Difference between revisions of "Part:BBa K1907001"
| (20 intermediate revisions by 3 users not shown) | |||
| Line 3: | Line 3: | ||
<partinfo>BBa_K1907001 short</partinfo> | <partinfo>BBa_K1907001 short</partinfo> | ||
| + | <br> | ||
<p><b>Introduction</b></p> | <p><b>Introduction</b></p> | ||
This part can be used to introduce microcystinase (MlrA) from <i>Sphingomonas</i> sp. ACM-3962 to yeast. MlrA is an enzyme that degrades some cyanobacterial toxins, most notably microcystin-LR (MC-LR). To facilitate detection and purification of the enzyme, a 6X Histidine tag is fused on the C-terminus of MlrA, with a short linker in between. The full protein-coding sequence has been codon-optimized for expression in <i>Saccharomyces cerevisiae </i> by GeneArt. | This part can be used to introduce microcystinase (MlrA) from <i>Sphingomonas</i> sp. ACM-3962 to yeast. MlrA is an enzyme that degrades some cyanobacterial toxins, most notably microcystin-LR (MC-LR). To facilitate detection and purification of the enzyme, a 6X Histidine tag is fused on the C-terminus of MlrA, with a short linker in between. The full protein-coding sequence has been codon-optimized for expression in <i>Saccharomyces cerevisiae </i> by GeneArt. | ||
| − | + | <br> | |
<p> | <p> | ||
| Line 15: | Line 16: | ||
<p></p> | <p></p> | ||
| − | |||
| − | |||
<br> | <br> | ||
| + | <p><b>Background</b></p> | ||
Microcystinase, or MlrA, is a member of a set of enzymes that degrade microcystins. These enzymes - MlrA, MlrB, MlrC and MlrD - are found naturally in some gram-negative bacteria, for example <i>Sphingomonas</i> sp. They each have a different function in the overall process of microcystin degradation. MlrA is first in the sequence to act. It hydrolyzes the peptide bond between the Adda and arginine amino acids in microcystin-LR (MC-LR). The breakage of this bond linearizes the toxin (Figure 1). Following this, MlrB (a serine protease) and MlrC (a metalloprotease) are able to further degrade the linearized product into individual amino acids, which are then used in the cell’s metabolism. Therefore, the toxin is likely used as a carbon source. MlrD’s function is not fully understood at this point, but it may have a role in importing the toxins. (Bourne et al., 2001) | Microcystinase, or MlrA, is a member of a set of enzymes that degrade microcystins. These enzymes - MlrA, MlrB, MlrC and MlrD - are found naturally in some gram-negative bacteria, for example <i>Sphingomonas</i> sp. They each have a different function in the overall process of microcystin degradation. MlrA is first in the sequence to act. It hydrolyzes the peptide bond between the Adda and arginine amino acids in microcystin-LR (MC-LR). The breakage of this bond linearizes the toxin (Figure 1). Following this, MlrB (a serine protease) and MlrC (a metalloprotease) are able to further degrade the linearized product into individual amino acids, which are then used in the cell’s metabolism. Therefore, the toxin is likely used as a carbon source. MlrD’s function is not fully understood at this point, but it may have a role in importing the toxins. (Bourne et al., 2001) | ||
| + | <br><br> | ||
| + | ------------------------------------------------------------------------------------------------------------------- | ||
| + | <p><b>Update from the 2021 Lethbridge Collegiate Team</b></p> | ||
| + | <br> | ||
| + | Xu <i>et al</i>., 2019 has completed a comprehensive analysis of the active site behaviour through computational and experimental methods. Their work demonstrates that the originally hypothesized active site as part of the HEXHH zinc-binding motif is sterically hindered and likely unable to cleave microcystins. They instead propose another active site and mechanism which categorizes MlrA as a type II CAAX prenyl protease, which was first noted due to the high similarity of the homology template MmRce1. The proposed mechanism involves Glu172 and His205 first activating a water molecules which permits nucleophilic attack of the MC-LR ADDA-Arg peptide bond, with Trp176 and Trp201 potentially raising the pKa for accelerating reaction kinetics, and finally the His260 and Asn264 from the metalloprotease-hypothesized active site serving to stabilize the transition state as an oxyanion hole. Furthermore, their experimental studies which involved comparative EDTA enzymatic inhibition assays also suggest mlrA is likely not a metalloprotease, but a glutamate intramembrane protease. This was further validated by predictive docking simulations for the bound complex and experimental studies of mutated active site residues (Xu et al., 2019). | ||
| + | <br> | ||
| + | Our team completed molecular dynamics (MD) simulations to further examine the behaviour of a very closely related MlrA enzyme (from the strain <i>Sphingomonas</i> sp. USTB-05). Our analysis on the trajectories reveals structural information, which will have similar active site behaviour to the MlrA from <i>Sphingomonas</i> sp. ACM-3962. Furthermore, both enzymes from each strain have higher sequence similarities with the type II CAAX prenyl proteases than metalloproteases (Xu <i>et al</i>., 2019). Our modeling results indicate that hydrogen atoms of water molecules are occupied within a 1.9 Angstroms distance for a significant amount of the simulations. This result supports the hypothesis by Xu <i>et al</i>., 2019 however it can be improved with further docking simulations on the trajectory ensembles generated from our MD production simulations and MC-LR ligand poses calculated through conformational search software designed for macrocyclic peptides. More information can be found on the modeling section of our wiki. | ||
| + | ----------------------------------------------------------------------------------------------------------------<br><br><br> | ||
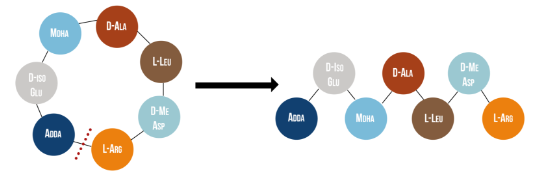
| − | [[File:Linearization.png|center|]] | + | [[File:Linearization.png|600px|thumb|center|Figure 1: Linearization of microcystin-LR occurs when MlrA hydrolyzes the bond between two of the toxin’s amino acids, Adda and arginine. It has been reported that the linearization of MC-LR renders the toxin 160 times less toxic (Bourne et al., 1996).]] |
<p> | <p> | ||
| − | |||
</p> | </p> | ||
| − | < | + | <br> |
| − | + | ||
| − | + | ||
<p> | <p> | ||
| − | + | Based on a homologous structure model (Figure 2) obtained from the Phyre2 tool (Kelley et al., 2015), it seems that the enzyme is a transmembrane protein consisting of eight hydrophobic α-helices that form a channel, with the presumed active site (Bourne et al., 2001) residing in the middle. It is worth pointing out, however, that the model does not cover the whole length of the MlrA sequence, as only very few homologs were found for the sequence - the homology model is mostly based on two homologs that didn’t cover the very ends of either terminus. Other than this evidence, there are previous studies pointing to the direction of MlrA being a membrane protein (Pei et al., 2011). However, when the enzyme was heterologously produced in <i> E. coli </i>, it was noticed that it had localised mostly in the soluble fraction of the cytosol, in an active state (Dziga et al., 2012). | |
</p> | </p> | ||
| + | |||
| + | [[File:homology2.png|300px|thumb|center|Figure 2: A homology-based structure model of MlrA made with Phyre2 and visualised with UCSF Chimera. Based on the model, it seems that MlrA has a typical membrane protein structure consisting of set of hydrophobic transmembrane helices.. The presumed active site (Bourne et al., 2001) resides in the middle of a channel formed by the helices.]] | ||
| + | |||
| + | <br> | ||
<p>The mechanism of MlrA’s function is not yet fully understood. The homologous structure model gives some suggestions as to how the enzyme might work: it seems that one end of the channel is wider than the other, and that the active site resides in the middle of it. Consequently, if the enzyme is located on the outer membrane of the cells, it might actually be a transporter which the cells use to import the toxin and simultaneously linearize it. | <p>The mechanism of MlrA’s function is not yet fully understood. The homologous structure model gives some suggestions as to how the enzyme might work: it seems that one end of the channel is wider than the other, and that the active site resides in the middle of it. Consequently, if the enzyme is located on the outer membrane of the cells, it might actually be a transporter which the cells use to import the toxin and simultaneously linearize it. | ||
</p> | </p> | ||
| + | |||
| + | <br> | ||
| + | <p><b>Previous use</b></p><p> | ||
| + | This part was used in <i>S. cerevisiae</i> strain SS328-leu. It was used in the pRS415-GAL plasmid backbone, with the insert being cloned into the plasmid using the SpeI and XhoI restriction sites (5' and 3', respectively). These restriction sites flanked the protein-coding site directly. The part was modified to be flanked by the BioBrick standard prefix and suffix for shipping to the registry. | ||
| + | </p> | ||
| + | <br> | ||
<p><b>Expression</b></p> | <p><b>Expression</b></p> | ||
| − | + | To produce the protein, a 20 mL pre-culture was prepared in SD-leu + 2% glucose and grown overnight at 30 °C. The culture was then diluted into a final volume of 500 mL using SD-leu + 0.5% glucose, to an OD600 value of 0.2. This culture was refreshed in 30 °C for 6 h, and then induced by adding galactose to a final concentration of 2%. Expression was continued overnight, then cells were harvested. Samples from different stages of cell culture and cell lysis were collected for SDS-PAGE and analysis with western blot utlizing anti-His antibodies. The results of this analysis are presented in figure 3. It was observed that after cell lysis, the protein localized mainly in the insoluble fraction. | |
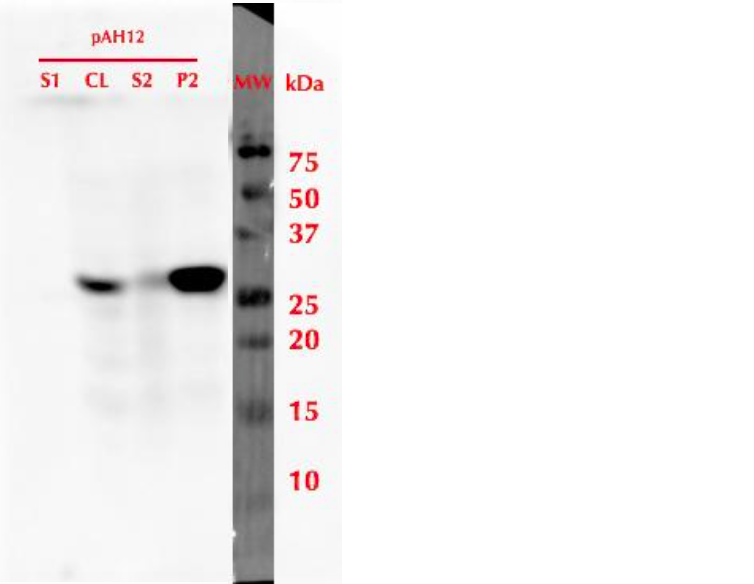
| − | [[File:MlrAY_WB.png|300px|thumb|center|]] | + | [[File:MlrAY_WB.png|300px|thumb|center|Figure 3: Western blot of MlrA expression in yeast. S1 - growth medium before lysing cells, CL - cell lysate, S2 - supernatant from cell lysate and P2 - pellet from cell lysate.]] |
| − | < | + | <br> |
| − | + | ||
| − | + | ||
| + | <p><b>Localization</b></p> | ||
| + | For determining the location of the protein, yeast cells were lysed and fractions were further processed to obtain information on the different cellular components. All of these fractions were then analyzed by western blotting (Figure 4). MlrA was detected in the fraction comprising inclusion bodies, plasma membrane and cell wall (P2 fraction) as well as fraction containing only cell wall and plasma membrane (P3 fraction), indicating that the protein is localized in the plasma membrane. Unlike previously, no enzyme was detected in the soluble fraction. | ||
| + | |||
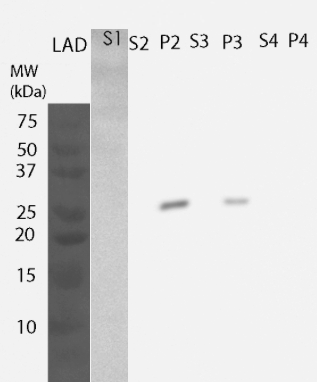
| + | [[File:Localisation.png|300px|thumb|center|Figure 4. Localisation assay of MlrA. S1 - growth medium, S2 - supernatant after cell lysis, P2 - pellet after cell lysis, S3 - re-folded proteins, P3 - plasma membrane + cell wall, S4 - soluble protein, P4 - inner membranes.]] | ||
| + | |||
| + | <br> | ||
<p><b>Activity measurements</b></p> | <p><b>Activity measurements</b></p> | ||
| Line 54: | Line 73: | ||
<p> | <p> | ||
| − | The results we acquired from this construct show that the amount of microcystin decreases, which means that the enzyme is active (Figure | + | The results we acquired from this construct show that the amount of microcystin decreases, which means that the enzyme is active (Figure 5). The results also show that for example the 1:10 dilution degrades the microcystin-LR about ten times faster than the 1:100 dilution. |
</p> | </p> | ||
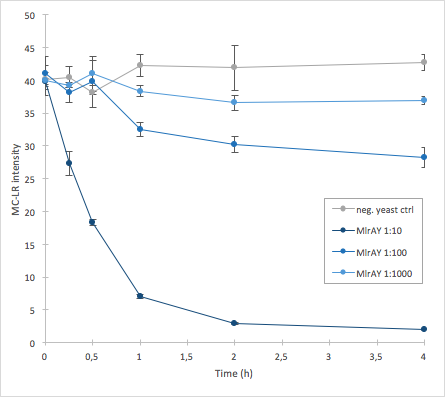
| − | [[File:MlrAY_activity.png|300px|thumb|center|]] | + | [[File:MlrAY_activity.png|300px|thumb|center|Figure 5: Enzymatic activity of MlrA produced in <i>S. cerevisiae </i>, plotted values are means of three replicates. Results show that the 1:10 dilution has the highest activity whereas 1:1000 has the lowest.]] |
| − | < | + | <br> |
| − | + | <br> | |
| − | + | ||
| + | For more details on the experiments done with this BioBrick, visit http://2016.igem.org/Team:Aalto-Helsinki/Laboratory | ||
| − | |||
<br> | <br> | ||
| + | |||
| + | <p><b>References</b> </p> | ||
| + | |||
| + | Bourne DG, Jones GJ, Blakeley RL, Jones A, Negri AP, Riddles P., 1996. Enzymatic Pathway for the Bacterial Degradation of the Cyanobacterial Cyclic Peptide Toxin Microcystin LR. Appl. Environ. Microbiol. 62(11), pp. 4086-4094. | ||
| + | |||
Bourne DG, Riddles P, Jones GJ, Smith W, Blakely RL., 2001. Characterisation of a gene cluster involved in bacterial degradation of the cyanobacterial toxin microcystin LR. Environ Toxicol. 16(6), pp. 523-34. | Bourne DG, Riddles P, Jones GJ, Smith W, Blakely RL., 2001. Characterisation of a gene cluster involved in bacterial degradation of the cyanobacterial toxin microcystin LR. Environ Toxicol. 16(6), pp. 523-34. | ||
Latest revision as of 02:51, 22 October 2021
Microcystinase (MlrA) for S. cerevisiae
Introduction
This part can be used to introduce microcystinase (MlrA) from Sphingomonas sp. ACM-3962 to yeast. MlrA is an enzyme that degrades some cyanobacterial toxins, most notably microcystin-LR (MC-LR). To facilitate detection and purification of the enzyme, a 6X Histidine tag is fused on the C-terminus of MlrA, with a short linker in between. The full protein-coding sequence has been codon-optimized for expression in Saccharomyces cerevisiae by GeneArt.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 166
Illegal BglII site found at 907 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 849
Background
Microcystinase, or MlrA, is a member of a set of enzymes that degrade microcystins. These enzymes - MlrA, MlrB, MlrC and MlrD - are found naturally in some gram-negative bacteria, for example Sphingomonas sp. They each have a different function in the overall process of microcystin degradation. MlrA is first in the sequence to act. It hydrolyzes the peptide bond between the Adda and arginine amino acids in microcystin-LR (MC-LR). The breakage of this bond linearizes the toxin (Figure 1). Following this, MlrB (a serine protease) and MlrC (a metalloprotease) are able to further degrade the linearized product into individual amino acids, which are then used in the cell’s metabolism. Therefore, the toxin is likely used as a carbon source. MlrD’s function is not fully understood at this point, but it may have a role in importing the toxins. (Bourne et al., 2001)
Update from the 2021 Lethbridge Collegiate Team
Xu et al., 2019 has completed a comprehensive analysis of the active site behaviour through computational and experimental methods. Their work demonstrates that the originally hypothesized active site as part of the HEXHH zinc-binding motif is sterically hindered and likely unable to cleave microcystins. They instead propose another active site and mechanism which categorizes MlrA as a type II CAAX prenyl protease, which was first noted due to the high similarity of the homology template MmRce1. The proposed mechanism involves Glu172 and His205 first activating a water molecules which permits nucleophilic attack of the MC-LR ADDA-Arg peptide bond, with Trp176 and Trp201 potentially raising the pKa for accelerating reaction kinetics, and finally the His260 and Asn264 from the metalloprotease-hypothesized active site serving to stabilize the transition state as an oxyanion hole. Furthermore, their experimental studies which involved comparative EDTA enzymatic inhibition assays also suggest mlrA is likely not a metalloprotease, but a glutamate intramembrane protease. This was further validated by predictive docking simulations for the bound complex and experimental studies of mutated active site residues (Xu et al., 2019).
Our team completed molecular dynamics (MD) simulations to further examine the behaviour of a very closely related MlrA enzyme (from the strain Sphingomonas sp. USTB-05). Our analysis on the trajectories reveals structural information, which will have similar active site behaviour to the MlrA from Sphingomonas sp. ACM-3962. Furthermore, both enzymes from each strain have higher sequence similarities with the type II CAAX prenyl proteases than metalloproteases (Xu et al., 2019). Our modeling results indicate that hydrogen atoms of water molecules are occupied within a 1.9 Angstroms distance for a significant amount of the simulations. This result supports the hypothesis by Xu et al., 2019 however it can be improved with further docking simulations on the trajectory ensembles generated from our MD production simulations and MC-LR ligand poses calculated through conformational search software designed for macrocyclic peptides. More information can be found on the modeling section of our wiki.
Based on a homologous structure model (Figure 2) obtained from the Phyre2 tool (Kelley et al., 2015), it seems that the enzyme is a transmembrane protein consisting of eight hydrophobic α-helices that form a channel, with the presumed active site (Bourne et al., 2001) residing in the middle. It is worth pointing out, however, that the model does not cover the whole length of the MlrA sequence, as only very few homologs were found for the sequence - the homology model is mostly based on two homologs that didn’t cover the very ends of either terminus. Other than this evidence, there are previous studies pointing to the direction of MlrA being a membrane protein (Pei et al., 2011). However, when the enzyme was heterologously produced in E. coli , it was noticed that it had localised mostly in the soluble fraction of the cytosol, in an active state (Dziga et al., 2012).
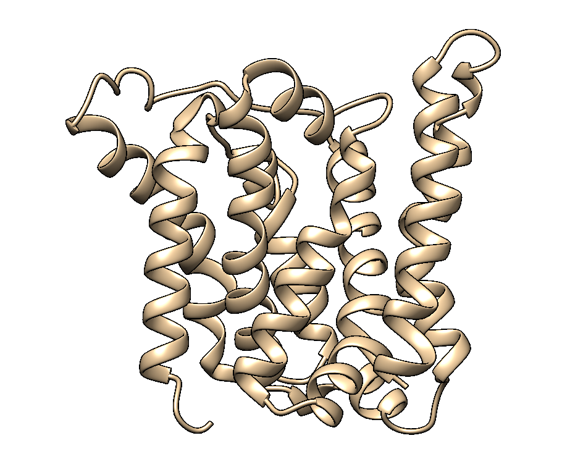
The mechanism of MlrA’s function is not yet fully understood. The homologous structure model gives some suggestions as to how the enzyme might work: it seems that one end of the channel is wider than the other, and that the active site resides in the middle of it. Consequently, if the enzyme is located on the outer membrane of the cells, it might actually be a transporter which the cells use to import the toxin and simultaneously linearize it.
Previous use
This part was used in S. cerevisiae strain SS328-leu. It was used in the pRS415-GAL plasmid backbone, with the insert being cloned into the plasmid using the SpeI and XhoI restriction sites (5' and 3', respectively). These restriction sites flanked the protein-coding site directly. The part was modified to be flanked by the BioBrick standard prefix and suffix for shipping to the registry.
Expression
To produce the protein, a 20 mL pre-culture was prepared in SD-leu + 2% glucose and grown overnight at 30 °C. The culture was then diluted into a final volume of 500 mL using SD-leu + 0.5% glucose, to an OD600 value of 0.2. This culture was refreshed in 30 °C for 6 h, and then induced by adding galactose to a final concentration of 2%. Expression was continued overnight, then cells were harvested. Samples from different stages of cell culture and cell lysis were collected for SDS-PAGE and analysis with western blot utlizing anti-His antibodies. The results of this analysis are presented in figure 3. It was observed that after cell lysis, the protein localized mainly in the insoluble fraction.
Localization
For determining the location of the protein, yeast cells were lysed and fractions were further processed to obtain information on the different cellular components. All of these fractions were then analyzed by western blotting (Figure 4). MlrA was detected in the fraction comprising inclusion bodies, plasma membrane and cell wall (P2 fraction) as well as fraction containing only cell wall and plasma membrane (P3 fraction), indicating that the protein is localized in the plasma membrane. Unlike previously, no enzyme was detected in the soluble fraction.
Activity measurements
The activity of MlrA was measured by incubating the enzyme in different dilutions with microcystin extract. The final concentration of microcystin-LR was estimated to be 1 μg/L. Samples were taken at different time points and they were analysed with liquid chromatography/mass spectrometry (LC-MS). This gives data of the relative abundance of microcystin-LR in each sample. The results were then plotted against time, together with a negative control which consisted of microcystin extract and cell lysate from S. cerevisiae SS328-leu strain which did not have an MlrA construct.
The results we acquired from this construct show that the amount of microcystin decreases, which means that the enzyme is active (Figure 5). The results also show that for example the 1:10 dilution degrades the microcystin-LR about ten times faster than the 1:100 dilution.
For more details on the experiments done with this BioBrick, visit http://2016.igem.org/Team:Aalto-Helsinki/Laboratory
References
Bourne DG, Jones GJ, Blakeley RL, Jones A, Negri AP, Riddles P., 1996. Enzymatic Pathway for the Bacterial Degradation of the Cyanobacterial Cyclic Peptide Toxin Microcystin LR. Appl. Environ. Microbiol. 62(11), pp. 4086-4094.
Bourne DG, Riddles P, Jones GJ, Smith W, Blakely RL., 2001. Characterisation of a gene cluster involved in bacterial degradation of the cyanobacterial toxin microcystin LR. Environ Toxicol. 16(6), pp. 523-34.
Dziga D, Wladyka B, Zielińska G, Meriluoto J, Wasylewski M., 2012. Heterologous expression and characterisation of microcystinase. Toxicon, 59(5), pp. 578-586.
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJE., 2015. The Phyre2 web portal for protein modeling, prediction and analysis. Nature Protocols 10, pp. 845–858.
Pei J, Mitchell DA, Dixon JE, Grishin NV., 2011. Expansion of type II CAAX proteases reveals evolutionary origin of g-secretase subunit APH-1. J. Mol. Biol. 410:18–26.