Difference between revisions of "Part:BBa K3332013"
AnnaTaylor (Talk | contribs) (→Usage) |
|||
| (3 intermediate revisions by 2 users not shown) | |||
| Line 3: | Line 3: | ||
<partinfo>BBa_K3332013 short</partinfo> | <partinfo>BBa_K3332013 short</partinfo> | ||
| − | We anchor GRHPR protein onto membranes through Lpp-OmpA to catalyze the reaction of reducing glyoxalic acid and consuming NADPH. We use | + | We anchor GRHPR protein onto membranes through Lpp-OmpA to catalyze the reaction of reducing glyoxalic acid and consuming NADPH. We use <partinfo>BBa_K880005</partinfo> to construct the expression system and anchor GRHPR on the surface of ''E.coli''. |
| + | |||
| + | |||
| + | ===Biology=== | ||
| + | |||
| + | Lpp-OmpA is an anchor protein from ''E. coli'', which can anchor its passenger protein to the cell membrane. It has been widely used in cell-surface display. GRHPR, a Glyoxylate reductase from human liver, can reduce glyoxylic acid when NADPH is used as cofactor. GRHPR is fused at N terminal with Lpp-OmpA so that GRHPR can be displayed on the surface of ''E. coli''.<ref>Rumsby G, Cregeen D P. Identification and expression of a cDNA for human hydroxypyruvate/glyoxylate reductase[J]. Biochimica et Biophysica Acta (BBA) - Gene Structure and Expression, 1999, 1446(3): 383-388.</ref><ref>http://2016.igem.org/Team:TJUSLS_China</ref> | ||
| + | <html> | ||
| + | <figure> | ||
| + | <img src="https://2020.igem.org/wiki/images/8/82/T--XMU-China--XMU-China_2020-GRHPR%E9%94%9A%E5%AE%9A.png" width="45%" style="float:center"> | ||
| + | <figcaption> | ||
| + | <p style="font-size:1rem"> | ||
| + | </p> | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </html> | ||
| + | |||
| + | :'''Fig 1.''' mechanism. | ||
| + | |||
| + | |||
| + | ===Usage=== | ||
| + | |||
| + | Here, we used <partinfo>BBa_K880005</partinfo> to construct the expression system and demonstrate the effect of Lpp-OmpA-GRHPR on the surface of ''E. coli''. We obtained the composite part <partinfo>BBa_K3332058</partinfo>. Due to the limited time, we didn’t get the gene in time. As a result, there is a lack of data about this part. The progress of this part remains in the stage of design. | ||
| + | |||
| + | <html> | ||
| + | <figure> | ||
| + | <img src="https://2020.igem.org/wiki/images/a/aa/T--XMU-China--XMU-China_2020-J23100_B0034_lpp-ompA-grhpr_B0015.png" width="35%" style="float:center"> | ||
| + | <figcaption> | ||
| + | <p style="font-size:1rem"> | ||
| + | </p> | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </html> | ||
| + | |||
| + | :'''Fig 2.''' Gene circuit of Lpp-OmpA-GRHPR. | ||
| + | |||
| + | ===References=== | ||
| + | <references/> | ||
| − | |||
| − | |||
<!-- --> | <!-- --> | ||
Latest revision as of 22:57, 27 October 2020
Lpp-OmpA-GRHPR
We anchor GRHPR protein onto membranes through Lpp-OmpA to catalyze the reaction of reducing glyoxalic acid and consuming NADPH. We use BBa_K880005 to construct the expression system and anchor GRHPR on the surface of E.coli.
Biology
Lpp-OmpA is an anchor protein from E. coli, which can anchor its passenger protein to the cell membrane. It has been widely used in cell-surface display. GRHPR, a Glyoxylate reductase from human liver, can reduce glyoxylic acid when NADPH is used as cofactor. GRHPR is fused at N terminal with Lpp-OmpA so that GRHPR can be displayed on the surface of E. coli.[1][2]
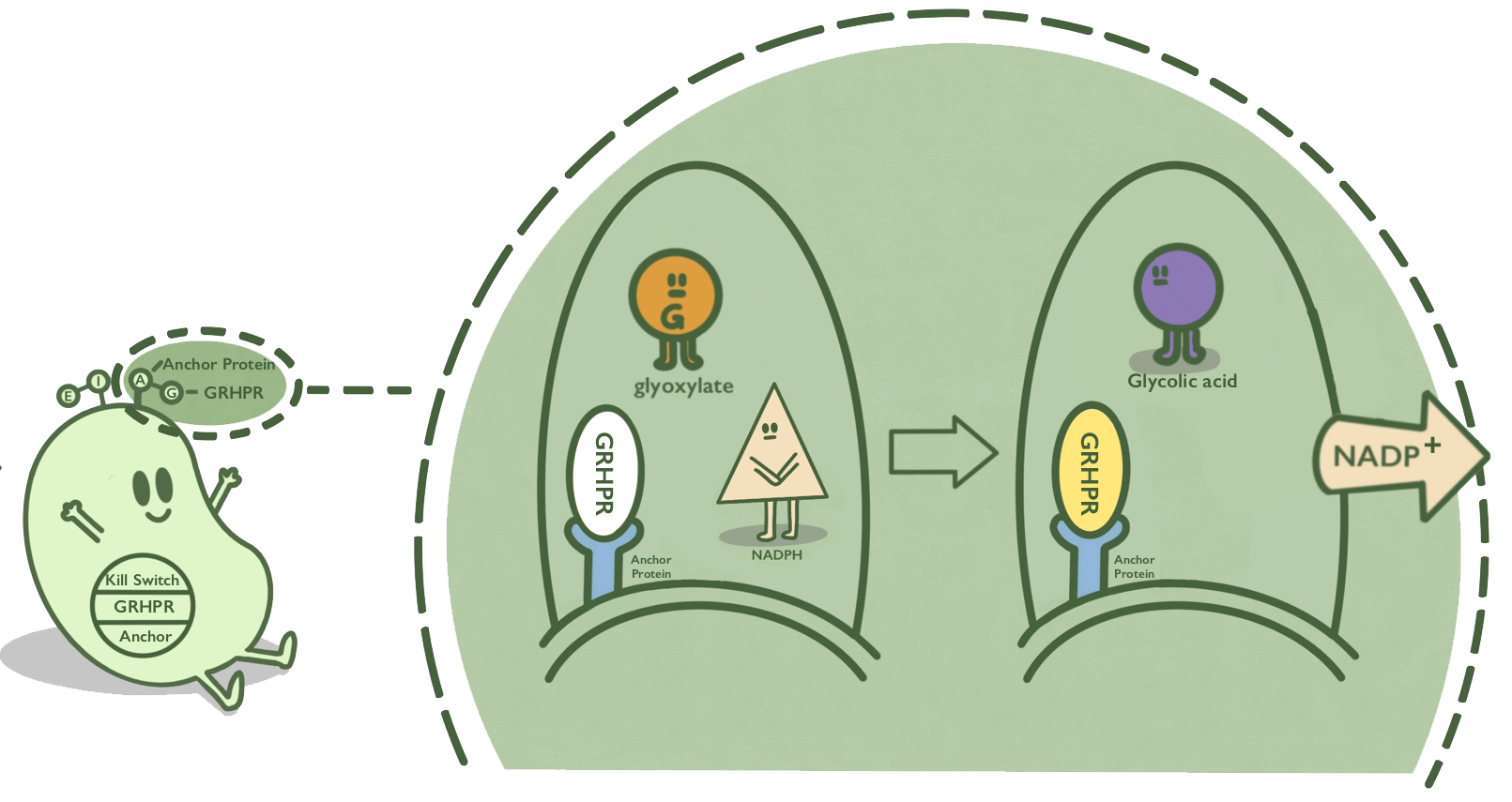
- Fig 1. mechanism.
Usage
Here, we used BBa_K880005 to construct the expression system and demonstrate the effect of Lpp-OmpA-GRHPR on the surface of E. coli. We obtained the composite part BBa_K3332058. Due to the limited time, we didn’t get the gene in time. As a result, there is a lack of data about this part. The progress of this part remains in the stage of design.

- Fig 2. Gene circuit of Lpp-OmpA-GRHPR.
References
- ↑ Rumsby G, Cregeen D P. Identification and expression of a cDNA for human hydroxypyruvate/glyoxylate reductase[J]. Biochimica et Biophysica Acta (BBA) - Gene Structure and Expression, 1999, 1446(3): 383-388.
- ↑ http://2016.igem.org/Team:TJUSLS_China
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 1012
- 1000COMPATIBLE WITH RFC[1000]
