Difference between revisions of "Part:BBa K1675002"
SquarePants (Talk | contribs) |
|||
| Line 34: | Line 34: | ||
<p style="text-align:center;">[[File:T - SCUT China- MR in ybaS.jpeg|500px]]</p> | <p style="text-align:center;">[[File:T - SCUT China- MR in ybaS.jpeg|500px]]</p> | ||
<p style="text-align:center;">Fig.5 Growth of strains MG,MGR and MGR-ybaS under acid stress.</p> | <p style="text-align:center;">Fig.5 Growth of strains MG,MGR and MGR-ybaS under acid stress.</p> | ||
| + | |||
| + | =PuiChing_Macau 2022= | ||
| + | ==Pasr-glsA (BBa_K4340603)pH maintenance functional test== | ||
| + | |||
| + | This is a composite part that can neutralize an acidic environment with ammonia. | ||
| + | |||
| + | <p>These are the results of our Pasr-glsA in pET11a plasmid test (comparing with sfGFP_pET11a). We measured the pH, OD 600, and fluorescence of the cell culture of the transformed E.coli. We use these three indicators to validate the pH neutralizing function and the survival and growth curve of the E.coli transformed with Pasr-glsA_pET11a plasmid.</p> | ||
| + | ===1. pH change test=== | ||
| + | <table> | ||
| + | <tr> | ||
| + | <td> | ||
| + | [[File:Glsa-ph-ph5.png|250px|thumb|center| Figure 1. The pH change in 24 hours of glsA compared to sfGFP (BBa_K4340605) as the control group starting from initial pH 5.]] | ||
| + | </td> | ||
| + | <td> | ||
| + | [[File:Glsa-ph-ph7.png |250px|thumb|center| Figure 2. The pH change in 24 hours of glsA compared to sfGFP (BBa_K4340605) as the control group starting from initial pH 7.]] | ||
| + | </td> | ||
| + | <td> | ||
| + | [[File:glsa-ph-ph9.png |250px|thumb|center| Figure 3. The pH change in 24 hours of glsA compared to sfGFP (BBa_K4340605) as the control group starting from initial pH 9.]] | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p>As the Pasr-glsA_pET11a plasmid is an acid shooting circuit that functions at a low pH environment, the pH change of Figure 2 is very significant in that the pH converges to pH 7 after 24 hours. For figure 3, which is in a pH 7 environment, the pH of Pasr-glsA_pET11a culture drops to pH 6.5 in the first three hours and increases firmly from the third to the ninth hour. Then, starting from the ninth to the 24th hour, the pH increases to around pH 7.4. Overall, the Pasr-glsA_pET11a plasmid does not make a shift change in the pH 7 environment, which is the same result as predicted. In Figure 4, which is in a pH 9 environment where Pasr-glsA_pET11a should not function, the pH first drops to around pH 7.4 in the first nine hours and climbs up to pH 7.8 slowly from the ninth hour to the 24th hour. At the same time, the control group sfGFP (BBa_K4340605)follows the same pattern, which indicates that the Pasr-glsA_pET11a does not function in a high pH environment.</p> | ||
| + | |||
| + | [[File:GlsA plate.png|200px|thumb|center|Photo 1: The glsA transformed E.coli plate.]] | ||
| + | [[File:GlsA plate2.png|400px|thumb|center|Photo 2: The glsA transformed E.coli plate.]] | ||
| + | |||
| + | ===2. OD change test=== | ||
| + | <table> | ||
| + | <tr> | ||
| + | <td> | ||
| + | [[File:glsa-od-ph5.png |250px|thumb|center| Figure 4. The OD changes of Pasr-glsA-pET11a and sfGFP-pET11a transformed E.coli in a pH 5 environment.]] | ||
| + | </td> | ||
| + | <td> | ||
| + | [[File:glsa-od-ph7.png |250px|thumb|center| Figure 5. The OD changes of Pasr-glsA-pET11a and sfGFP-pET11a transformed E.coli in a pH 7 environment.]] | ||
| + | </td> | ||
| + | <td> | ||
| + | [[File:glsa-od-ph9.png |250px|thumb|center| Figure 6. The OD changes of Pasr-glsA-pET11a and sfGFP-pET11a transformed E.coli in a pH 9 environment.]] | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | |||
| + | <p>We also tested the OD value of the E.coli transformed with Pasr-glsA_pET11a and the sfGFP_pET11a (as a control group). In the glsA group, E.coli grows best at pH 7, following pH 5, and finally at pH 9. Since Pasr-glsA constructs can only work to neutralize a low pH environment, the result is as predicted. However, in the sfGFP control group, the highest OD600 rate is in the pH7 environment.</p> | ||
| + | |||
| + | ===3. Fluorescence test=== | ||
| + | <table> | ||
| + | <tr> | ||
| + | <td> | ||
| + | [[File:glsa-fl-ph5.png |250px|thumb|center| Figure 7. The Fluorescence Rate in pH 5 initial environment of sfGFP_pET11a and Pasr-glsA_pET11a.]] | ||
| + | </td> | ||
| + | <td> | ||
| + | [[File:glsa-fl-ph7.png |250px|thumb|center| Figure 8. The Fluorescence Rate in pH 7 initial environment of sfGFP_pET11a and Pasr-glsA_pET11a.]] | ||
| + | </td> | ||
| + | <td> | ||
| + | [[File:glsa-fl-ph9.png |250px|thumb|center| Figure 9. The Fluorescence Rate in pH 9 initial environment of sfGFP_pET11a and Pasr-glsA_pET11a.]] | ||
| + | |||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p>The fluorescence of sfGFP in pH 5 is significantly higher than the fluorescence of Pasr-glsA. It is possibly because the protein size of the sfGFP is smaller than Pasr-glsA, which at the same time produces ammonia to neutralize the environment. In a pH 7 environment, the fluorescence of both sfGFP and Pasr-glsA is relatively similar and reached the same point at the ninth hour. In a pH 9 environment, both fluorescence of sfGFP and Pasr-glsA are low compared to data in pH 5 and 7. This proved that sfGFP and Pasr-glsA, which have the same acid promoter (asr) show low fluorescence in a high-pH environment.</p> | ||
| + | ===4. real-time PCR result=== | ||
| + | [[FIle:glsa-pcr.png |200px|thumb|center| Figure 10. The real-time PCR test result of Pasr-glsA.]] | ||
| + | <p>In our real-time quantitative PCR test, we can see that the fold change of glsA in pH 6.0 is the highest, followed by pH 6.0 and pH 7.0. The glsA mRNA expressed in acid and weak acid environment, which demonstrates that glsA expressed appropriately as predicted.</p> | ||
| + | |||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
Revision as of 13:31, 12 October 2022
glsA, glutaminase1
GlsA encodes an acid-activated glutaminase, which is sufficient for an acid resistance system, and it is able to catalyze a reaction converting glutamate to glutamine and releasing the free gaseous ammonia. The free gaseous ammonia will consume the proton and increase the intracellular pH (Fig.1). The robust glutaminase activity only exists at pH 6.0 or lower. The highest activity is obtained at pH 4.0, followed by pH 5.0 and 6.0. In contrast, GlsA is not activated at pH 7.0 and 8.0. The dependence on pH makes it an effective tool to respond when necessary.
Fig.1 The mechanism of GlsA
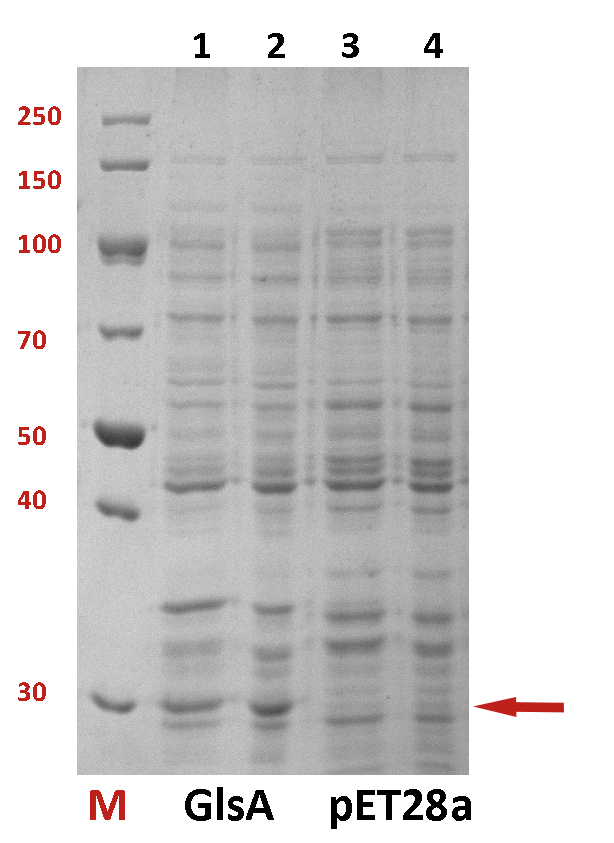
The difference between the testing group and the control group is not evident. We employed a strong promoter T7 to check whether the protein has been expressed. The functional gene GlsA was constructed on plasmid pET28a and the plasmid was transformed into BL21(DE3). 0.5% IPTG was added to induce the expression of the protein. The following is the picture of SDS-PAGE (Fig.2). It shows that our target protein has been expressed successfully.
Fig.2 The SDS-PAGE of GlsA and pET28a
Characterization from SCUT_China 2019:
GlsA is an acid tolerant factor also named ybaS which has a great influence on the acid resistance of the strain. SCUT_China 2019 have expressed this gene and tested their influence on the acid tolerance of E.coli MG1655-T7 RNAP (MGR). T7 RNA polymerase was integrated into the genome of E. coli MG1655(MG) by SCUT_China 2019 to test their VerProS system.
The functional gene ybaS was constructed on plasmid pET30a(+) and the plasmid was transformed into MGR. IPTG(0.2 mM) was added to induce the expression of the protein. Inoculated the MGR with pET30a(+)-ybaS in 10ML LB medium,37 ℃,250rpm for 12 hours,and then 1:100 transferred it to 12.5ml medium with IPTG (0.2 mM) for 18 hours. The following is the picture of SDS-PAGE (Fig.3) which shows that target protein has been expressed successfully.
Fig.3 The SDS-PAGE of gadB and ybaS
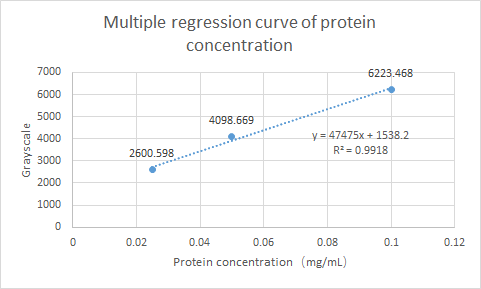
What’s more, SCUT_China 2019 have tested the Protein expression of ybaS. Using ImageJ for gray scale comparison, the multiple regression curve was drawn with the BSA of 0.0125m to 0.1m as the reference, as follow figure:
Fig.4 Multiple regression curve of protein concentration
Finally, the expression of ybaS was calculated as 0.0665 mg/ml.
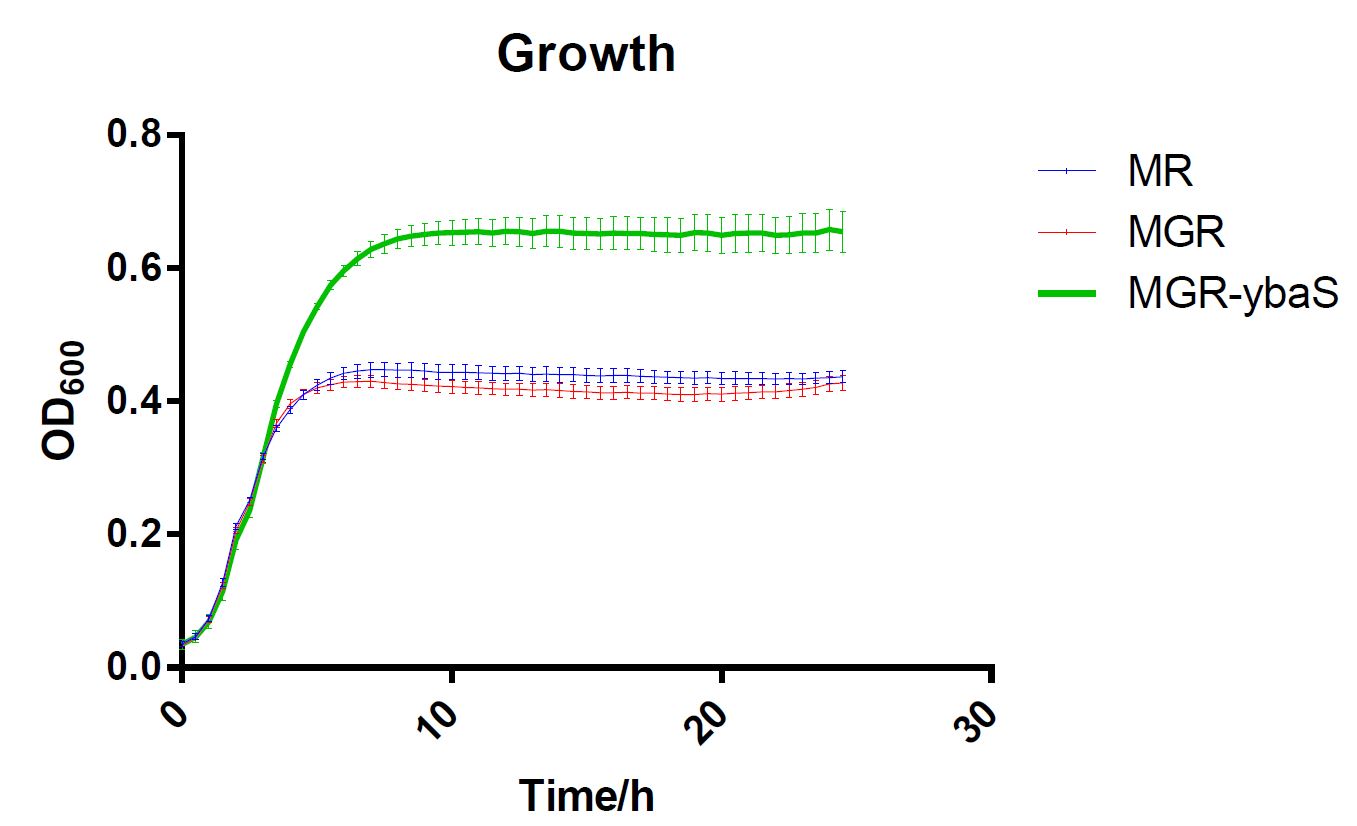
The last, SCUT_China 2019 have tested its influence on the acid tolerance of MGR. MG, MGR and MGR expressing ybaS were grown overnight (about 16 h) in LBG medium of pH 7.0 at 37 °C. The cultures were then diluted to initial OD600 0.05 in 300 μL LBG medium of pH 7.0, LBG medium acidified by HCl or succinic acid to pH 4.5. Then the cultures were incubated at 37 °C in 100-well Honeycomb microplates using an automated turbidimeter (Bioscreen C, Oy Growth Curves Ab Ltd., Helsinki, Finland) for online monitoring of OD600 for 24 h. A growth assay under moderate acid stress were performed to investigate the effect of overexpression ybaS or not on acid tolerance. Under moderate acid stress, the final OD600 value of strain MGR-ybaS(the strain overexpressing ybaS) was 53% higher than that of the wild type strain (MG) (Fig. 5).
Fig.5 Growth of strains MG,MGR and MGR-ybaS under acid stress.
PuiChing_Macau 2022
Pasr-glsA (BBa_K4340603)pH maintenance functional test
This is a composite part that can neutralize an acidic environment with ammonia.
These are the results of our Pasr-glsA in pET11a plasmid test (comparing with sfGFP_pET11a). We measured the pH, OD 600, and fluorescence of the cell culture of the transformed E.coli. We use these three indicators to validate the pH neutralizing function and the survival and growth curve of the E.coli transformed with Pasr-glsA_pET11a plasmid.
1. pH change test
As the Pasr-glsA_pET11a plasmid is an acid shooting circuit that functions at a low pH environment, the pH change of Figure 2 is very significant in that the pH converges to pH 7 after 24 hours. For figure 3, which is in a pH 7 environment, the pH of Pasr-glsA_pET11a culture drops to pH 6.5 in the first three hours and increases firmly from the third to the ninth hour. Then, starting from the ninth to the 24th hour, the pH increases to around pH 7.4. Overall, the Pasr-glsA_pET11a plasmid does not make a shift change in the pH 7 environment, which is the same result as predicted. In Figure 4, which is in a pH 9 environment where Pasr-glsA_pET11a should not function, the pH first drops to around pH 7.4 in the first nine hours and climbs up to pH 7.8 slowly from the ninth hour to the 24th hour. At the same time, the control group sfGFP (BBa_K4340605)follows the same pattern, which indicates that the Pasr-glsA_pET11a does not function in a high pH environment.
2. OD change test
We also tested the OD value of the E.coli transformed with Pasr-glsA_pET11a and the sfGFP_pET11a (as a control group). In the glsA group, E.coli grows best at pH 7, following pH 5, and finally at pH 9. Since Pasr-glsA constructs can only work to neutralize a low pH environment, the result is as predicted. However, in the sfGFP control group, the highest OD600 rate is in the pH7 environment.
3. Fluorescence test
The fluorescence of sfGFP in pH 5 is significantly higher than the fluorescence of Pasr-glsA. It is possibly because the protein size of the sfGFP is smaller than Pasr-glsA, which at the same time produces ammonia to neutralize the environment. In a pH 7 environment, the fluorescence of both sfGFP and Pasr-glsA is relatively similar and reached the same point at the ninth hour. In a pH 9 environment, both fluorescence of sfGFP and Pasr-glsA are low compared to data in pH 5 and 7. This proved that sfGFP and Pasr-glsA, which have the same acid promoter (asr) show low fluorescence in a high-pH environment.
4. real-time PCR result
In our real-time quantitative PCR test, we can see that the fold change of glsA in pH 6.0 is the highest, followed by pH 6.0 and pH 7.0. The glsA mRNA expressed in acid and weak acid environment, which demonstrates that glsA expressed appropriately as predicted.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]