0034-RFP-B0015. Designed NNNNNN in -35 region of Ptet promoter. | ||
| + | |||
| + | PCR amplification | ||
| + | |||
| + | Plasmid construction through digestion and ligation | ||
| + | |||
| + | Selection through colony PCR | ||
| + | |||
| + | Incubation for over 12 hours | ||
| + | |||
| + | Measuring the RFP intensity using enzyme labeled instrument | ||
| + | |||
| + | Sequencing the plasmids | ||
| + | |||
| + | |||
| + | |||
| + | [[File:BIT-CHINA-PARTS-MUTATION-6.png|600px|thumb|center]] | ||
===Applications of BBa_R0040=== | ===Applications of BBa_R0040=== | ||
| Line 60: | Line 132: | ||
|width='60%' valign='top'| | |width='60%' valign='top'| | ||
The initial gel worked although the obvious confirmation was not possible due to the very small fragment size (54bp).But later it was confirmed following its usage by forming the part BBa_K182001 through the PCR gel analysis and the sequencing.The plasmid miniprep also worked. | The initial gel worked although the obvious confirmation was not possible due to the very small fragment size (54bp).But later it was confirmed following its usage by forming the part BBa_K182001 through the PCR gel analysis and the sequencing.The plasmid miniprep also worked. | ||
| + | |} | ||
| + | |||
| + | {|width='80%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | <partinfo>BBa_R0040 AddReview 5</partinfo> | ||
| + | <I>Youri</I> | ||
| + | |width='60%' valign='top'| | ||
| + | This promoter was used for high constitutive GFP expression and functioned properly. | ||
| + | |} | ||
| + | |||
| + | <html> | ||
| + | <a name='pTetPV11'></a> | ||
| + | </html> | ||
| + | {|width='80%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | <partinfo>BBa_R0040 AddReview 5</partinfo> | ||
| + | <I>UNIPV-Pavia iGEM 2011</I> | ||
| + | |width='60%' valign='top'| | ||
| + | <html> | ||
| + | |||
| + | <ol> | ||
| + | <ul> | ||
| + | <li> <A HREF="https://parts.igem.org/wiki/index.php/Part: BBa_K516230 "> BBa_K516230 </a> p<sub>Tet</sub>-RBS30-mRFP-TT </li> | ||
| + | <li> <A HREF="https://parts.igem.org/wiki/index.php/Part: BBa_K516231 "> BBa_K516231 </a> p<sub>Tet</sub>-RBS31-mRFP-TT </li> | ||
| + | <li> <A HREF="https://parts.igem.org/wiki/index.php/Part: BBa_K516232 "> BBa_K516232 </a> p<sub>Tet</sub>-RBS32-mRFP-TT | ||
| + | </li> | ||
| + | <li> <A HREF="https://parts.igem.org/wiki/index.php/Part: BBa_I13521 "> BBa_I13521 </a> p<sub>Tet</sub>-RBS34-mRFP-TT | ||
| + | </li></ul> | ||
| + | </ul> | ||
| + | </ol> | ||
| + | |||
| + | <em> | ||
| + | NB: All tests were performed in <a href='http://2011.igem.org/Team:UNIPV-Pavia/Protocols#MG1655Z1'><em>E. coli</em> MGZ1</a> in M9 supplemented medium at 37°C in low copy plasmid <A HREF="https://parts.igem.org/wiki/index.php/Part: pSB4C5">pSB4C5</a>. This strain constitutively expresses TetR, so pTet promoter can be induced with aTc. | ||
| + | </em> | ||
| + | |||
| + | <p>PTet promoter was assembled upstream of different mRFP coding sequences, containing an RBS from the Community collection.</p> | ||
| + | <p>The assembled RBSs are:</p> | ||
| + | <br> | ||
| + | <div align='center'> | ||
| + | <table class='data' border='1'> | ||
| + | <tr><td class="row"><b>BioBrick code</b></td><td><b> Declared efficiency</b></td></tr> | ||
| + | <tr><td class="row">BBa_B0030 </td><td class="row"> 0,6</td></tr> | ||
| + | <tr><td class="row">BBa_B0031 </td><td class="row"> 0,07</td></tr> | ||
| + | <tr><td class="row">BBa_B0032 </td><td class="row"> 0,3</td></tr> | ||
| + | <tr><td class="row">BBa_B0034 </td><td class="row"> 1</td></tr> | ||
| + | </table></div> | ||
| + | <br> | ||
| + | <div align="justify"><p>For an inducible device, the RBS variation has the purpose to stretch the induction curve, thus modulating its PoPs-OUT range.</p> | ||
| + | <p>The complex RBS-promoter acts as a whole regulatory element and determines the amount of translated protein. | ||
| + | RBSs have been reported to have an un-modular behavior, since the translational efficiency is not independent on the coding sequences, but variates as an effect of different mRNA structure stability [Salis et al., Nat Biotec, 2009]. It is not possible to separate the effects of the sole promoter and of the sole RBS on the total amount/activity of gene product (in this case study, mRFP).</p> | ||
| + | <p>For this reason, every combination 'Promoter+RBS' was studied as a different regulatory element. Regulatory elements were characterized using mRFP reporter protein for different RBSs in terms of Synthesis rate per Cell (<b>S<sub>cell</sub></b>) and <b>R.P.U.s</b> (Relative Promoter Units) as explained in <a href='http://2011.igem.org/Team:UNIPV-Pavia/Measurements'>measurements</a> section.</p> | ||
| + | </p> | ||
| + | |||
| + | <p align='justify'> | ||
| + | The Hill function relating the induction to the S<sub>cell</sub> is reported below:<br> | ||
| + | <p align='center'> | ||
| + | S<sub>cell</sub>=α * ( δ + (1-δ)/(1+(K/Induction)<sup>η</sup>) ) | ||
| + | |||
| + | </p> | ||
| + | |||
| + | </p> | ||
| + | |||
| + | <p>Operative parameters of the promoter are derived from the estimated Hill equations obtained by <em>nonlinear least squares</em> fitting (<em>lsqnonlin</em> Matlab routine) of the <a href='http://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#Equations_for_gene_networks'>Hill function</a> expressed in RPUs:</p> | ||
| + | |||
| + | <p><ol><ul><li><b> | ||
| + | RPU<sub>max</sub></b> is equal to the α and represents the maximum promoter activity</p> | ||
| + | </li><p><li><b> | ||
| + | RPU<sub>min</sub></b> is equal to the α * δ represents the minimum promoter activity</p> | ||
| + | </li><p><li> | ||
| + | <b>Switch point</b> is computed as the abscissa of the inflection point of the Hill curve and it is representative of the position of linear region</p> | ||
| + | </li> | ||
| + | <p><li> | ||
| + | <b>Linearity boundaries</b> are determined as the intersection between the tangent line to the inflection point and the upper and lower horizontal boundaries of the Hill curve.</div></li></p> | ||
| + | </ul></ol> | ||
| + | </p> | ||
| + | |||
| + | |||
| + | <div align="justify"> | ||
| + | <p>The protocols for the characterization of p<sub>Tet</sub> promoter are reported in the <a href='http://2011.igem.org/Team:UNIPV-Pavia/Measurements#pTet_protocol'>p<sub>Tet</sub> measurement section</a>.</p> | ||
| + | <p>The data collected from the mRFP measurement systems were processed as described in <a href='http://2011.igem.org/Team:UNIPV-Pavia/Measurements'> data analysis section</a>. The induction curves were obtained by fitting a Hill function as described in <a href='http://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#Ptet_&_Plux'>modelling section</a> and the estimated <a href='http://2011.igem.org/Team:UNIPV-Pavia/Project/Modelling#Table_of_parameters'>parameters</a> for p<sub>Tet</sub> are reported in the pictures and in table below. </p> | ||
| + | <p>This promoter is widely studied and characterized usually using the strong RBS BBa_B0034. Here we have characterized its transcriptional strength as a function of aTc induction (ng/ml) for different RBSs. Three different induction curves were obtained and are reported in figure:</p></div> | ||
| + | |||
| + | <center><a href="https://static.igem.org/mediawiki/2011/a/af/E32_RPU_80.jpg" class="image"> | ||
| + | <img alt="" src="https://static.igem.org/mediawiki/2011/a/af/E32_RPU_80.jpg" class="thumbimage" width="50%"></a> | ||
| + | </center> | ||
| + | |||
| + | |||
| + | <!-- <td width='50%'> | ||
| + | <div style="text-align:justify"><div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/9/93/E33_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/9/93/E33_RPU_80.jpg" class="thumbimage" width="100%"></a></div></div> | ||
| + | </td> --> | ||
| + | |||
| + | <table width='100%'> | ||
| + | <tr><td width='50%'> | ||
| + | <div style="text-align:justify"><div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/9/99/E34_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/9/99/E34_RPU_80.jpg" class="thumbimage" width="100%"></a></div></div> | ||
| + | </td> | ||
| + | <td width='50%'> | ||
| + | <div style="text-align:justify"><div class="thumbinner" width='100%'><a href="https://static.igem.org/mediawiki/2011/e/e4/E35_RPU_80.jpg" class="image"><img alt="" src="https://static.igem.org/mediawiki/2011/e/e4/E35_RPU_80.jpg" class="thumbimage" width="100%"></a></div></div> | ||
| + | </td></tr> | ||
| + | </table> | ||
| + | |||
| + | |||
| + | <p align='justify'> | ||
| + | The estimated parameters of the Hill curves described in the figures are summarized in the table below: | ||
| + | </p> | ||
| + | <br> | ||
| + | <table class='data' width='100%' title='parameter value'> | ||
| + | <tr> | ||
| + | <td class="row"><b>RBS</b></td> | ||
| + | <td class="row"><b>α<sub>p<sub>Tet</sub></sub> [(AUr/min)/cell]</b></td> | ||
| + | <td class="row"><b>δ<sub>p<sub>Tet</sub></sub> [-]</b></td> | ||
| + | <td class="row"><b>η<sub>p<sub>Tet</sub></sub> [-]</b></td> | ||
| + | <td class="row"><b>k<sub>p<sub>Tet</sub></sub> [nM]</b></td> | ||
| + | </tr> | ||
| + | <tr><td class="row">BBa_B0030</td> | ||
| + | <td class="row">230.67 [3.7]</td> | ||
| + | <td class="row">0.028 [91.61]</td> | ||
| + | <td class="row">4.61 [23.73]</td> | ||
| + | <td class="row">8.75 [4.16]</td> | ||
| + | </tr> | ||
| + | <tr><td class="row">BBa_B0031</td> | ||
| + | <td class="row">ND</td> | ||
| + | <td class="row">ND</td> | ||
| + | <td class="row">ND</td> | ||
| + | <td class="row">ND</td> | ||
| + | </tr> | ||
| + | <tr><td class="row">BBa_B0032</td> | ||
| + | <td class="row">55.77 [12]</td> | ||
| + | <td class="row">1.53E-11 [>>100]</td> | ||
| + | <td class="row">4.98 [57.62]</td> | ||
| + | <td class="row">7.26 [14.98]</td> | ||
| + | </tr> | ||
| + | <tr><td class="row">BBa_B0034</td> | ||
| + | <td class="row">120 [5.95]</td> | ||
| + | <td class="row">0.085 [40.6]</td> | ||
| + | <td class="row">24.85 [47.6]</td> | ||
| + | <td class="row">9 [5.43]</td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | Data are provided as average [CV%] | ||
| + | <br><br> | ||
| + | |||
| + | <p align='justify'> | ||
| + | The measurement system p<sub>Tet</sub>-B0031-mRFP-TT couldn't be assayed because its fluorescence output is under the detectability threshold of our measurement instrument. For this reason, the parameters of the corresponding Hill curve couldn't be estimated and are reported as 'Not Determined' ND. | ||
| + | </p> | ||
| + | |||
| + | <div align='justify'> | ||
| + | <p>α parameter (representing the maximum trascriptional rate in the studied range of induction) varies as expected with the RBS variation and also the δ and η parameters are quite different among the RBS variations.</p> | ||
| + | <p>The k<sub>p<sub>Tet</sub></sub> parameter is quite constant among the RBS variations, thus suggesting that in this case the RBS variation doesn't substantially affect the switch point of the Hill curve, even if the amplitude and the maximum slope are not quite maintained (for the η parameter, maybe fitting problems). | ||
| + | |||
| + | </p> | ||
| + | <p align='justify'> | ||
| + | The operative parameters are summarized in the table below: | ||
| + | </p></div> | ||
| + | |||
| + | <table align='center' class='data' width='100%'> | ||
| + | <tr> | ||
| + | <td class='row'><b>RBS</b></td> | ||
| + | <td class='row'><b>RPU<sub>max</sub></b></td> | ||
| + | <td class='row'><b>RPU<sub>min</sub></b></td> | ||
| + | <td class='row'><b>Switch point [ng/ml]</b></td> | ||
| + | <td class='row'><b>Linear boundaries [MIN; MAX] [ng/ml]</b></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='row'>B0030</td><td class='row'>1.53</td><td class='row'>~0</td><td class='row'>7.95</td><td class='row'>[4.66;11.99]</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='row'>B0031</td><td class='row'>ND</td><td class='row'>ND</td><td class='row'>ND</td><td class='row'>ND</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='row'>B0032</td><td class='row'>3.16</td><td class='row'>~0</td><td class='row'>6.7</td><td class='row'>[4.45;10.05]</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='row'>B0034</td><td class='row'>2.73</td><td class='row'>0.23</td><td class='row'>8.96</td><td class='row'>[8.27;9.71]</td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | |||
| + | <p align='justify'> | ||
| + | From these parameters, it is evident that whilst the switch-point is almost maintained for all the RBSs, the linear boundaries are similar for RBS30 and RBS32 but for RBS34 are moved on the right of one order of magnitude. | ||
| + | </p></html> | ||
| + | |} | ||
| + | |||
| + | {|width='80%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | <partinfo>BBa_R0040 AddReview 7</partinfo> | ||
| + | <I>iGEM2016_<br>Hong_Kong_HKUST</I> | ||
| + | |width='60%' valign='top'| | ||
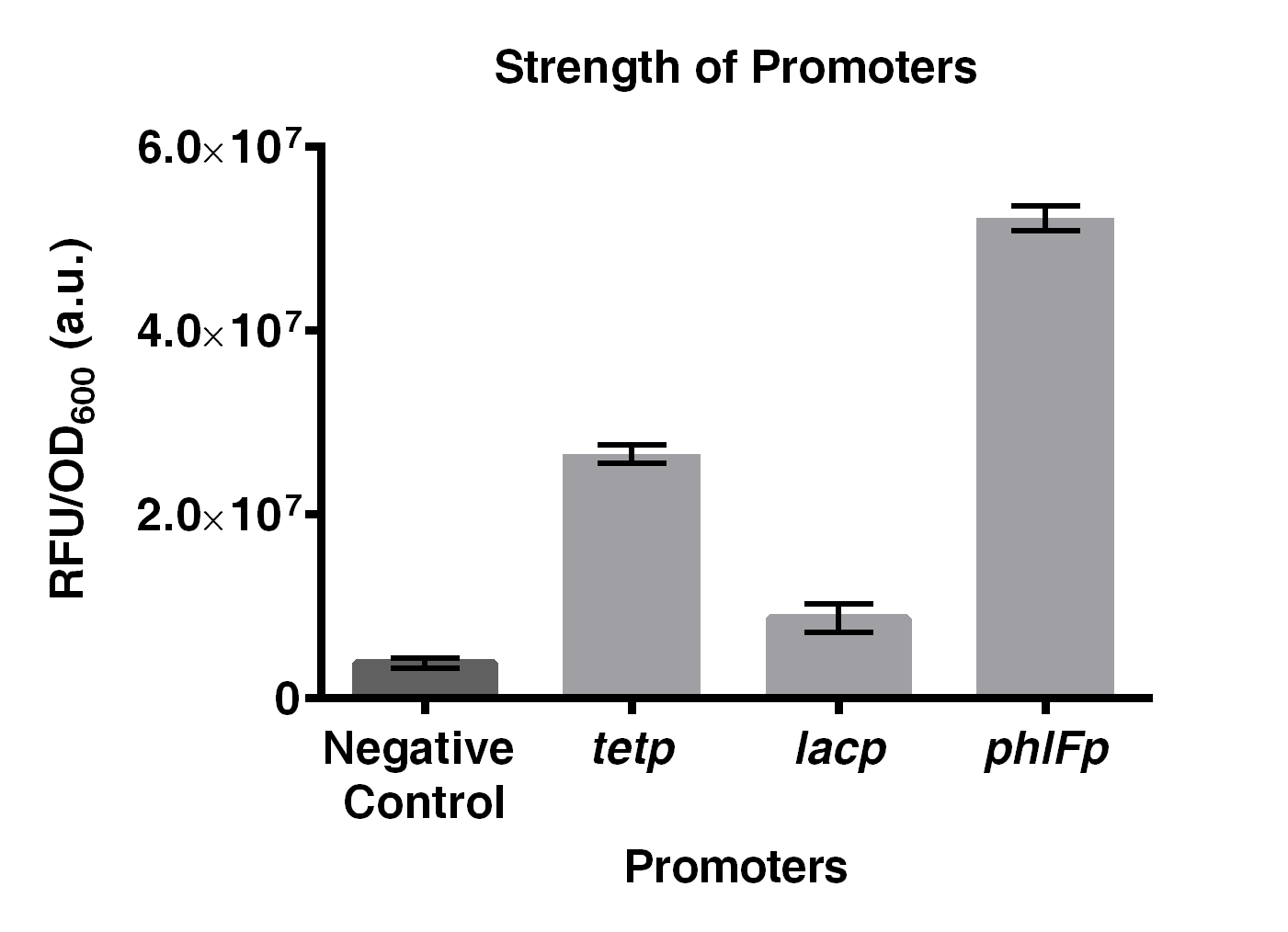
| + | The project of our team this year is to build a Tristable switch. Our design aims to achieve three main characteristics: stable and distinct signal output, and fast state-switching rate. In which stability is the major goal that we would like to achieve. As a results, the strength of the promoters need to be similar. As a result, we compared the promoter strength of <i>phlFp</i>, <i>tetp</i> and <i>lacp</i> by ligating them with a GFP reporter gene, BBa_E0240, in BioBrick RFC10 standard. | ||
| + | <br><br><b>Results</b> | ||
| + | <br><br> | ||
| + | Experiments on comparing the strength of <i>phlFp</i>, <i>tetp</i> and <i>lacp</i>, were performed. Results indicated that <i>phlFp</i> is the strongest among the three, with 1.96 times and 6 times stronger than that of <i>tetp</i> and <i>lacp</i>, respectively. | ||
| + | [[File:IGEM2016 HKUST constructA.png|thumb|500px|center|<b>Fig 2. Comparison on the strength of <i>phlFp</i>, <i>tetp</i> and <i>lacp</i>.</b> Negative control represents BBa_E0240. Characterization was done using <i>E. coli</i> strain JW0336. Cells were first precultured overnight and were subcultured to mid-log phase where GFP emission measurements were made using an EnVision® multilabel reader. This result was obtained by combining 3 characterization data obtained in 3 different days. Error bar present SD from 3 biological replicates.]] | ||
| + | |||
| + | |}; | ||
| + | <!-- End of the user review template --> | ||
<!-- DON'T DELETE --><partinfo>BBa_R0040 EndReviews</partinfo> | <!-- DON'T DELETE --><partinfo>BBa_R0040 EndReviews</partinfo> | ||
| Line 82: | Line 351: | ||
<br style="clear:both;"/> | <br style="clear:both;"/> | ||
| + | |||
| + | |||
| + | {|width='80%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | <I>[http://2013.igem.org/Team:UCSF UCSF 2013]</I> | ||
| + | |width='60%' valign='top'| | ||
| + | <html> | ||
| + | |||
| + | <regulartext1>The UCSF iGEM Team characterized this part as an inducible promoter controlling the expression of GFP. The constitutive tetR repressor was cloned upstream of pTET-GFP, transformed into Top10 cells and the following data was obtained: <br> | ||
| + | |||
| + | <br> | ||
| + | |||
| + | |||
| + | |||
| + | <img align="center" style="margin-bottom:5px; width: 500px; margin-top:20px; padding:2; margin-left:55px;" src="https://static.igem.org/mediawiki/2013/8/8f/PTET_Time_Induction_Curve.png"> <br> | ||
| + | |||
| + | Time curve for pTET promoter induced with different amount of inducers (0, 0.004, 0.01, 0.04, 0.06, 0.11 uM aTc). Cells were grow to mid-log phase and then start induction. OD600 value and GFP fluorescence level of each sample were measured by plate reader over time. GFP fluorescence were corrected for OD600 value. Error bars indicate standard deviation calculated on the basis of technical replicates. | ||
| + | |||
| + | |||
| + | |||
| + | <img align="center" style="margin-bottom:5px; width: 500px; margin-top:50px; padding:2; margin-left:55px;" src="https://static.igem.org/mediawiki/2013/0/04/PTET_Dose_Response_Curve.png"> <br> | ||
| + | |||
| + | Dose-response curve for pTET promoter induced with different amount of inducers (0, 0.004, 0.01, 0.04, 0.06, 0.11 uM aTc). Cells were grow to mid-log phase and then start induction. OD600 value and GFP fluorescence level of each sample were measured by plate reader after saturation. GFP fluorescence were corrected for OD600 value. The red line indicated Hill function fit of the dose-response curve and error bars indicate standard deviation calculated on the basis of technical replicates. | ||
| + | |||
| + | Details on the protocols used can be found on our webpage: http://2013.igem.org/Team:UCSF/Protocols2 | ||
| + | |||
| + | </html> | ||
Latest revision as of 02:27, 12 October 2023
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_R0040
2023 University of Warwick iGEM (Team BioMonix)
We utilised the pTet sequence as a part of a larger construct, in the pJBEI-6410 backbone.

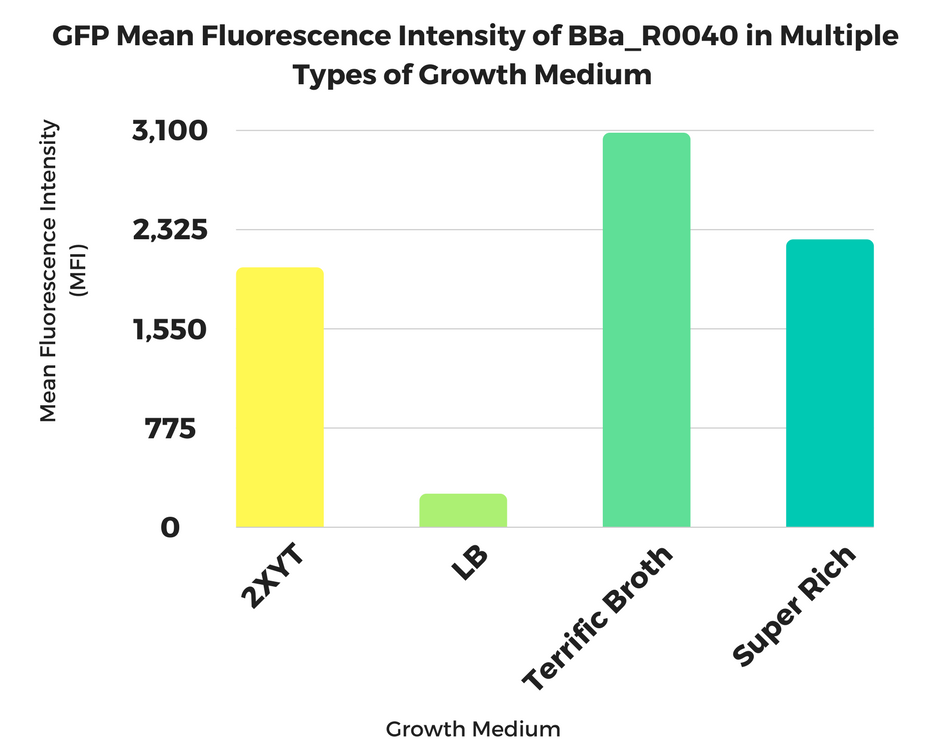
2017 Georgia State iGEM Team
Georgia State Growth Medium Study
Group: 2016 BIT-China iGEM team
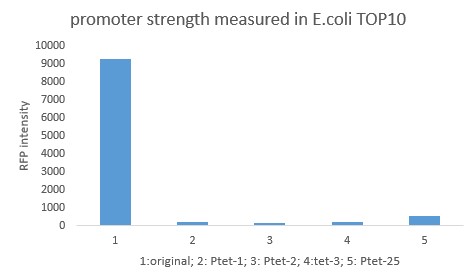
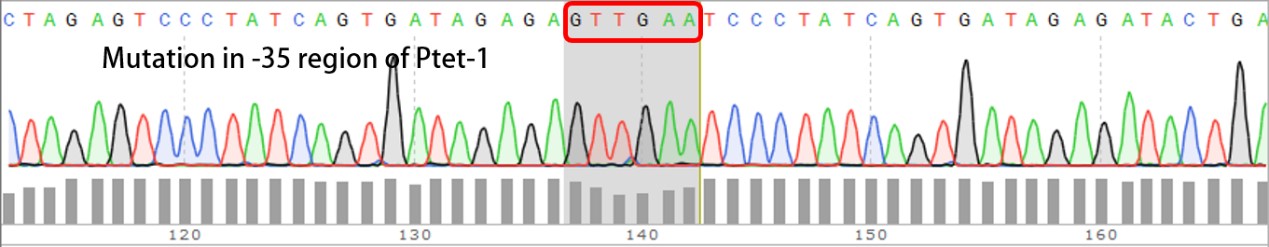
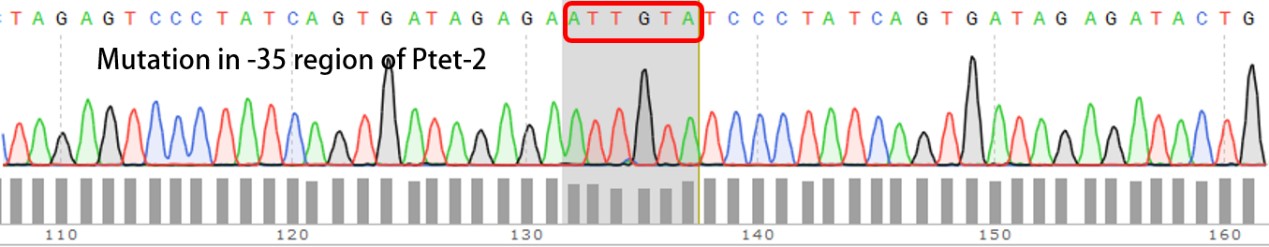
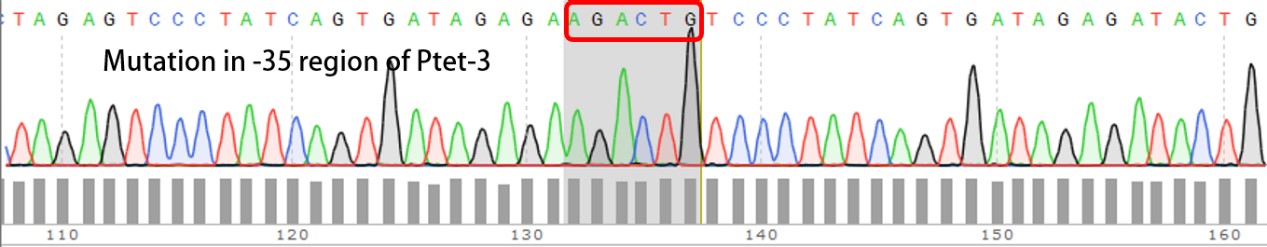
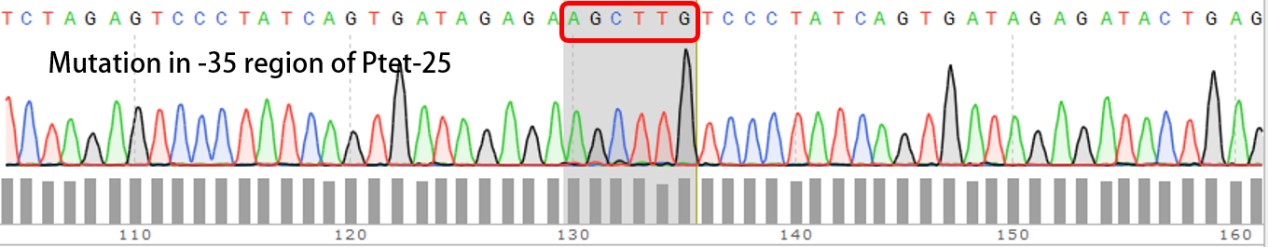
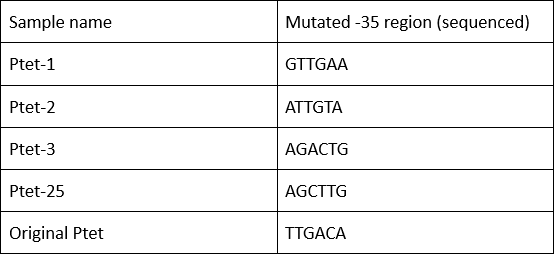
Summary: Characterizing Part:BBa_R0040 through measuring red fluorescence intensity in E.coli TOP10 and comparing the promoter strength with mutants. The mutation is designed in -35 region of the Ptet promoter.
Documentation
Purpose: We used Ptet promoter to expression the killer gene in our project. In order to decrease the expression level of toxins, we mutated the -35 region to get Ptet promoters with lower strength. Results: Measuring the RFP intensity using enzyme labeled instrument.
Sequencing the plasmids
Methods:
Random primers are synthesized by GENEWIZ.
Template is Ptet-B0034-RFP-B0015. Designed NNNNNN in -35 region of Ptet promoter.
PCR amplification
Plasmid construction through digestion and ligation
Selection through colony PCR
Incubation for over 12 hours
Measuring the RFP intensity using enzyme labeled instrument
Sequencing the plasmids
Applications of BBa_R0040
Used this part to create constituitively on GFP and constituitively on OriTf, OriTr. This was one of our most-often used and reliable parts. Have not tried to de-activate it using TetR yet. Worked very well with our parts. [smelissali 6/7/06]
User Reviews
UNIQ666d698b2cfc8300-partinfo-00000001-QINU
|
•••••
Antiquity |
This review comes from the old result system and indicates that this part worked in some test. |
|
•••••
smelissali |
|
••••• |
BBa_R0040, when used to regulate transcription of GFP in BBa_I7100, demonstrated the expected behavior. BBa_R0040 also worked well in the constitutive wintergreen odor generator (BBa_J45120) and banana odor generator (BBa_J45200). |
|
••••
wmholtz |
I have used this promoter both in inducible and constitutive contexts. It generally works great, however it does contain ~20 base pairs of exactly repeated sequence. Several times I've seen recA- strains delete one of these repeated sections. |
|
•••••
Aberdeen_Scotland 2009 |
The initial gel worked although the obvious confirmation was not possible due to the very small fragment size (54bp).But later it was confirmed following its usage by forming the part BBa_K182001 through the PCR gel analysis and the sequencing.The plasmid miniprep also worked. |
|
•••••
Youri |
This promoter was used for high constitutive GFP expression and functioned properly. |
|
•••••
UNIPV-Pavia iGEM 2011 |
PTet promoter was assembled upstream of different mRFP coding sequences, containing an RBS from the Community collection. The assembled RBSs are:
For an inducible device, the RBS variation has the purpose to stretch the induction curve, thus modulating its PoPs-OUT range. The complex RBS-promoter acts as a whole regulatory element and determines the amount of translated protein. RBSs have been reported to have an un-modular behavior, since the translational efficiency is not independent on the coding sequences, but variates as an effect of different mRNA structure stability [Salis et al., Nat Biotec, 2009]. It is not possible to separate the effects of the sole promoter and of the sole RBS on the total amount/activity of gene product (in this case study, mRFP). For this reason, every combination 'Promoter+RBS' was studied as a different regulatory element. Regulatory elements were characterized using mRFP reporter protein for different RBSs in terms of Synthesis rate per Cell (Scell) and R.P.U.s (Relative Promoter Units) as explained in measurements section.
The Hill function relating the induction to the Scell is reported below: Scell=α * ( δ + (1-δ)/(1+(K/Induction)η) ) Operative parameters of the promoter are derived from the estimated Hill equations obtained by nonlinear least squares fitting (lsqnonlin Matlab routine) of the Hill function expressed in RPUs:
The protocols for the characterization of pTet promoter are reported in the pTet measurement section. The data collected from the mRFP measurement systems were processed as described in data analysis section. The induction curves were obtained by fitting a Hill function as described in modelling section and the estimated parameters for pTet are reported in the pictures and in table below. This promoter is widely studied and characterized usually using the strong RBS BBa_B0034. Here we have characterized its transcriptional strength as a function of aTc induction (ng/ml) for different RBSs. Three different induction curves were obtained and are reported in figure: 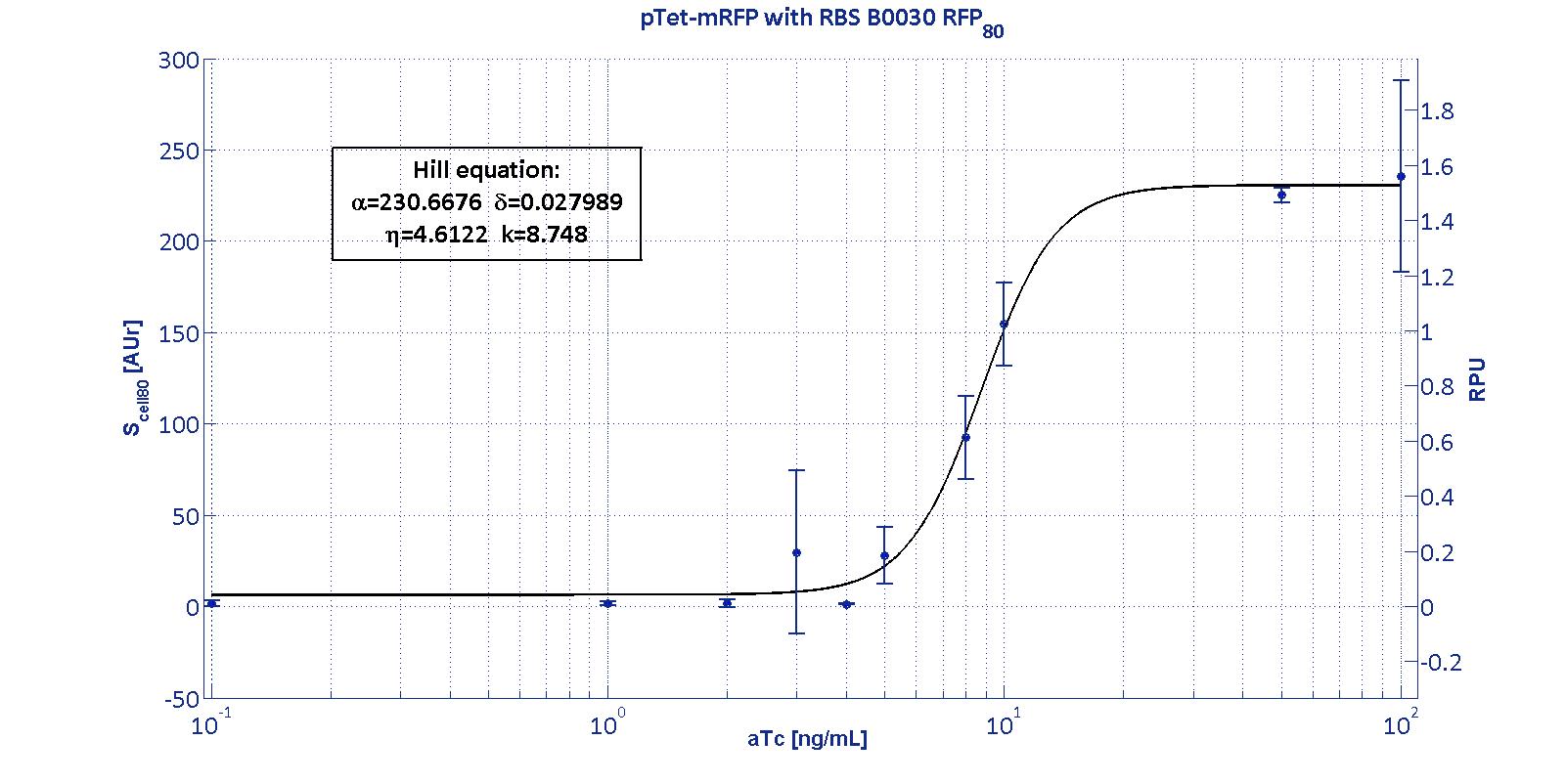
The estimated parameters of the Hill curves described in the figures are summarized in the table below:
The measurement system pTet-B0031-mRFP-TT couldn't be assayed because its fluorescence output is under the detectability threshold of our measurement instrument. For this reason, the parameters of the corresponding Hill curve couldn't be estimated and are reported as 'Not Determined' ND. α parameter (representing the maximum trascriptional rate in the studied range of induction) varies as expected with the RBS variation and also the δ and η parameters are quite different among the RBS variations. The kpTet parameter is quite constant among the RBS variations, thus suggesting that in this case the RBS variation doesn't substantially affect the switch point of the Hill curve, even if the amplitude and the maximum slope are not quite maintained (for the η parameter, maybe fitting problems). The operative parameters are summarized in the table below:
From these parameters, it is evident that whilst the switch-point is almost maintained for all the RBSs, the linear boundaries are similar for RBS30 and RBS32 but for RBS34 are moved on the right of one order of magnitude. |
UNIQ666d698b2cfc8300-partinfo-00000011-QINU
Characterization
Transcriptional control of GFP generator

We successfully designed, constructed and tested transcriptional control devices for constitutive, stationary phase dependent and exponential phase dependent protein production (A-C). To test and verify function of our three transcriptional control devices, we assembled each control device with the GFP protein generator BBa_E0840 and monitored the fluorescence of E. coli cultures with each device over time. For each device, we plot the change in fluorescence per unit time (normalized GFP synthesis rate) versus the cell density (OD600nm) (D). The constitutive transcriptional control device produced a high GFP synthesis rate irrespective of cell density. The stationary phase transcriptional control device produced a low initial GFP synthesis rate which increased with culture cell density. The exponential phase transcriptional control device produced an initially high GFP synthesis rate which dropped off as cell density increased. Data shown are averages of triplicate measurements of cultures grown from three individual colonies of each device. Error bars are the standard deviation of the three individual cultures.
Transcriptional control of banana odor enzyme generator
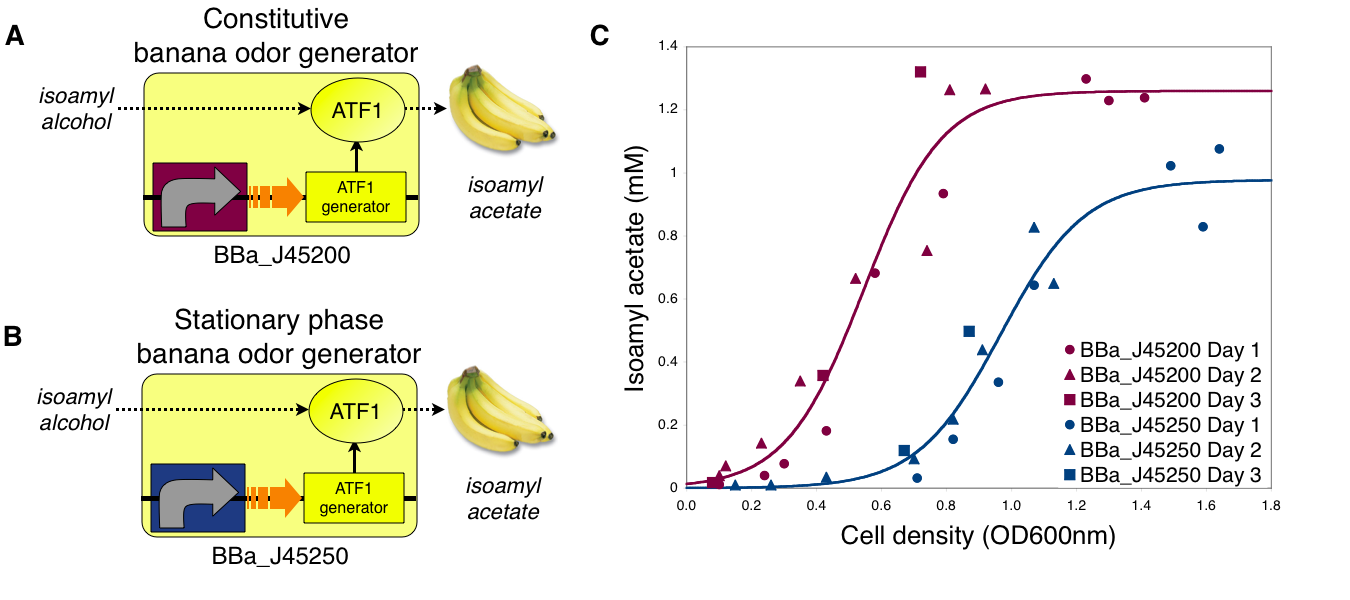
To demonstrate growth phase dependent banana odor production, we compared the behavior of constitutive and stationary phase dependent banana odor generators (A and B, respectively). We measured isoamyl acetate concentrations of cultures of the constitutive and stationary phase banana odor generators at different culture cell densities (OD600nm) (C). As expected, the stationary phase banana odor generator produced very little isoamyl acetate at low cell densities but its isoamyl acetate production increased with cell density. By comparison, the constitutive banana odor generator produced more isoamyl acetate at lower cell densities than the stationary phase banana odor generator. To visually aid comparison of the two odor generators, an empirical fit to the data for each device is shown.
Transcriptional control of wintergreen odor generator
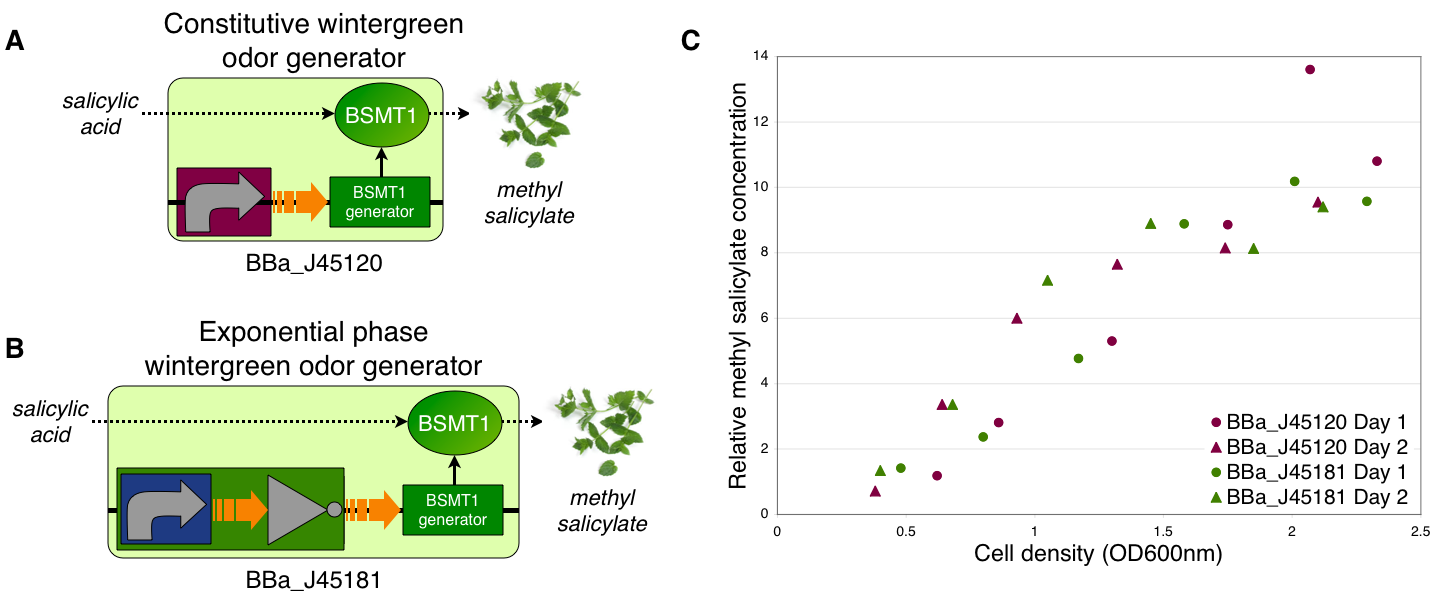
To demonstrate growth phase dependent wintergreen odor production, we compared the behavior of constitutive and exponential phase dependent wintergreen odor generators (A and B, respectively). We measured methyl salicylate concentration (relative to the pentachloronitrobenzene internal standard) of cultures of the constitutive and exponential phase wintergreen odor generators at different culture cell densities (OD600nm) (C). The constitutive and exponential phase wintergreen odor generators produced similar levels of methyl salicylate at all cell densities examined. Thus, the exponential phase wintergreen odor generator does not work as intended.
|
[http://2013.igem.org/Team:UCSF UCSF 2013] |
|