Difference between revisions of "Part:BBa K2010999"
| Line 54: | Line 54: | ||
=DUT_China 2021 improvement= | =DUT_China 2021 improvement= | ||
| − | Compared to the old part BBa_K2010999, expressing PETase, we design a new part [https://parts.igem.org/wiki/index.php?title=Part:BBa_K3898152 | + | Compared to the old part BBa_K2010999, expressing PETase, we design a new part [https://parts.igem.org/wiki/index.php?title=Part:BBa_K3898152 BBa_K3898152], which is expressing RIDD-PETase. PETase in our research was optimized, and it shows better enzyme activity and considerable degradation effect. <br> |
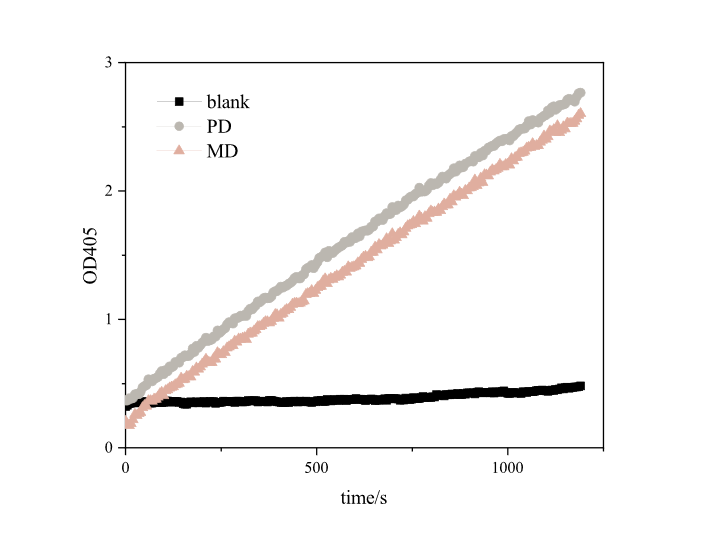
[[Image:T--DUT_China--es22.png|400px|thumb|left|'''Fig.2''' Enzyme activity analysis of pNPB . PD: RIDD-PETase enzyme solution, ; MD: RIDD-MHETase enzyme solution; Blank: normal E.coli BL21’s broken medium. '''(30℃, pH=8.0,1000μM pNPB)''']] | [[Image:T--DUT_China--es22.png|400px|thumb|left|'''Fig.2''' Enzyme activity analysis of pNPB . PD: RIDD-PETase enzyme solution, ; MD: RIDD-MHETase enzyme solution; Blank: normal E.coli BL21’s broken medium. '''(30℃, pH=8.0,1000μM pNPB)''']] | ||
Based on the results, we found that RIDD-PETase have esterase activity. After 20 minutes of reaction, each resulted in absorption peaks as high as 3.4873 at 405nm. Our optimized RIDD-PETase enzyme activity is 1.5 times higher than theirs in BBa_K2010999. <br> | Based on the results, we found that RIDD-PETase have esterase activity. After 20 minutes of reaction, each resulted in absorption peaks as high as 3.4873 at 405nm. Our optimized RIDD-PETase enzyme activity is 1.5 times higher than theirs in BBa_K2010999. <br> | ||
Revision as of 01:07, 22 October 2021
PETase (PET-degrading enzyme, origin I. sakaiensis)
PETase is a poly(ethylene terehphthalate)-degrading enzyme first identified in Ideonella sakaiensis. BBa_K2010999 is the sequence for the E. coli K12 optimized DNA sequence for PETase. The catalytic activity of IsPETase can be characterized using a p-nitrophenol butyrate (pNPB) assay, in which PETase cleaves the ester bond of pNPB to release p-nitrophenolate. The absorbance of this compound can be measured at 405 nm to reflect IsPETase activity.

This SDS-PAGE gel demonstrates successful protein purification of PETase. PETase has a molecular weight of approximately 30 kDa.

This graph demonstrates PETase activity with various substrate concentrations at a pH of 7.0.

This graph demonstrates PETase activity with various substrate concentrations at a pH of 8.0.

This graph demonstrates PETase activity with various substrate concentrations at a pH of 9.0.

This graph demonstrates PETase activity as a function of pH. PETase is most active at a pH of 9.0, but activity at 8.0 and 9.0 is practically indistinguishable.
Contribution
Group: KEYSTONE 2020
Summary: According to a study by Brandon C. Knott et al., PETase can be linked with MHETase in an two-enzyme system to increase the efficiency of degradation significantly. The deconstruction of recalcitrant polymers is done in nature by synergistic enzyme cocktails. There is a recent discovery on a type of two-enzyme system for the structure of polyethylene terephthalate(PET). This system used a kind of enzyme to transform the polymer to a soluble intermediate and another kind of enzyme for constituent PET monomer production. This discovery suggests that nature may start to involve with the deconstruction of synthetic plastics. The evolution of microbes is with the ability to use synthetic polymers as sources of carbon and energy. Ideonella sakaiensis was found recently to secrete a two-enzyme system. The sakaiensis PETase depolymerizes PET, and release soluble products, such as 2-hydroxyethy, MHET that was cleaved from the terephthalic acid and ethylene glycol by MHETase. 1.6 Å resolution MHETase structure was recorded, which means MHETase core domain is covered by a lid domain and has similarity with PETase. Simulations of catalytic itinerary predict the MHETase uses the typical two-step serine hydrolase mechanism. The Bioinformatics analysis said the MHETase is a result of the evolution of ferulic acid esterases, and the two homologous enzymes show the exhibit of MHET turnover. The result of two homologous enzymes and MHETase S131G also shows the residue is very important for the accommodation of MHET about the active site. MHETase lid is also important to hydrolysis of MHET, and MHETase does not turnover mono-furanoate of mono-isophalate. Also, all exhibit improved PET and MHET show turnover to free enzymes of the PETase chimeric proteins of linker lengths. These results suggested information for future work on the two-enzyme PET depolymerization, and can contribute to future deconstruction of plastics through biological ways.
References: Brandon C. Knott (13 Oct. 2020.). Characterization and engineering of a two-enzyme system for plastics depolymerization. PNAS. Retrieved from https://www.pnas.org/content/117/41/25476
Contribution
Group: Austin UTexas
Summary: PETase’s ability to depolymerize PET is limited by its low stability in environmental conditions. However, PETase has increased stability when displayed through a yeast cell-surface display system designed by Chen et al. This group found that PETase is most stable at a pH of 9 and a temperature of 40 °C. Below this temperature conformation changes lead to reduced stability. Crystallinity was also found to affect PETase’s function in plastic-degradation. PETase is less effective at degrading samples of PET with more crystallized molecular structures. These results also indicated that protein concentration has a great influence on the turnover rate of PETase. Increasing protein concentration over a threshold causes enzyme oversaturation, since not all PETase molecules can react with a given sample of PET at once.1
References: Chen, Z., Wang, Y., Cheng, Y., Wang, X., Tong, S., Yang, H., & Wang, Z. (2020). Efficient biodegradation of highly crystallized polyethylene terephthalate through cell surface display of bacterial PETase. Science Of The Total Environment, 709, 136138. doi: 10.1016/j.scitotenv.2019.136138
DUT_China 2021 improvement
Compared to the old part BBa_K2010999, expressing PETase, we design a new part BBa_K3898152, which is expressing RIDD-PETase. PETase in our research was optimized, and it shows better enzyme activity and considerable degradation effect.
Based on the results, we found that RIDD-PETase have esterase activity. After 20 minutes of reaction, each resulted in absorption peaks as high as 3.4873 at 405nm. Our optimized RIDD-PETase enzyme activity is 1.5 times higher than theirs in BBa_K2010999.
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NotI site found at 67
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]