Difference between revisions of "Part:BBa K2753002"
Cassandrashi (Talk | contribs) |
Cassandrashi (Talk | contribs) |
||
| (4 intermediate revisions by the same user not shown) | |||
| Line 91: | Line 91: | ||
<p> In the case of pTac, it fits into the scenario when the gene expression is lower than the optimum: higher copy produced more enzymes to catalyze geraniol synthesis reaction. As for TALE stabilized promoters, the expression level of enzymes would remain unchanged regardless of copy number due to the stabilization nature of TALE (More about TALE: [http://2018.igem.org/Team:GreatBay_China/Part_Collection]). So in low copy vector where the cellular burden isn’t significant, the product yield is only related to the strength of the promoter, explaining why TALE sp2 has better performance than TALE sp1 as it’s closer to the optimum. But when the copy number increases, the expression level is unaffected while the cellular burden rises sharply because much more TALE would be made to stabilize expression, leaving less energy available for geraniol synthesis. And it in turns answers why a negative trend of production is shown when regulated by TALE.</p><br> | <p> In the case of pTac, it fits into the scenario when the gene expression is lower than the optimum: higher copy produced more enzymes to catalyze geraniol synthesis reaction. As for TALE stabilized promoters, the expression level of enzymes would remain unchanged regardless of copy number due to the stabilization nature of TALE (More about TALE: [http://2018.igem.org/Team:GreatBay_China/Part_Collection]). So in low copy vector where the cellular burden isn’t significant, the product yield is only related to the strength of the promoter, explaining why TALE sp2 has better performance than TALE sp1 as it’s closer to the optimum. But when the copy number increases, the expression level is unaffected while the cellular burden rises sharply because much more TALE would be made to stabilize expression, leaving less energy available for geraniol synthesis. And it in turns answers why a negative trend of production is shown when regulated by TALE.</p><br> | ||
| + | ===Usage and Biology: Characterization=== | ||
| + | <span style="font-size: 130%;font-weight:bold">Group: KEYTSONE 2020 </span> | ||
| + | <span style="font-size: 130%;font-weight:bold">Summary: We used this part in the composite part pR6K-patc-GPPS-bLIS, and were able to produce Linalool from GPP with bLIS.</span> | ||
| − | |||
| − | |||
<span style="font-size: 130%;font-weight:bold">Description</span> | <span style="font-size: 130%;font-weight:bold">Description</span> | ||
| − | <p>GPPS- | + | |
| + | <p>GPPS-bLIS is composed of two coding sequences, GPPS (https://parts.igem.org/Part:BBa_K2753002) and bLIS (https://parts.igem.org/Part:BBa_K3478890). Coding GPPS is used to produce Geranyl pyrophosphate synthase that produce GPP from DMAPP, which is a substrate for linalool synthase that produces linalool. in our experiment, we constructed pR6K-ptac-GPPS-bLIS to produce linalool. The vector of the part is pR6K, with the promoter ptac, and composite part GPPS-bLIS. We first amplified the gene fragments using PCR (figure 1A, B, C and D) , and purified them to get pure gene fragments for pR6K-ptac-GPPS and bLIS. </p > | ||
https://2020.igem.org/wiki/images/7/76/T--KEYSTONE--mva1.png | https://2020.igem.org/wiki/images/7/76/T--KEYSTONE--mva1.png | ||
| Line 102: | Line 104: | ||
https://2020.igem.org/wiki/images/c/c3/T--KEYSTONE--gpps4.png | https://2020.igem.org/wiki/images/c/c3/T--KEYSTONE--gpps4.png | ||
| − | <p>Figure figure 1: the gel DNA gel electrophoresis results. (A) amplification result of gene segments of p15A-MVA-ptac-GPPS-bLIS (BLIS-1 (1015 bp), MVA-vector (6372 bp), MVA-GPPS (6970 bp)), pR6K-ptac-GPPS- | + | <p>Figure figure 1: the gel DNA gel electrophoresis results. (A) amplification result of gene segments of p15A-MVA-ptac-GPPS-bLIS (BLIS-1 (1015 bp), MVA-vector (6372 bp), MVA-GPPS (6970 bp)), pR6K-ptac-GPPS-bLIS (BLIS-2 (1016 bp), pR6K (5958 bp)) , pSB1K3-ptac-GPPS-bLIS (BLIS-3 (1016 bp), Lac1-Ptac-GPPS (2371 bp), PSBIC3 (2111 bp)) , p15A-MVA (MVA-1 (5177 bp), MVA-2 (6922 bp)). (B) result of gene segment purification. BLIS-1 (1015 bp), MVA-vector (6372 bp), MVA-GPPS (6970 bp), BLIS-2 (1016 bp), BLIS-3 (1016 bp), Lac1-Ptac-GPPS (2371 bp), (MVA-1 (5177 bp), MVA-2 (6922 bp). (C) amplification result of pR6K (5958 bp). (D) purification result of pR6K (5958 bp). </p > |
| − | <p>Gibson assembly was used to create plasmid pR6K-patc-GPPS- | + | <p>Gibson assembly was used to create plasmid pR6K-patc-GPPS-bLIS. We transformed the pR6K-patc-GPPS-bLIS to E. Coli DH5 α. The success of transformation is verified by a colony PCR. And induced it with IPTG, which allows it to produce linalool synthase (figure.2) </p > |
https://2020.igem.org/wiki/images/c/cc/T--KEYSTONE--gpps5.png | https://2020.igem.org/wiki/images/c/cc/T--KEYSTONE--gpps5.png | ||
| Line 110: | Line 112: | ||
| − | <p>Figure 2: (A) gel electrophoresis result of pR6K-ptac-GPPS- | + | <p>Figure 2: (A) gel electrophoresis result of pR6K-ptac-GPPS-bLIS (14358 bp). |
(B) protein electrophoresis results of linalool synthase (65.6 kDa) | (B) protein electrophoresis results of linalool synthase (65.6 kDa) | ||
| − | However, pR6K-patc-GPPS- | + | However, pR6K-patc-GPPS-bLIS alone in the E. Coli can’t produce linalool, because the substrate of GPP synthase, DMPP, is produced by MVA pathway, so we also constructed a plasmid p15A-MVA (figure 3), which allows the bacteria to have MVA pathway in it and produce DMPP, so that linalool synthesize can be achieved. </p > |
https://2020.igem.org/wiki/images/d/de/T--KEYSTONE--gpps7.png | https://2020.igem.org/wiki/images/d/de/T--KEYSTONE--gpps7.png | ||
<p>Figure 3: gene sequencing result of p15A-MVA. </p > | <p>Figure 3: gene sequencing result of p15A-MVA. </p > | ||
| − | <p>We transformed the two plasmid in We cotransformed P15A-MVA and pR6K-ptac-GPPS- | + | <p>We transformed the two plasmid in We cotransformed P15A-MVA and pR6K-ptac-GPPS-bLIS plasmid to E.coli DH5α. we induced them with IPTG and two of them with extra glucose. after the inducing process, the bacteria solution was treated with n-hexane to extract purified linalool(figure 4). We can clearly smell the strong fragrance of linalool in those samples, and samples with glucose have a stronger fragrance than samples that are not treated with glucose. This indicates that glucose may promote the synthesis of linalool in E.coli DH5α. We do not need to knock out the gene in E.coli producing stinky smell because the aroma overcomes the smell of E.coli. Therefore, our E.coli won’t affect the local environment and appearance. </p > |
<p>Besides, we also compared the smell of our sample to standard linalool sample, and standard geraniol, which is the product of pR6K-ptac-GPPS-GES (the plasmid before our improvement) Based on our observation the aroma of geraniol contains sweetness; whereas, the aroma of standard linalool sample has a slight peppery smell. The two smells are quite distinguishable. The small of our samples also have a slightly peppery aroma, which is close to the linalool standard sample. Thus, we have successfully produced linalool. | <p>Besides, we also compared the smell of our sample to standard linalool sample, and standard geraniol, which is the product of pR6K-ptac-GPPS-GES (the plasmid before our improvement) Based on our observation the aroma of geraniol contains sweetness; whereas, the aroma of standard linalool sample has a slight peppery smell. The two smells are quite distinguishable. The small of our samples also have a slightly peppery aroma, which is close to the linalool standard sample. Thus, we have successfully produced linalool. | ||
In addition, through visual observation, we also confirmed that glucose can promote the synthesis of linalool in E.coli. The transparent liquids in the test tubes are purified linalool (figure 4). The volume of the liquid in samples treated with glucose is larger than samples that aren’t treated by glucose. Therefore, E.coli treated with glucose may be able to produce more linalool in the same condition thanE.coli that is not. Our next step may be finding the optimal condition for linalool synthesis in E.coli. However, due to the Covid-19 situation, we do not have enough time in the lab, so we can only do those experiments in the future. </p > | In addition, through visual observation, we also confirmed that glucose can promote the synthesis of linalool in E.coli. The transparent liquids in the test tubes are purified linalool (figure 4). The volume of the liquid in samples treated with glucose is larger than samples that aren’t treated by glucose. Therefore, E.coli treated with glucose may be able to produce more linalool in the same condition thanE.coli that is not. Our next step may be finding the optimal condition for linalool synthesis in E.coli. However, due to the Covid-19 situation, we do not have enough time in the lab, so we can only do those experiments in the future. </p > | ||
https://2020.igem.org/wiki/images/3/3a/T--KEYSTONE--mva4.png | https://2020.igem.org/wiki/images/3/3a/T--KEYSTONE--mva4.png | ||
| − | <p>Figure 4: E.coli DH5α with P15A-MVA and pR6K-ptac-GPPS- | + | <p>Figure 4: E.coli DH5α with P15A-MVA and pR6K-ptac-GPPS-bLIS was propagated in four separated conical flasks. After the bacteria solutions reach the suitable OD (0.6-0.8), we added 2.5 ml glucose to two of the mediums during and added 25mM of IPTG to By adding n-hexane (4.5ml) to the induced bacteria solution, we can extract pure linalool (transparent layer) </p > |
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
<partinfo>BBa_K2753002 SequenceAndFeatures</partinfo> | <partinfo>BBa_K2753002 SequenceAndFeatures</partinfo> | ||
Latest revision as of 03:24, 28 October 2020
pSB1C3-GPPS cds
GPPS: Geranyl diphosphate (GPP), the entry point to the formation of terpene moiety, is a product of the condensation of isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP) by GPP synthase (GPPS)[1]. IPP and DMAPP are supplied by either MVA pathway or MEP pathway or both depending on species. This parts originates from the species Abies Grandis, and it’s codon-optimized for the use in E. coli.

Characterization

Figure. 1 The biosynthetic route for obtaining geraniol from acetyl-coA, which requires either a MVA or MEP pathway, a GPPS, and a GES.
In our study, we aim to achieve geraniol synthesis in E. coli, so at the beginning, we expressed a geraniol synthesis operon (https://parts.igem.org/Parts:BBa_K2753016) containing an Abies Grandis geranyl pyrophosphate synthase (https://parts.igem.org/Parts:BBa_K2753002) and an Ocimum basilicum geraniol synthase (https://parts.igem.org/Parts:BBa_K2753003) regulated by a pTac promoter in E. coli. Assuming that higher expression leads to higher yield, the operon was assembled on a high copy vector pUC20 whose copy number is approximately 500 per cell. However, when carrying out shake-flask fermentation with this strain, after induced by 25uM IPTG for 24 hours2, no geraniol production was detected using gas chromatography. (see detailed methods in our notebook: http://2018.igem.org/Team:GreatBay_China/Notebook)
We infer that the failure to produce geraniol was due to the lack of substrates IPP and DMAPP. Thus we decided to co-express an MVA pathway in addition to the geraniol synthesis operon. The pathway can either be added by cotransformation of a plasmid containing only the MVA pathway and the other plasmid with only GPPS and GES, or by constructing an all-in-one plasmid containing the MVA pathway, GPPS, and GES. We obtained a plasmid pMVA-only which was a gift from Taek Soon Lee, and have added GPPS and GES onto pMVA-only to acquire a pMVA-GPPS-GES.

Figure. 2 Geraniol production analysis by gas chromatography. (A) GPPS and GES are arranged in operon regulated by pTac, placed on a high copy vector pUC20. (B) The MVA pathway is split into two clusters and placed on a low copy vector with the upper cluster containing three genes and the downstream containing four. (C) A combined plasmid with GPPS&GES operon and MVA pathway with a low copy p15A origin. (D) Gas chromatography for geraniol produced by E. coli expressing pMVA only (middle trace) or pMVA-GPPS-GES (bottom trace) whose peaks coincide with a geraniol standard (top trace). (E) Geraniol yield is noticeable with only the heterologous MVA pathway. Upon introduction of GPPS and GES, the yield doubled relative to the negative control pMVA only.
Under the same fermentation condition, pMVA-GPPS-GES gave a geraniol titer of 11.4mg/L while the negative control pMVA-only as well produced geraniol titer of about 5.65mg/L. Then we are assured that a stable supply of precursors IPP and DMAPP are crucial for geraniol production. But still, there is room for higher geraniol yield. Therefore, to gain maximum flux to geraniol, we tuned the expression level of GPPS and GES using three promoters:
- TALE sp1 (https://parts.igem.org/Part:BBa_K2753030), copy number-independent promoter of medium strength
- TALE sp2 (https://parts.igem.org/Part:BBa_K2753018), copy number-independent promoter of low strength
- pTac (https://parts.igem.org/Part:BBa_K180000), IPTG inducible promoter
And three vectors of unique copy number:
- pUC20: ~500
- pR6K: ~15
- pSC101: ~1

Figure. 3 Geraniol production assay of TALEsp relative to pTac IPTG-inducible promoter. (A) Twelve combinations of promoters and plasmid backbones used to express a geraniol synthesis operon for characterization. pSC101 is the low copy vector, pR6K is the medium copy vector and pUC20 is the high copy vector. The combinations are transformed and tested in E. coli already harboring pMVA plasmids. (B) Geraniol yield achieved under the regulation of pTac, TALE1 sp1, TALE2 sp1, and TALE2 sp6 across different copy numbers is shown. The vertical error bars represents standard deviations of geraniol yield measured, and the horizontal error bars signify the reported range of copy number of plasmid backbones used. The portion of graph belong to TALE2sp6 is not shown as there is no detectable geraniol production.
Having transferred the constructs above along with pMVA-only into strain E. coli DH5α, the same shake-flask fermentation experiment was conducted. The result shows detectable geraniol production by each strain.
With pTac promoters, geraniol yield increased with the copy number of the vector, showing positively correlated relation. However, when TALE stabilized promoter (TALEsp) was used, higher the copy number of the vector, lower the production of geraniol, being the opposite of pTac. And the stronger TALEsp promoter, TALE1sp1 gave generally reduced yield compared to its weaker counterpart TALE2sp1. Interestingly, the graphs of the two TALEsp promoters appeared to be seemingly parallel to each other.

Figure. 4 Gas chromatography analysis for geraniol produced regulated by pTac and TALE2sp1 compared with the peak of geraniol standard. (A) Geraniol production accomplished by pTac regulation across low (pSC101), medium (pR6K) and high (pUC20) copy number. The area of the peak which appears at around 4.41s concurring with that of the geraniol standard increases along the copy number. (B) Geraniol yield assay using TALE2sp1 with the same set of copy number. The largest peak of geraniol emerges on the low copy vector pSC101. The peak shrinks in size on higher copy.
We surmised that the yield of geraniol was affected by two factors: the expression level of enzymes and the cellular burden. As for enzyme expression, there exists an optimal gene expression level that produces just enough enzyme to metabolize all the substrate. The yield would be the greatest at this level. But if lower than this level, production would increase with enzyme expression since there is a surplus of substrates. And if the enzyme expression is higher, the more enzymes now becomes a cellular burden as it no longer contributes to more product.
In the case of pTac, it fits into the scenario when the gene expression is lower than the optimum: higher copy produced more enzymes to catalyze geraniol synthesis reaction. As for TALE stabilized promoters, the expression level of enzymes would remain unchanged regardless of copy number due to the stabilization nature of TALE (More about TALE: [http://2018.igem.org/Team:GreatBay_China/Part_Collection]). So in low copy vector where the cellular burden isn’t significant, the product yield is only related to the strength of the promoter, explaining why TALE sp2 has better performance than TALE sp1 as it’s closer to the optimum. But when the copy number increases, the expression level is unaffected while the cellular burden rises sharply because much more TALE would be made to stabilize expression, leaving less energy available for geraniol synthesis. And it in turns answers why a negative trend of production is shown when regulated by TALE.
Usage and Biology: Characterization
Group: KEYTSONE 2020 Summary: We used this part in the composite part pR6K-patc-GPPS-bLIS, and were able to produce Linalool from GPP with bLIS.
Description
GPPS-bLIS is composed of two coding sequences, GPPS (https://parts.igem.org/Part:BBa_K2753002) and bLIS (https://parts.igem.org/Part:BBa_K3478890). Coding GPPS is used to produce Geranyl pyrophosphate synthase that produce GPP from DMAPP, which is a substrate for linalool synthase that produces linalool. in our experiment, we constructed pR6K-ptac-GPPS-bLIS to produce linalool. The vector of the part is pR6K, with the promoter ptac, and composite part GPPS-bLIS. We first amplified the gene fragments using PCR (figure 1A, B, C and D) , and purified them to get pure gene fragments for pR6K-ptac-GPPS and bLIS.
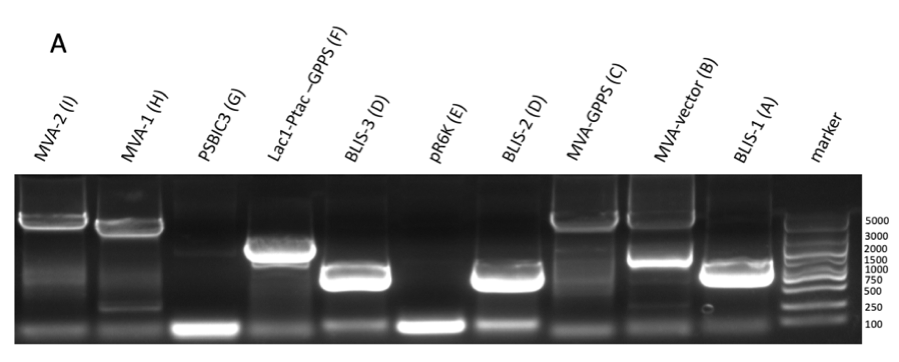
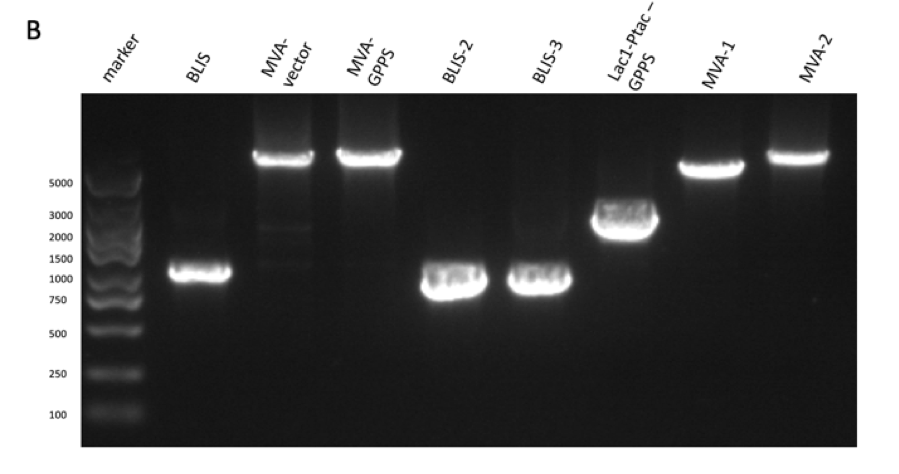
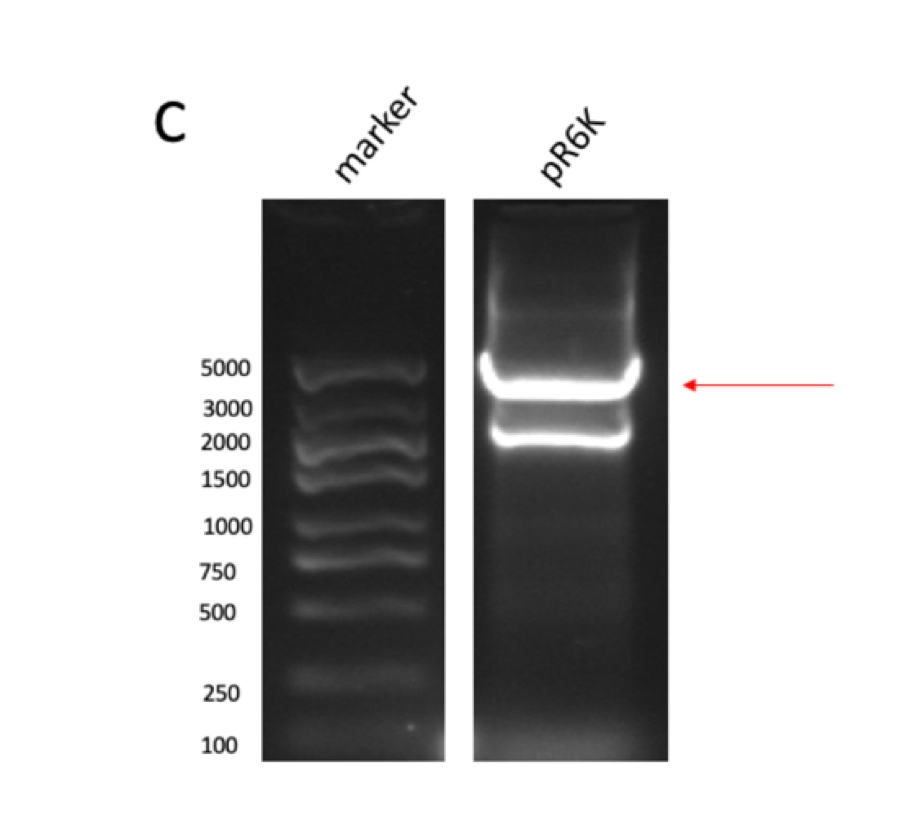
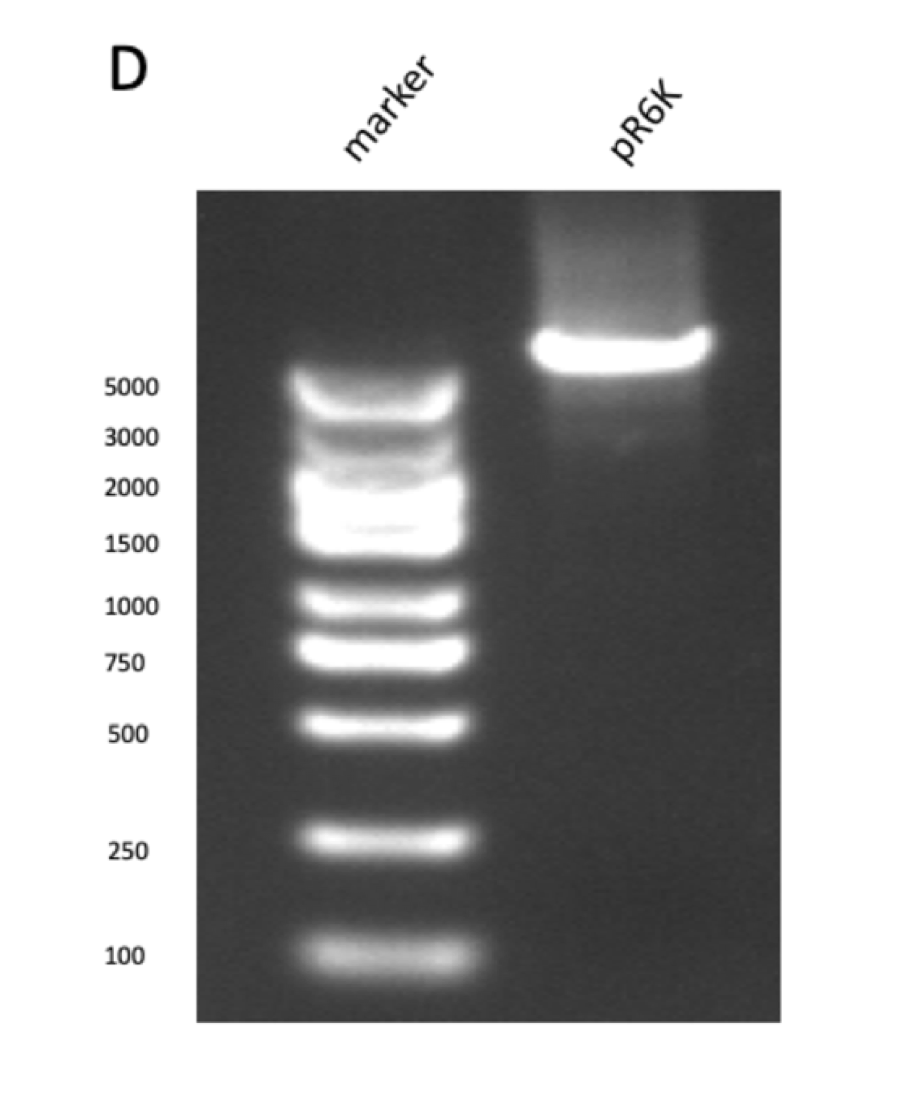
Figure figure 1: the gel DNA gel electrophoresis results. (A) amplification result of gene segments of p15A-MVA-ptac-GPPS-bLIS (BLIS-1 (1015 bp), MVA-vector (6372 bp), MVA-GPPS (6970 bp)), pR6K-ptac-GPPS-bLIS (BLIS-2 (1016 bp), pR6K (5958 bp)) , pSB1K3-ptac-GPPS-bLIS (BLIS-3 (1016 bp), Lac1-Ptac-GPPS (2371 bp), PSBIC3 (2111 bp)) , p15A-MVA (MVA-1 (5177 bp), MVA-2 (6922 bp)). (B) result of gene segment purification. BLIS-1 (1015 bp), MVA-vector (6372 bp), MVA-GPPS (6970 bp), BLIS-2 (1016 bp), BLIS-3 (1016 bp), Lac1-Ptac-GPPS (2371 bp), (MVA-1 (5177 bp), MVA-2 (6922 bp). (C) amplification result of pR6K (5958 bp). (D) purification result of pR6K (5958 bp).
Gibson assembly was used to create plasmid pR6K-patc-GPPS-bLIS. We transformed the pR6K-patc-GPPS-bLIS to E. Coli DH5 α. The success of transformation is verified by a colony PCR. And induced it with IPTG, which allows it to produce linalool synthase (figure.2)
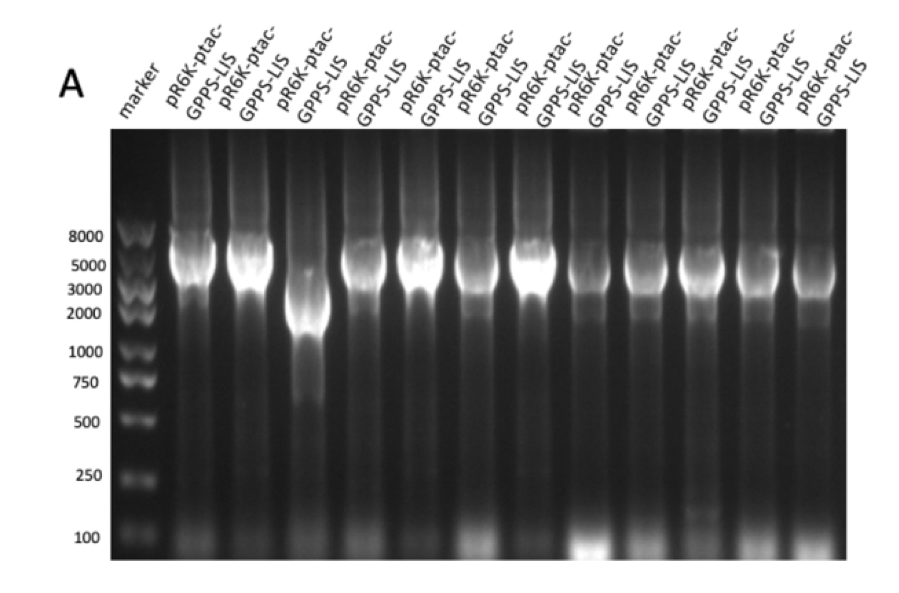
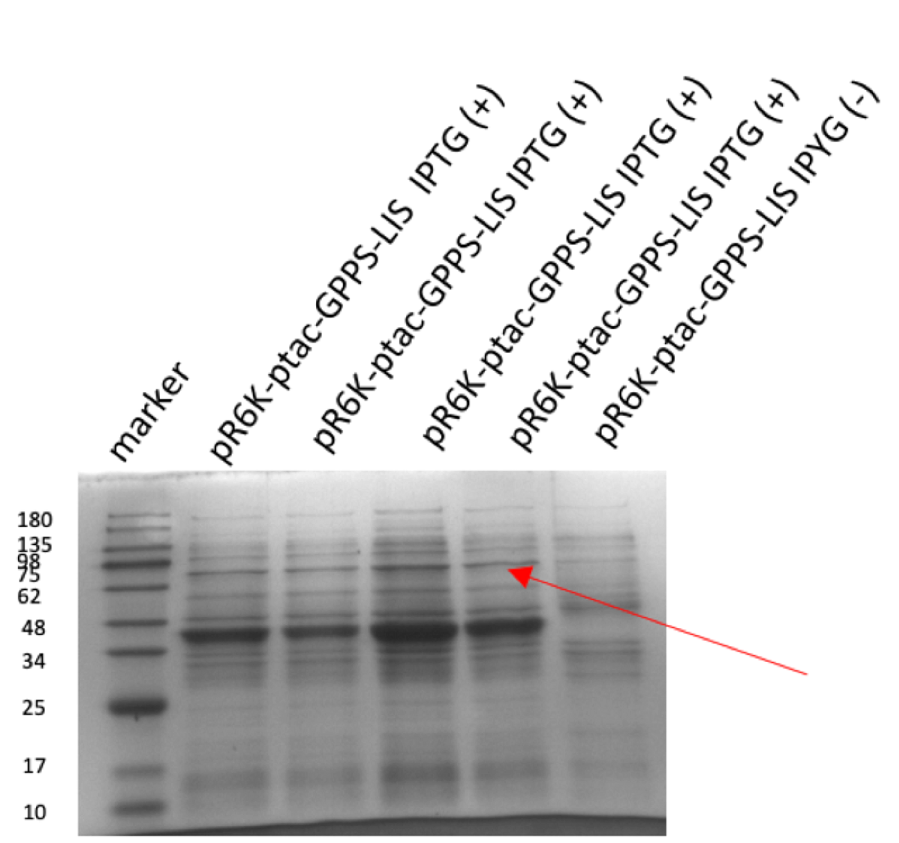
Figure 2: (A) gel electrophoresis result of pR6K-ptac-GPPS-bLIS (14358 bp). (B) protein electrophoresis results of linalool synthase (65.6 kDa) However, pR6K-patc-GPPS-bLIS alone in the E. Coli can’t produce linalool, because the substrate of GPP synthase, DMPP, is produced by MVA pathway, so we also constructed a plasmid p15A-MVA (figure 3), which allows the bacteria to have MVA pathway in it and produce DMPP, so that linalool synthesize can be achieved.
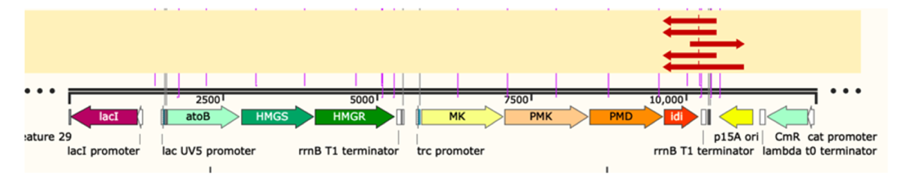
Figure 3: gene sequencing result of p15A-MVA.
We transformed the two plasmid in We cotransformed P15A-MVA and pR6K-ptac-GPPS-bLIS plasmid to E.coli DH5α. we induced them with IPTG and two of them with extra glucose. after the inducing process, the bacteria solution was treated with n-hexane to extract purified linalool(figure 4). We can clearly smell the strong fragrance of linalool in those samples, and samples with glucose have a stronger fragrance than samples that are not treated with glucose. This indicates that glucose may promote the synthesis of linalool in E.coli DH5α. We do not need to knock out the gene in E.coli producing stinky smell because the aroma overcomes the smell of E.coli. Therefore, our E.coli won’t affect the local environment and appearance.
Besides, we also compared the smell of our sample to standard linalool sample, and standard geraniol, which is the product of pR6K-ptac-GPPS-GES (the plasmid before our improvement) Based on our observation the aroma of geraniol contains sweetness; whereas, the aroma of standard linalool sample has a slight peppery smell. The two smells are quite distinguishable. The small of our samples also have a slightly peppery aroma, which is close to the linalool standard sample. Thus, we have successfully produced linalool. In addition, through visual observation, we also confirmed that glucose can promote the synthesis of linalool in E.coli. The transparent liquids in the test tubes are purified linalool (figure 4). The volume of the liquid in samples treated with glucose is larger than samples that aren’t treated by glucose. Therefore, E.coli treated with glucose may be able to produce more linalool in the same condition thanE.coli that is not. Our next step may be finding the optimal condition for linalool synthesis in E.coli. However, due to the Covid-19 situation, we do not have enough time in the lab, so we can only do those experiments in the future.
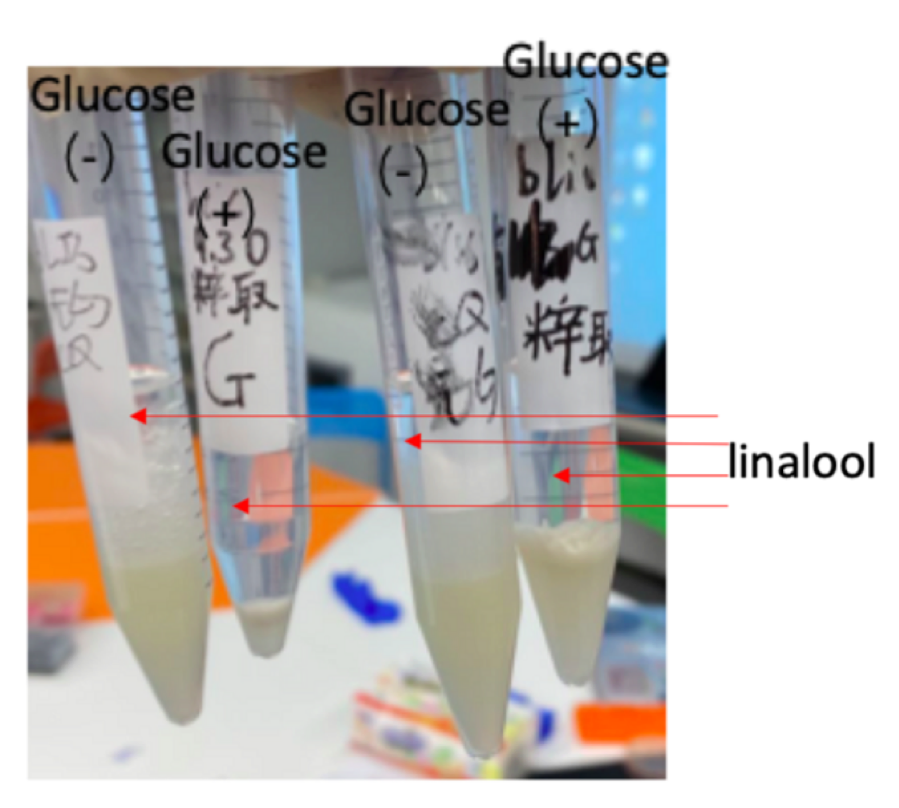
Figure 4: E.coli DH5α with P15A-MVA and pR6K-ptac-GPPS-bLIS was propagated in four separated conical flasks. After the bacteria solutions reach the suitable OD (0.6-0.8), we added 2.5 ml glucose to two of the mediums during and added 25mM of IPTG to By adding n-hexane (4.5ml) to the induced bacteria solution, we can extract pure linalool (transparent layer)
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Referece
[1]: Rai, A., Smita, S. S., Singh, A. K., Shanker, K., & Nagegowda, D. A. (2013). Heteromeric and Homomeric Geranyl Diphosphate Synthases from Catharanthus roseus and Their Role in Monoterpene Indole Alkaloid Biosynthesis. Molecular Plant, 6(5), 1531–1549. doi:10.1093/mp/sst058
[2]: Zebec, Z., Wilkes, J., Jervis, A. J., Scrutton, N. S., Takano, E., & Breitling, R. (n.d.). Towards synthesis of monoterpenes and derivatives using synthetic biology. Current Opinion in Chemical Biology, 34, 37–43. https://doi.org/10.1016/j.cbpa.2016.06.002
