Difference between revisions of "Part:BBa K302033"
(→Improvement from XJTU-China 2022) |
(→Profile) |
||
| Line 82: | Line 82: | ||
=Contribution & Improvement from XJTU-China 2022= | =Contribution & Improvement from XJTU-China 2022= | ||
==Profile== | ==Profile== | ||
| + | ===Name=== | ||
| + | Lysis-phi X174 | ||
| + | |||
===Base Pairs=== | ===Base Pairs=== | ||
273 | 273 | ||
Revision as of 07:08, 12 October 2022
mazF
Encodes a stable non-specific ribonuclease in Bacillus Subtilis. It is used in conjungtion with mazE in Bacillus Subtilis to provide a toxin-antitoxin kill switch in various stressful conditions. When expression of both genes is turned off (as both are under contol of same promoter) mazE will be degraded faster than mazF. There is then no inhibiton of mazF, killing the cell.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Characterized by BNU-China 2019
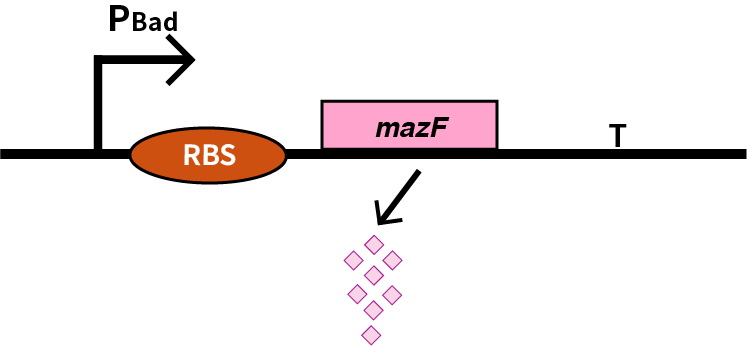
We characterize mazF (BBa_K302033) by an induced suicide system, in which the downstream mazF (BBa_K302033) gene encoding toxin MazF is put under the control of L-arabinose induced promoter pBAD (BBa_K206000). MazF is an endoribonuclease that cleaves RNAs at ACA sites and causes the death of microbe as a reporter [1]. As a result, we can characterize mazF in a cell density-dependent manner in Escherichia coli K-12.
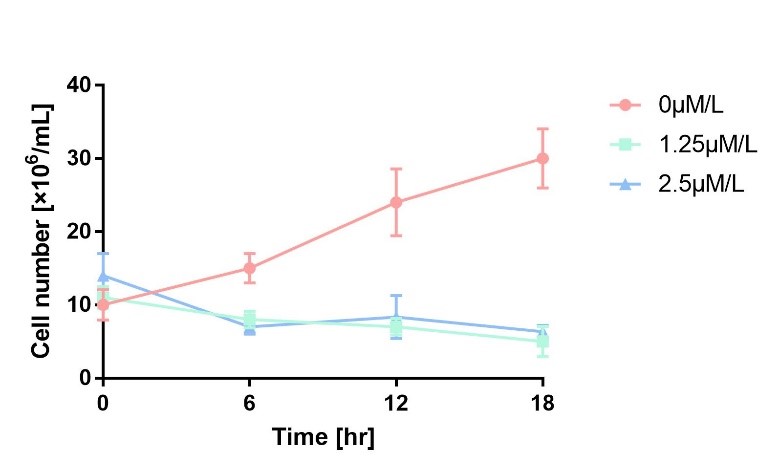
In order to characterize the toxicity of protein encoded by mazF, we take engineered microbe without induction as control group. As is shown in Fig.1, the cell number of experimental groups show a significant decrease upon induction, which indicates the protein encoded by mazF is lethal to E. coli.
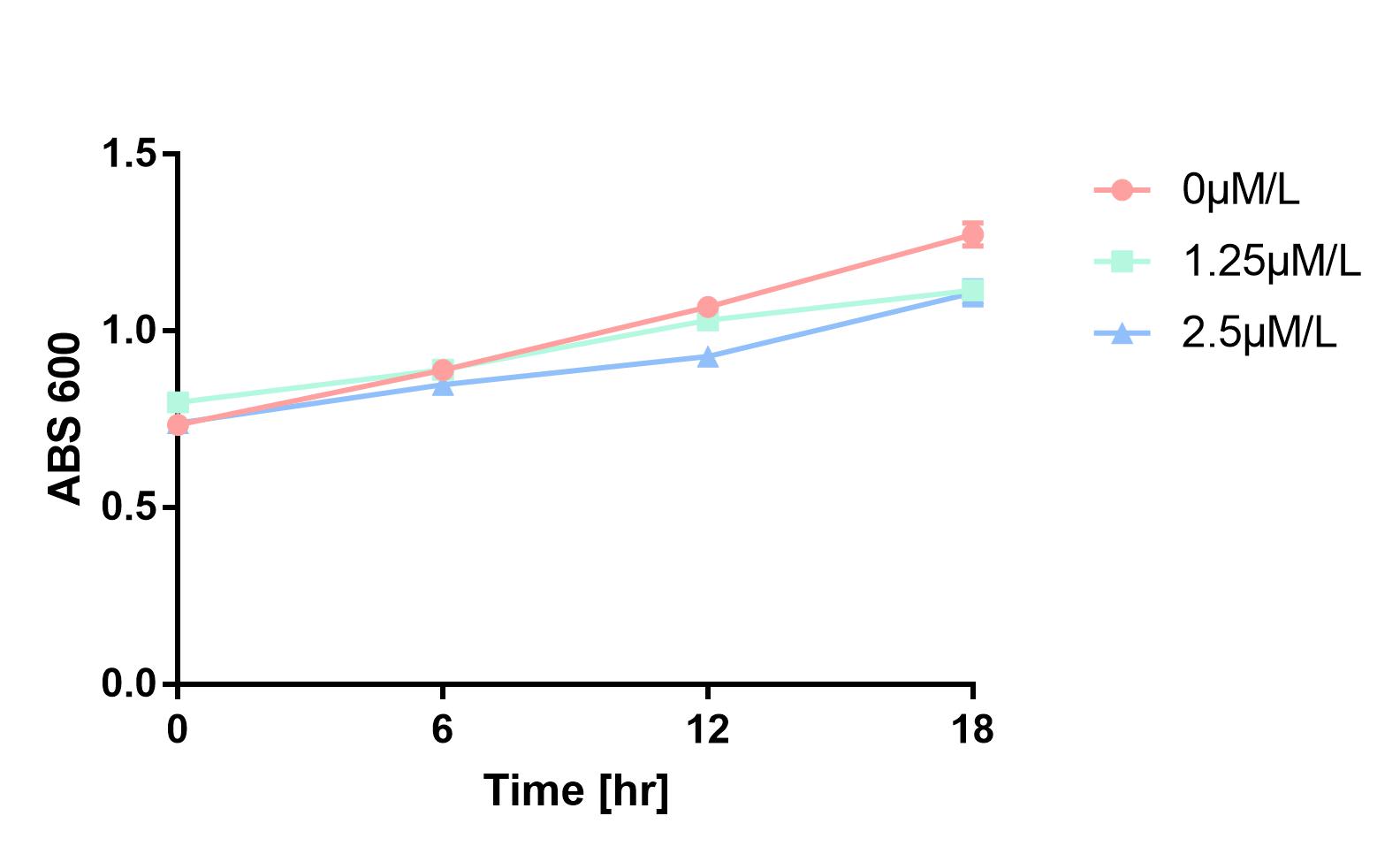
Furthermore, in order to prove that MazF kills the cell without lysing it, we measure OD600 of each sample to give an overall number of intact bacteria, dead and alive. As is shown in Fig.2, there is little difference between control and experimental groups, although it is validated numbers of alive cells differ. Hence, we reach a conclusion that protein encoded by mazF mediates suicide of cells without causing lysis of bacterial cells.
Experimental approach
1. Transform the plasmids into E. coli DH5α competent cells.
2. The engineered bacteria are cultured in 200mL LB-ampicillin (50 ng/µl) medium overnight at 37℃, 200rpm;
3. Equally divide the culture into 90 centrifuge tubes, which is 1mL respectively. Centrifuge them at 4000rpm for 5 minutes. Discard the liquid.
4. Resuspend 30 tubes of collected bacteria with LB-ampicillin (50 ng/µl) containing 1.25μM/L and 2.5μM/L L-arabinose respectively as experimental groups. Resuspend 30 tubes of bacteria with pure LB-ampicillin (50 ng/µl) medium.
5. Collect 3 tubes of all groups every 6 hours, dilute all of the samples to 10^7 times and then spread them on solid LB-ampicillin (50 ng/µl) medium separately. At the same time, refresh the medium to maintain the concentration of L-arabinose.
6. Count the number of colonies in 5 cm^2 per plate after cultured for 24 hours at 37℃
7. Three repicas are tested in each group.
Reference
[1] Diana Széliová, Ján Krahulec, Martin Šafránek, et al. Modulation of heterologous expression from PBAD promoter in Escherichia coli production strains[J]. Journal of Biotechnology, 2016, 236:1-9.
Characterized by XJTU-China 2020
Characterization
This year, our team designed a suicide switch mediated by arabinose operon for biosafety. We use MazF to kill the bacteria. We have supplemented the relevant information of this element in the iGEM Parts and added test and characterization to it.
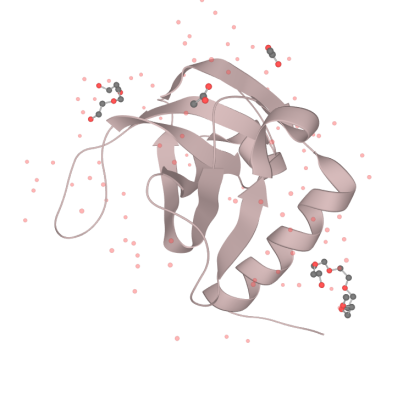
Our MazF comes from the genome of Bacillus Subtilis. By regulating the expression of MazF, we inhibit the life of bacteria when needed to prevent the possible strains and genetic leakage. MazF is a kind of mRNA interferas. Specifically, it specifically cleaves five-base U^ACAU sequence (^indicating the cleavage site) on RNA[1]. MazF achieves its catalytic function by forming a special MazF-RNA complex. One molecule of 9-mer RNA is bound to one MazF dimer, and two subunits of MazF form the dimer related by local 2-fold symmetry.The RNA in an extended alignment is bound along the RNA binding interface between subunits of the MazF dimer, covering part of this dimeric interface, and MazF interacts extensively with the pentad target sequence present in the RNA chain by interactions with bases[2]. Therefore, the backbone phosphate moieties project outward and away from the protein surface. In this way, scissile phosphate will be attacked by other side chain groups, which causes breaking of P-O bond.
What’s more, it is an important part of the Bacillus Subtilis toxin-antitoxin system. Which is essential for the programmed cell death of this developmental bacterium. In normally growing cells, MazF forms a stable complex with its cognate antitoxin, MazE, however, under stress conditions, unstable MazE is preferentially degraded to release free MazF in the cells, which then cleaves cellular mRNAs to inhibit protein synthesis, leading to growth arrest. The protein-protein interaction interface between MazE and MazF (2,843 Å2) is larger than the MazF-RNA interaction interface (2,153 Å2), suggesting that MazF is likely to have a higher affinity for MazE over its RNA substrate. In the B.S MazE-MazF complex structure, MazE binds to MazF along the dimer interface, and it even occupies part of the putative modeled RNA binding site on the second subunit of MazF[3]. Thus, in the presence of MazE, MazF cannot bind to or cleave substrate RNA.
Biochemical properties of MazF
divalent cation[4].
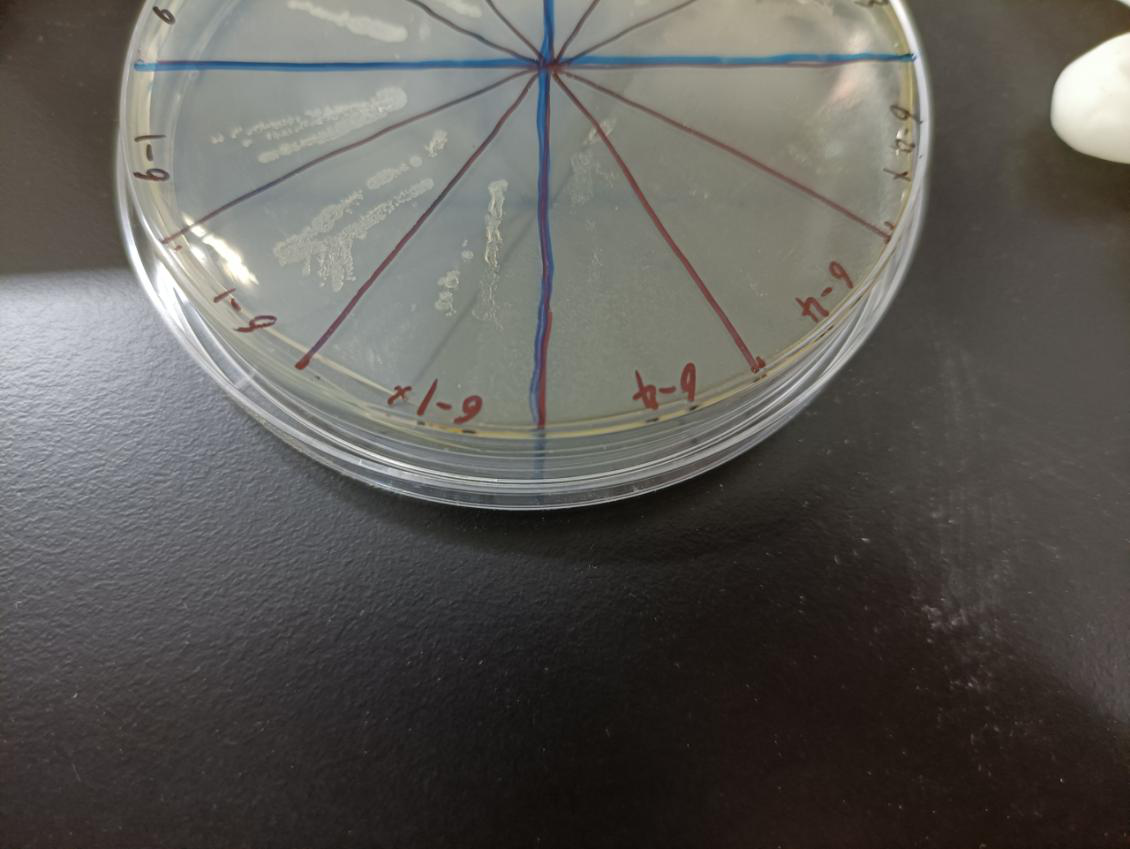
In the figure, the bacteria in this plate contained plasmids of pBAD-MazF. Three of 6-1 groups were the control group without arabinose induction, and three of 6-4 groups were the experimental group with 1.5 M/L arabinose induction. Where, the dilution factor of the group with * is 10^4, and the dilution factor of the group without * is 10^6.
Result: The bacteria without arabinose induction grow normally, while with arabinose induction, the bacteria suicide.
Reference
[1]Park, Jung-Ho, Yamaguchi, Yoshihiro and Inouye, Masayori(2011), Bacillus subtilis MazF‐bs (EndoA) is a UACAU‐specific mRNA interferase, FEBS Letters, 585, doi: 10.1016/j.febslet.2011.07.008 [2]Dhirendra K. Simanshu, Yoshihiro Yamaguchi, Jung-Ho Park, et al. Structural Basis of mRNA Recognition and Cleavage by Toxin MazF and Its Regulation by Antitoxin MazE in Bacillus subtilis. 2013, 52(3):447-458 [3]Stein T. Bacillus subtilis antibiotics: structures, syntheses and specific functions. Mol. Microbiol. 2005; 56:845–857. [PubMed: 15853875] [4] Pellegrini O, Mathy N, Gogos A, Shapiro L, Condon C. The Bacillus subtilis ydcDE operon encodes an endoribonuclease of the MazF/PemK family and its inhibitor. Mol Microbiol. 2005 Jun;56(5):1139-48. doi: 10.1111/j.1365-2958.2005.04606.x. PMID: 15882409.
Contribution & Improvement from XJTU-China 2022
Profile
Name
Lysis-phi X174
Base Pairs
273
Design Notes
The gene was optimized by E. coli codon
Source
Shigella flexneri 2a str. 301 (strain: 301, serotype: 2a)
Usage&Biology
Source and Principle
Biosafety is an important consideration when designing engineered bacteria. From the beginning, we designed the bacteria on the premise that it would work in the field soil, so we first needed to consider whether our product could be easily controlled for the time of its operation and whether there were potential risks to soil structure, crop growth, and the balance of soil microbiota. So we designed a "suicide system" at the genetic level to ensure that our engineered bacteria would not pose a potential biosecurity risk to the ecological environment.
The suicidal behavior of bacteria is a common phenomenon in nature, which is a programmed death mechanism of prokaryotes. quorum sensing (QS) is a form of communication between bacterial cells. Cells synthesize and secrete signal molecules. When the concentration of signal molecules in the environment reaches a certain threshold, a series of genes are activated, and the bacterial population synchronously realizes certain functional and behavioral changes. A quorum-sensing suicide gene circuit has been constructed, and the systematic study and precise regulation of this gene circuit are of great significance both in theory and application [1].
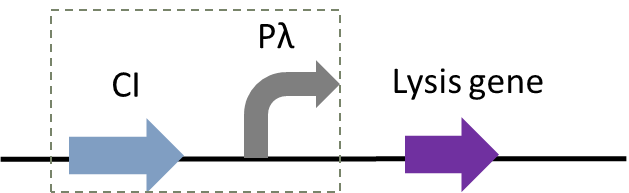
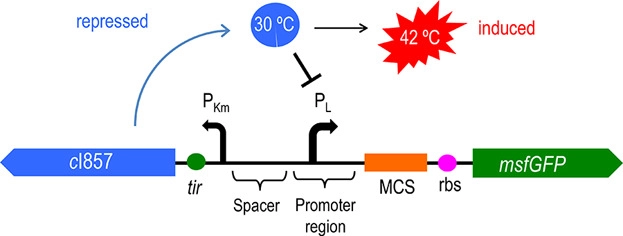
In addition to population-responsive suicide mechanisms, suicide systems with other regulatory modes can also be designed through synthetic biology. Here, we designed a temperature-responsive cleavage system to achieve temperature-controlled cleavage, that is, cleavage of thermoregulated lysis genes (Gene ID: IF654_RS00240) (Figure 1).
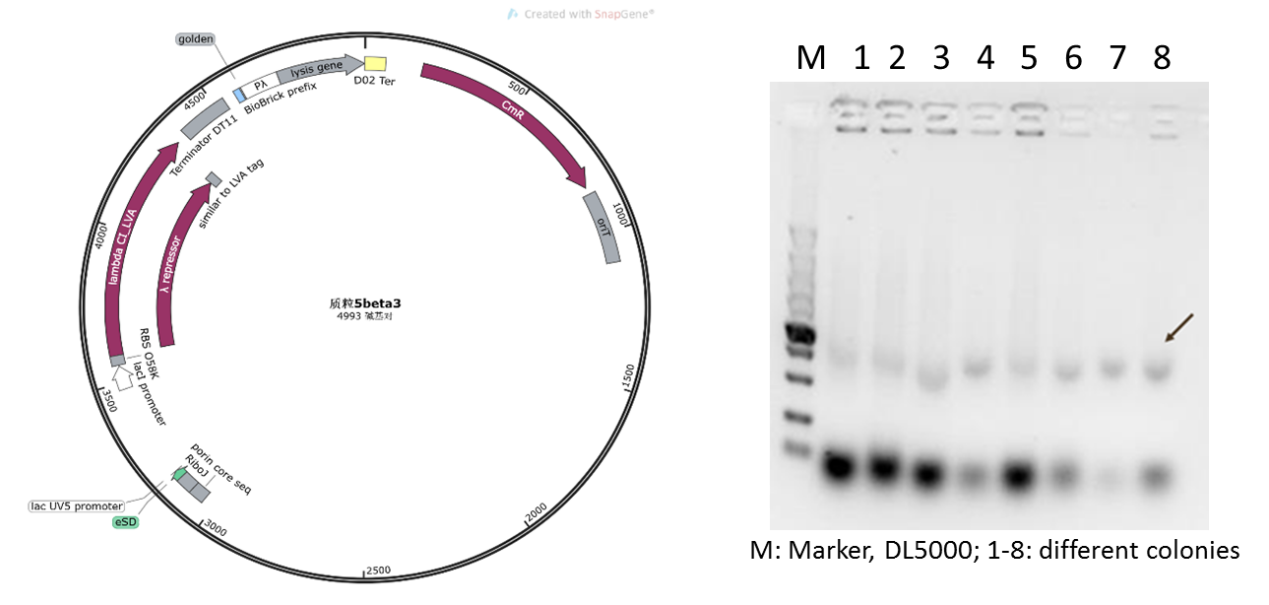
Figure 1: Circuit diagram of plasmid 5: Where CI is the C1857 suppression subsystem, Pλ is the promoter, and the temperature control system is in the dashed box.
Figure 2 shows the principle of the temperature control system [4]. When bacteria are at a low temperature, the c1857 gene expression protein binds to the Pλ promoter, making downstream genes unable to be translated. At 42℃, the protein will be cleaved, leading to the expression of downstream genes.
In conclusion, we wanted to take advantage of temperature changes as a variable environmental signal, allowing our engineered bacteria to function at lower temperatures and Lysis proteins to lysis the engineered E. coli cells at higher temperatures, resulting in control of the engineered bacteria and release of the product.
Codon improvement and optimization
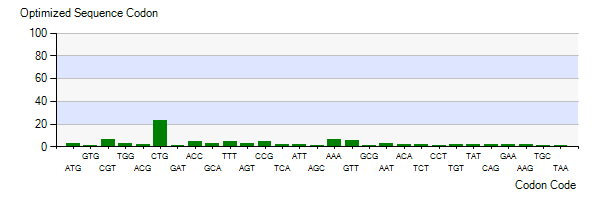
The Lysis gene we used was expressed in Pseudomonas lundensis. To better express the Lysis gene in engineered bacteria, we optimized the codon of the Lysis gene according to the codon preference of Escherichia coli. Figure 5-3 shows the number of codons we optimized to make our codons more in line with Escherichia coli preference. The modified Lysis gene is shown in BBa No.K4182007.
Figure 3: Optimized Sequence Codon in plasmid Ⅴ
Optimization of Plasmid
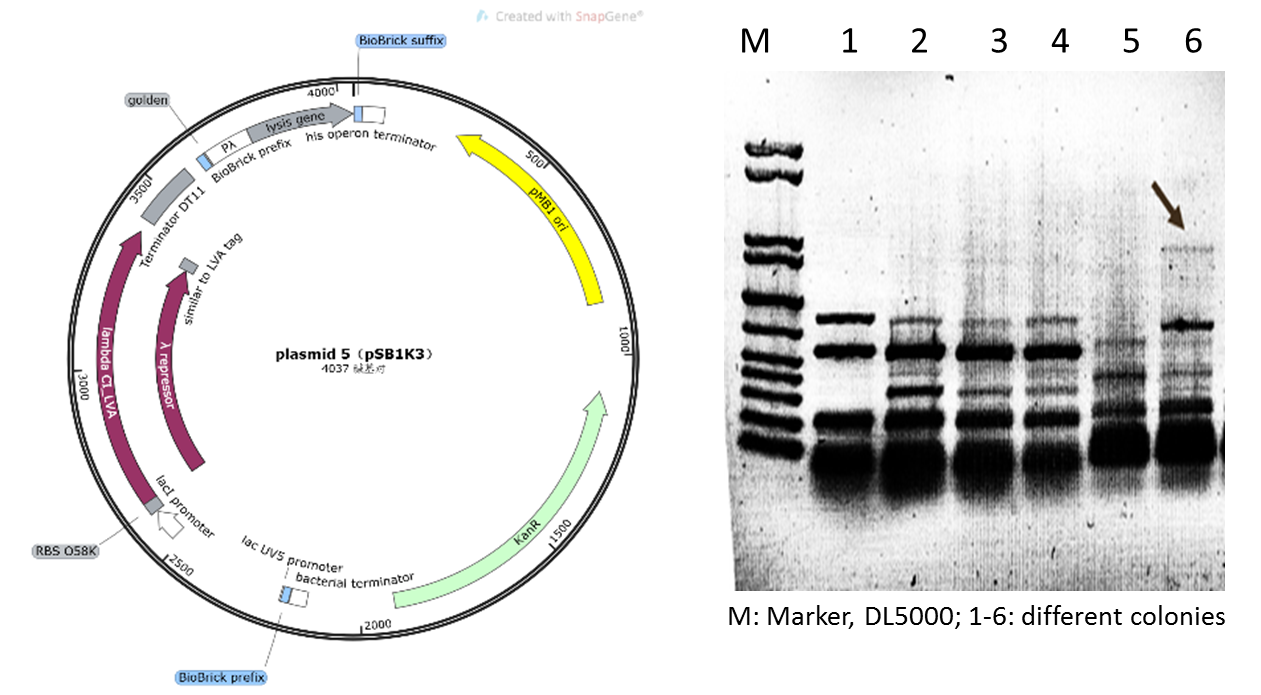
Initially, based on the design of the assay protocol, we planned to construct plasmid 5 using vector backbone pSB1K3. However, in our subsequent experiments, it was found that when the plasmids designed in this way were transferred to DH5α cells after Golden Gate cloning for expression, only dark target bands could be observed in colony PCR (Figure 5), and the extraction of plasmids and sequencing could not be completed due to the low concentration.
Figure 4: Plasmid 5 map based on pSB1K3 and its verification (he target band is approximately 1600bp)
Therefore, we supposed that due to the insufficient copy amount of pSB1K3 plasmid, we could not extract the product with a sufficient concentration in the engineered bacteria. Therefore, we replaced the vector of plasmid 5 with pSEVA341, a higher-copy-number vector and re-constructed the plasmid (Figure 6). As shown in Figure 6, the obvious target bands were observed, and the plasmid was further confirmed by sequencing.
Figure 5: new plasmid 5 with pSEVA341 backbone and its verification
The verification of heat triggered cell lysis and suicide
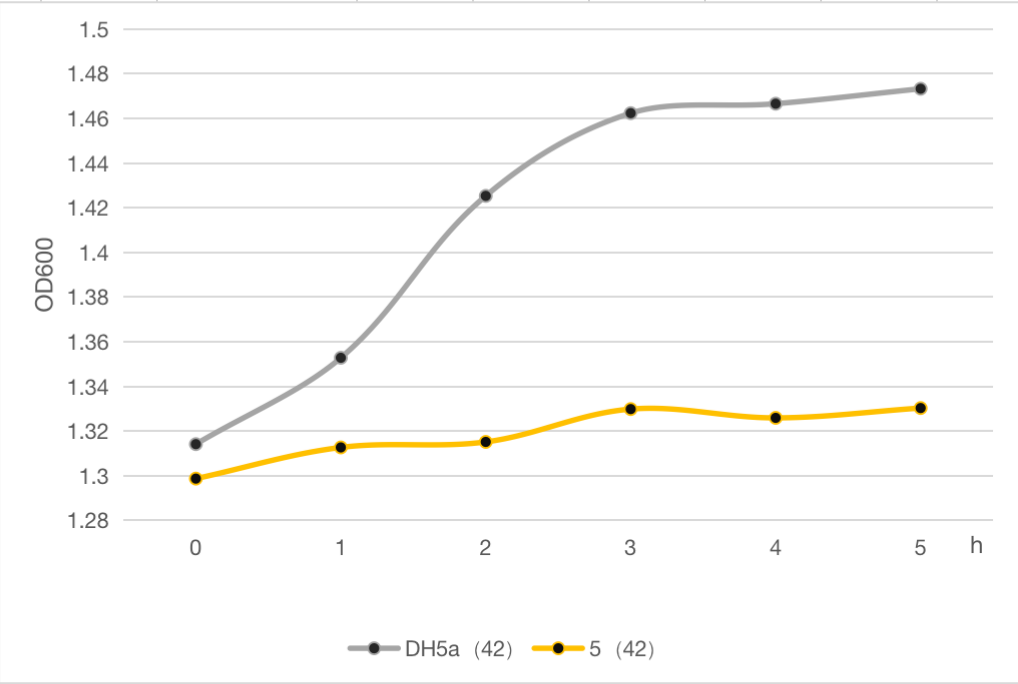
The engineered cell harboring plasmid 5 and blank vector respectively, were culture at 30℃ overnight, and then the temperature was shift to 42℃, while the control group were still cultured at 30℃. The OD600 of each group was detected every 1 h, and the growth curve of these strains were determined as follows.
Compared to the commonly used suicide protein MazF, the lysis protein in our study is shorter and easy to be manipulated, which can be used as an alternative and update for MazF.
Figure 6: Line chart of thallus concentration of DH5α and 5 at 42 ° C
References
1. Din, M.O., et al., Synchronized cycles of bacterial lysis for in vivo delivery. Nature, 2016. 536(7614): p. 81-85.
2. Saeidi, N., et al., Engineering microbes to sense and eradicate Pseudomonas aeruginosa, a human pathogen. Mol Syst Biol, 2011. 7: p. 521.
3. Restrepo-Pineda, S., et al., Thermoinducible expression system for producing recombinant proteins in Escherichia coli: advances and insights. FEMS Microbiol Rev, 2021. 45(6).
4. Aparicio, T., V. de Lorenzo, and E. Martínez-García, Improved Thermotolerance of Genome-Reduced Pseudomonas putida EM42 Enables Effective Functioning of the PL/cI857 System. Biotechnology Journal, 2019. 14(1): p. 1800483.
Characterized by BNUZH-China 2022
In this module, we replaced GOI with MazF. We hoped that when illuminated with the 660 nm light, MazF can be transcription and translation. Then MazF works, killing the engineered cells. Figure 1 illustrates the detailed design of the whole device.
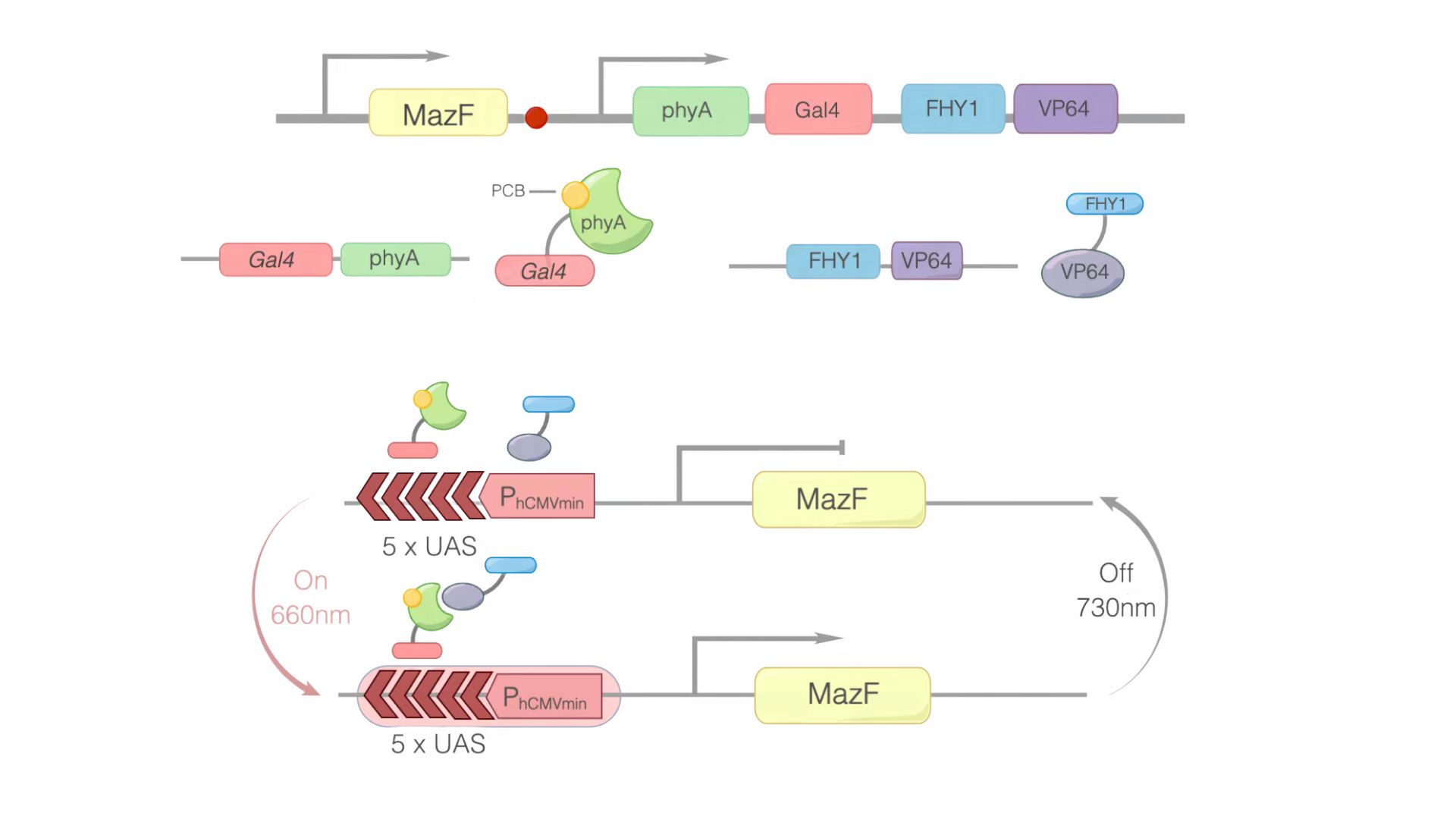
Figure 1: Construct design and the principle of the REDMAP-MazF. 5×UAS-P(hCMVmin) was fused to MazF. Gal4 was linked via a linker to PhyA. At the same time, PCB was combined with PhyA. PhyA turned to ΔPhyA. VP64 was linked with FHY1. When 660 nm light illuminating, ΔPhyA combines with VP64. Then, Gal4-ΔPhyA is transported into the nucleus by VP64-FHY1. After that, Gal4 will combine with 5×UAS-P(hCMVmin), activating the transcription of the downstream target gene: MazF. Then MazF works, killing the engineered cells.
Usage and Biology
We planned to use it to activate the transcription of MazF (BBa_K302033) to cause the engineered cells to commit suicide.
Experimental approach
For testing this device, we used HEK293T cells, which were seeded in 25cm2 flask. The PLVX vector was transiently transfected into HEK293T cells by Lipo8000™ transfection system and incubated for 43 h to ensure normal cell status. Then, phycocyanobilin (PCB) with a final concentration of 5 μM was added to the system, and the system was illuminated with red light (660 nm) for 3 h. After illumination, the cells were observed and the cell lysates were analyzed by SDS-PAGE and Western Blot.
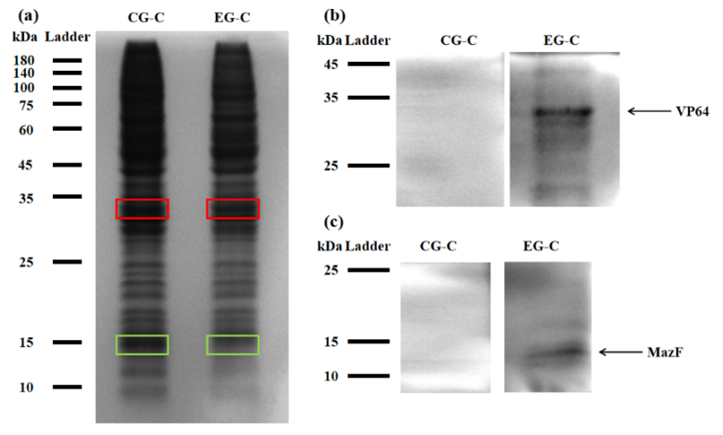
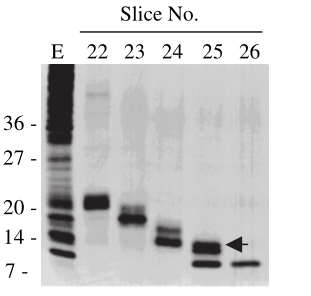
Figure 2: (a) REDMAP SDS-PAGE results. The VP64 strip is in the red box, and the MazF strip is in the green box. CG-C was the cell lysate of the control group, and EG-C was the cell lysate of the experimental group. (b) VP64 Western Blot diagram. CG-C was the cell lysate of the control group, EG-C was the cell lysate of the experimental group, and the theoretical size of VP64 was 30 kDa. (c) MazF Western Blot. CG-C was the cell lysate of the control group, EG-C was the cell lysate of the experimental group, and the theoretical MazF size was 13 kDa.
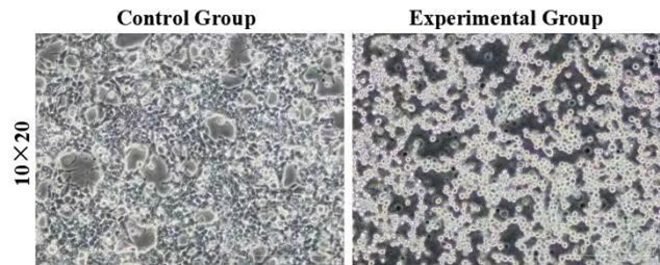
Figure 3: Morphology of control and experimental cells after red light verification. The control group was HEK293T cells without transient transfection, and the experimental group was HEK293T cells with transient transfection. 10×20 refer to the multiple of the eyepiece × the objective.
References
[1] Park, J. H., Yamaguchi, Y. & Inouye, M. Bacillus subtilis MazF-bs (EndoA) is a UACAU-specific mRNA interferase. FEBS Lett 585, 2526-2532 (2011). https://doi.org:10.1016/j.febslet.2011.07.008
[2] Zhou, Y., Kong, D., Wang, X. et al. A small and highly sensitive red/far-red optogenetic switch for applications in mammals. Nat Biotechnol 40, 262–272 (2022). https://doi.org/10.1038/s41587-021-01036-w
[3] Simanshu, D. K., Yamaguchi, Y., Park, J. H., Inouye, M. & Patel, D. J. Structural basis of mRNA recognition and cleavage by toxin MazF and its regulation by antitoxin MazE in Bacillus subtilis. Mol Cell 52, 447-458 (2013). https://doi.org:10.1016/j.molcel.2013.09.006
[4] Stein, T. Bacillus subtilis antibiotics: structures, syntheses and specific functions. Mol Microbiol 56, 845-857 (2005). https://doi.org:10.1111/j.1365-2958.2005.04587.x
[5] Pellegrini, O., Mathy, N., Gogos, A., Shapiro, L. & Condon, C. The Bacillus subtilis ydcDE operon encodes an endoribonuclease of the MazF/PemK family and its inhibitor. Mol Microbiol 56, 1139-1148 (2005). https://doi.org:10.1111/j.1365-2958.2005.04606.x