Difference between revisions of "Part:BBa K4365017"
Svetlana ia (Talk | contribs) (→Usage and Biology) |
|||
| (2 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
| − | |||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K4365017 short</partinfo> | <partinfo>BBa_K4365017 short</partinfo> | ||
| − | HFBI is a | + | HFBI is a protein found in <i>Trichoderma reesei</i> and belongs to a family of highly secreted proteins called hydrophobins. The signal peptide from the protein led to <html><a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365020">turboRFP</a></html> secretion in ''Saccharomyces cerevisiae''. |
| − | <!-- | + | {| class="wikitable" style="margin-left: auto; margin-right: auto; border: none; width:600px;" |
| − | === | + | |+ Collection of fungal signal peptides |
| + | |- | ||
| + | ! | ||
| + | ! ID | ||
| + | ! Source | ||
| + | ! Species | ||
| + | ! Prediction coefficient <ref name="signalp">José Juan Almagro Armenteros et al. (2019) SignalP 5.0 improves signal peptide predictions using deep neural networks Nature Biotechnology, 37, 420-423, doi: 10.1038/s41587-019-0036-z </ref> | ||
| + | ! turboRFP secretion | ||
| + | |- style="text-align:center;" | ||
| + | |sp1 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365006 BBa_K4365006] || MPGI || <i>Magnaporthe grisea</i> || 0.9562 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp2 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365007 BBa_K4365007] || RodA || <i>Aspergillus nidulans</i> || 0.9553 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp3 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365008 BBa_K4365008] || HYPI || <i>Aspergillus fumigatus</i> || 0.9703 || No | ||
| + | |- style="text-align:center;" | ||
| + | |sp4 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365009 BBa_K4365009] || SsgA || <i>Metarhizium anisopliae</i> || 0.9126 || No | ||
| + | |- style="text-align:center;" | ||
| + | |sp5 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365010 BBa_K4365010] || HCF-4 || <i>Cladosporium fulvum</i> || 0.9126 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp6 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365025 BBa_K4365025] || HCF-5 || <i>Cladosporium fulvum</i> || 0.9234 || NA | ||
| + | |- style="text-align:center;" | ||
| + | |sp7 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365026 BBa_K4365026] || CCG-2 || <i>Neurospora crassa</i> || 0.9566 || NA | ||
| + | |- style="text-align:center;" | ||
| + | |sp8 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365011 BBa_K4365011] || HCF-1 || <i>Cladosporium fulvum</i> || 0.6850 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp9 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365027 BBa_K4365027] || HCF-2 || <i>Cladosporium fulvum</i> || 0.8144 || NA | ||
| + | |- style="text-align:center;" | ||
| + | |sp10 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365012 BBa_K4365012] || HCF-3 || <i>Cladosporium fulvum</i> || 0.7934 || No | ||
| + | |- style="text-align:center;" | ||
| + | |sp11 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365013 BBa_K4365013] || SC3 || <i>Schizophyllum commune</i> || 0.6327 || No | ||
| + | |- style="text-align:center;" | ||
| + | |sp12 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365014 BBa_K4365014] || ABH3 || <i>Agaricus bisporus</i> || 0.9458 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp13 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365028 BBa_K4365028] || COH1 || <i>Coprinus cinereus</i> || 0.8545 || NA | ||
| + | |- style="text-align:center;" | ||
| + | |sp14 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365015 BBa_K4365015] || FBH1 || <i>Pleurotus ostreatus</i> || 0.3367 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp15 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365016 BBa_K4365016] || Aa-PRI2 || <i>Agrocybe aegerita</i> || 0.6356 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp16 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365033 BBa_K4365033] || Hyd-Pt1 || <i>Pisolithus tinctorius</i> || 0.8352 || NA | ||
| + | |- style="text-align:center;" | ||
| + | |sp17 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365017 BBa_K4365017] || HFBI || <i>Trichoderma reesei</i> || 0.9851 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp18 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365018 BBa_K4365018] || HFBII || <i>Trichoderma reesei</i> || 0.6862 || '''Yes''' | ||
| + | |- style="text-align:center;" | ||
| + | |sp19 || [https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365019 BBa_K4365019] || α-mating factor || <i>Saccharomyces cerevisiae</i> || 0.9988 || '''Yes''' | ||
| + | |} | ||
| − | < | + | <references /> |
| + | |||
| + | <br> | ||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
<partinfo>BBa_K4365017 SequenceAndFeatures</partinfo> | <partinfo>BBa_K4365017 SequenceAndFeatures</partinfo> | ||
| + | |||
| + | |||
| + | ===Usage and Biology=== | ||
| + | Secretion signal peptides can be fused to a protein to direct it through the secretory pathway out of the cell <ref name="2albert">Alberts, B. (2015) Molecular Biology of the Cell. 6th Edition, Garland Science, Taylor and Francis Group, New York.</ref>. Such secretion of recombinant proteins facilitates the protein purification process and the removal of GMOs <ref name="1kirsten">Kirsten Kottmeier, Kai Ostermann, Thomas Bley, Gerhard Rödel (2011) Hydrophobin signal sequence mediates efficient secretion of recombinant proteins in Pichia pastoris, Appl Microbiol Biotechnol 91, pages133–141, https://doi.org/10.1007/s00253-011-3246-y</ref>. However, commonly used signal sequences like the ''S. cerevisiae'' alpha-factor prepro-peptide <html><a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365019">K4365019</a></html> and the HSP150, are very long <ref name="1kirsten" /><ref>Russo P, Kalkkinen N, Sareneva H, Paakkola J, Makarow M. A heat shock gene from Saccharomyces cerevisiae encoding a secretory glycoprotein. Proc Natl Acad Sci U S A. 1992 May 1;89(9):3671-5. doi: 10.1073/pnas.89.9.3671. Erratum in: Proc Natl Acad Sci U S A. 1992 Sep 15;89(18):8857. PMID: 1570286; PMCID: PMC525552.</ref> which makes their use in genetic constructs limited. <!--This is a part to change for other peptides--> '''K4365017''' as a part of '''the collection of signal peptides''' represents a shorter sequence that could be inserted into the gene of interest simply by PCR and could display higher secretion efficiency. <!--This is a part to change for other peptides--> | ||
| + | |||
| + | <!--This is a part to change citations for other peptides--> | ||
| + | The sequences of the hydrophobic signal peptide was collected from literature <ref>M. G. Waters, E. A. Evans, G. Blobel (1988) Prepro-alpha-factor has a cleavable signal sequence, Journal of biological chemistry Volume 263, ISSUE 13, P6209-6214, https://doi.org/10.1016/S0021-9258(18)68773-3 </ref> and was extracted via analysis of their sequence using the SignalP - 5.0 signal peptide predictor tool <ref>José Juan Almagro Armenteros et al. (2019) SignalP 5.0 improves signal peptide predictions using deep neural networks Nature Biotechnology, 37, 420-423, doi: 10.1038/s41587-019-0036-z </ref>. | ||
| + | <!--This is a part to change citations for other peptides--> | ||
| + | |||
| + | Identified sequence was codon-optimized for ''S. cerevisiae'' using the codon optimization tool on GenScript. Then it was synthesized by IDT as a gBlock together with <html>a flexible glycine linker (<a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365005">K4365005</a>) and the turboRFP gene (<a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365020">K4365020</a>)</html> <ref>Merzlyak, E., Goedhart, J., Shcherbo, D. et al. (2007) Bright monomeric red fluorescent protein with an extended fluorescence lifetime. Nat Methods 4, 555–557 (2007). https://doi.org/10.1038/nmeth1062</ref>. | ||
| + | |||
| + | ====Hydrophobins==== | ||
| + | |||
| + | Hydrophobins are a group of small (~100 amino acids) proteins that are expressed in ascomycetes and basidiomycetes, which are very efficiently secreted <ref name="1kirsten" />. | ||
| + | |||
| + | After secretion from the fungus, they can assemble on the cell walls to form a hydrophobic coating. For this reason, the functions of hydrophobins are related to cell surface activity. For example, they can cover hyphae so the surface tension can be lowered and the hyphae can breach the water/air interface, thus escaping from the aqueous environment and growing into the aerial hyphae <ref name="5khal">Khalesi, M., Gebruers, K. & Derdelinckx, G. (2015) Recent Advances in Fungal Hydrophobin Towards Using in Industry, Protein J, 34, 243–255 . https://doi.org/10.1007/s10930-015-9621-2 </ref>. They also mediate the adhesion of the hyphae to hydrophobic surfaces such as in the case of pathogen-host interaction or symbiosis, as observed in lichens and in ectomycorrhizae (the symbiotic relationship between a fungal symbiont and the roots of a plant) <ref name="5khal" />. Hydrophobins also form a hydrophobic sheath to protect the surface of conidia, spores, and caps of fungi against wetting and desiccation <ref name="5khal" />. | ||
| + | |||
| + | ====Signal peptide secretion screening==== | ||
| + | |||
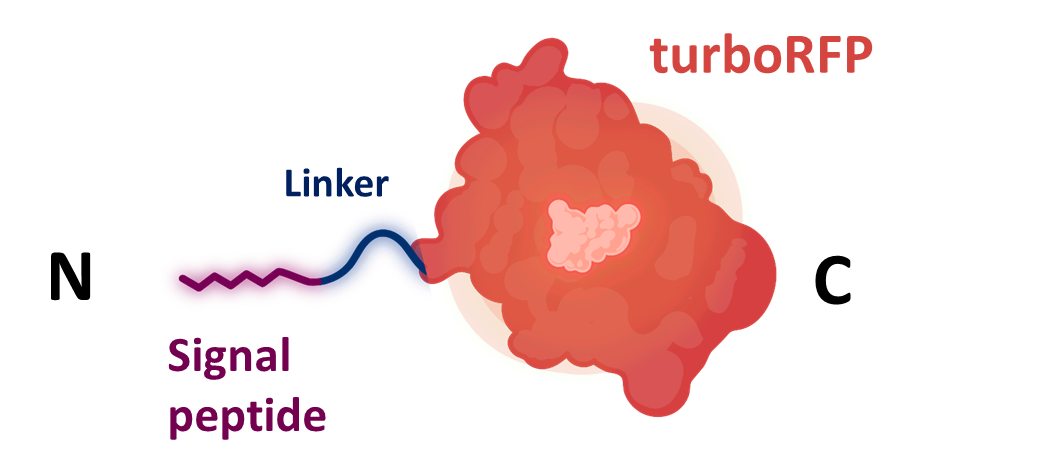
| + | Signal peptide siquence was cloned together with turboRFP <html><a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365020">K4365020</a> into the level 1 shuttle plasmid <a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365022">K4365022</a></html> (Figure 1). | ||
| + | |||
| + | [[File:BBa_K4365006_screening.png|500px|center|thumb|Figure 1: fusion protein used for the secretion assays composed of a signal peptide, a flexible linker, and the turboRFP fluorescent protein. Created with Biorender.com.]] | ||
| + | |||
| + | After the transformation with the shuttle plasmid, the yeast colonies were grown on W0 medium plates supplemented with histidine, leucine, and methionine and were used for secretion assays in flasks and in a 48-well plate. The α-mating factor pre-pro signal peptide <html><a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_K4365019">K4365019</a></html> was used as a positive control as it has been shown to be able to secrete proteins <ref name="19penas">Peñas M. M. et al. (1998) Identification, characterization, and In situ detection of a fruit-body-specific hydrophobin of Pleurotus ostreatus, Appl Environ Microbiol. 64(10):4028-34. doi: 10.1128/AEM.64.10.4028-4034.1998. PMID: 9758836; PMCID: PMC106595.</ref>. A turboRFP without signal peptide was employed as a negative control to account for the amount of free turboRFP present in the media due to cell lysis. | ||
| + | |||
| + | '''Secretion measurement protocols:''' | ||
| + | *[https://parts.igem.org/Part:BBa_K4365021#Secretion_assay_in_a_flask_protocol flask protocol] | ||
| + | *[https://parts.igem.org/Part:BBa_K4365021#Secretion_assay_in_a_48-well_plate_protocol 48-well plate protocol] | ||
| + | |||
| + | <!--This part different between sps--> | ||
| + | The first secretion experiment was conducted in flasks and showed (Figure 2A) that cell lysis was low as indicated by the low fluorescence of the negative control. In order to have consistent results and ensure confidence we set up biological triplicates. For this, turboRFP secretion was measured using 48-well plate approach. Cells with induced expression were pipetted into the wells of the plate and grown overnight. Starting at 1 hour after induction, samples were taken and the growth rate was monitored by measuring turbidity (NTU) and the development of fluorescence in the supernatant was measured on a plate reader. | ||
| + | |||
| + | K4365017 showed turboRFP secretion in both flask and 48-well plate measurement approaches (Figure 2). | ||
| + | |||
| + | <!--This part different between sps--> | ||
| + | |||
| + | [[File:BBa_K4365017_secretion.png|600px|center|thumb| Figure 2: '''A''' secretion by signal peptides in flasks. | ||
| + | '''B''' secretion assay in 48-well plate with an initial cell density of 1 OD. The canonical α-mating factor pre-pro signal peptide <ref name="19penas" /> (dark grey + control) was used as a positive control as it has been shown to be able to secrete proteins. A turboRFP without signal peptide was employed as a negative control to account for cell lysis (light grey, - control). The signal corresponding to K4365017 is depicted as a dashed line. Error bars represent mean ± SD.]] | ||
| + | |||
| + | ===References=== | ||
| + | |||
| + | |||
Latest revision as of 22:46, 11 October 2022
Signal peptide of HFBI from Trichoderma reesei
HFBI is a protein found in Trichoderma reesei and belongs to a family of highly secreted proteins called hydrophobins. The signal peptide from the protein led to turboRFP secretion in Saccharomyces cerevisiae.
| ID | Source | Species | Prediction coefficient [1] | turboRFP secretion | |
|---|---|---|---|---|---|
| sp1 | BBa_K4365006 | MPGI | Magnaporthe grisea | 0.9562 | Yes |
| sp2 | BBa_K4365007 | RodA | Aspergillus nidulans | 0.9553 | Yes |
| sp3 | BBa_K4365008 | HYPI | Aspergillus fumigatus | 0.9703 | No |
| sp4 | BBa_K4365009 | SsgA | Metarhizium anisopliae | 0.9126 | No |
| sp5 | BBa_K4365010 | HCF-4 | Cladosporium fulvum | 0.9126 | Yes |
| sp6 | BBa_K4365025 | HCF-5 | Cladosporium fulvum | 0.9234 | NA |
| sp7 | BBa_K4365026 | CCG-2 | Neurospora crassa | 0.9566 | NA |
| sp8 | BBa_K4365011 | HCF-1 | Cladosporium fulvum | 0.6850 | Yes |
| sp9 | BBa_K4365027 | HCF-2 | Cladosporium fulvum | 0.8144 | NA |
| sp10 | BBa_K4365012 | HCF-3 | Cladosporium fulvum | 0.7934 | No |
| sp11 | BBa_K4365013 | SC3 | Schizophyllum commune | 0.6327 | No |
| sp12 | BBa_K4365014 | ABH3 | Agaricus bisporus | 0.9458 | Yes |
| sp13 | BBa_K4365028 | COH1 | Coprinus cinereus | 0.8545 | NA |
| sp14 | BBa_K4365015 | FBH1 | Pleurotus ostreatus | 0.3367 | Yes |
| sp15 | BBa_K4365016 | Aa-PRI2 | Agrocybe aegerita | 0.6356 | Yes |
| sp16 | BBa_K4365033 | Hyd-Pt1 | Pisolithus tinctorius | 0.8352 | NA |
| sp17 | BBa_K4365017 | HFBI | Trichoderma reesei | 0.9851 | Yes |
| sp18 | BBa_K4365018 | HFBII | Trichoderma reesei | 0.6862 | Yes |
| sp19 | BBa_K4365019 | α-mating factor | Saccharomyces cerevisiae | 0.9988 | Yes |
- ↑ José Juan Almagro Armenteros et al. (2019) SignalP 5.0 improves signal peptide predictions using deep neural networks Nature Biotechnology, 37, 420-423, doi: 10.1038/s41587-019-0036-z
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Usage and Biology
Secretion signal peptides can be fused to a protein to direct it through the secretory pathway out of the cell [1]. Such secretion of recombinant proteins facilitates the protein purification process and the removal of GMOs [2]. However, commonly used signal sequences like the S. cerevisiae alpha-factor prepro-peptide K4365019 and the HSP150, are very long [2][3] which makes their use in genetic constructs limited. K4365017 as a part of the collection of signal peptides represents a shorter sequence that could be inserted into the gene of interest simply by PCR and could display higher secretion efficiency.
The sequences of the hydrophobic signal peptide was collected from literature [4] and was extracted via analysis of their sequence using the SignalP - 5.0 signal peptide predictor tool [5].
Identified sequence was codon-optimized for S. cerevisiae using the codon optimization tool on GenScript. Then it was synthesized by IDT as a gBlock together with a flexible glycine linker (K4365005) and the turboRFP gene (K4365020) [6].
Hydrophobins
Hydrophobins are a group of small (~100 amino acids) proteins that are expressed in ascomycetes and basidiomycetes, which are very efficiently secreted [2].
After secretion from the fungus, they can assemble on the cell walls to form a hydrophobic coating. For this reason, the functions of hydrophobins are related to cell surface activity. For example, they can cover hyphae so the surface tension can be lowered and the hyphae can breach the water/air interface, thus escaping from the aqueous environment and growing into the aerial hyphae [7]. They also mediate the adhesion of the hyphae to hydrophobic surfaces such as in the case of pathogen-host interaction or symbiosis, as observed in lichens and in ectomycorrhizae (the symbiotic relationship between a fungal symbiont and the roots of a plant) [7]. Hydrophobins also form a hydrophobic sheath to protect the surface of conidia, spores, and caps of fungi against wetting and desiccation [7].
Signal peptide secretion screening
Signal peptide siquence was cloned together with turboRFP K4365020 into the level 1 shuttle plasmid K4365022 (Figure 1).
After the transformation with the shuttle plasmid, the yeast colonies were grown on W0 medium plates supplemented with histidine, leucine, and methionine and were used for secretion assays in flasks and in a 48-well plate. The α-mating factor pre-pro signal peptide K4365019 was used as a positive control as it has been shown to be able to secrete proteins [8]. A turboRFP without signal peptide was employed as a negative control to account for the amount of free turboRFP present in the media due to cell lysis.
Secretion measurement protocols:
The first secretion experiment was conducted in flasks and showed (Figure 2A) that cell lysis was low as indicated by the low fluorescence of the negative control. In order to have consistent results and ensure confidence we set up biological triplicates. For this, turboRFP secretion was measured using 48-well plate approach. Cells with induced expression were pipetted into the wells of the plate and grown overnight. Starting at 1 hour after induction, samples were taken and the growth rate was monitored by measuring turbidity (NTU) and the development of fluorescence in the supernatant was measured on a plate reader.
K4365017 showed turboRFP secretion in both flask and 48-well plate measurement approaches (Figure 2).

References
- ↑ Alberts, B. (2015) Molecular Biology of the Cell. 6th Edition, Garland Science, Taylor and Francis Group, New York.
- ↑ 2.0 2.1 2.2 Kirsten Kottmeier, Kai Ostermann, Thomas Bley, Gerhard Rödel (2011) Hydrophobin signal sequence mediates efficient secretion of recombinant proteins in Pichia pastoris, Appl Microbiol Biotechnol 91, pages133–141, https://doi.org/10.1007/s00253-011-3246-y
- ↑ Russo P, Kalkkinen N, Sareneva H, Paakkola J, Makarow M. A heat shock gene from Saccharomyces cerevisiae encoding a secretory glycoprotein. Proc Natl Acad Sci U S A. 1992 May 1;89(9):3671-5. doi: 10.1073/pnas.89.9.3671. Erratum in: Proc Natl Acad Sci U S A. 1992 Sep 15;89(18):8857. PMID: 1570286; PMCID: PMC525552.
- ↑ M. G. Waters, E. A. Evans, G. Blobel (1988) Prepro-alpha-factor has a cleavable signal sequence, Journal of biological chemistry Volume 263, ISSUE 13, P6209-6214, https://doi.org/10.1016/S0021-9258(18)68773-3
- ↑ José Juan Almagro Armenteros et al. (2019) SignalP 5.0 improves signal peptide predictions using deep neural networks Nature Biotechnology, 37, 420-423, doi: 10.1038/s41587-019-0036-z
- ↑ Merzlyak, E., Goedhart, J., Shcherbo, D. et al. (2007) Bright monomeric red fluorescent protein with an extended fluorescence lifetime. Nat Methods 4, 555–557 (2007). https://doi.org/10.1038/nmeth1062
- ↑ 7.0 7.1 7.2 Khalesi, M., Gebruers, K. & Derdelinckx, G. (2015) Recent Advances in Fungal Hydrophobin Towards Using in Industry, Protein J, 34, 243–255 . https://doi.org/10.1007/s10930-015-9621-2
- ↑ 8.0 8.1 Peñas M. M. et al. (1998) Identification, characterization, and In situ detection of a fruit-body-specific hydrophobin of Pleurotus ostreatus, Appl Environ Microbiol. 64(10):4028-34. doi: 10.1128/AEM.64.10.4028-4034.1998. PMID: 9758836; PMCID: PMC106595.