Difference between revisions of "Part:BBa K3453055"
| (3 intermediate revisions by the same user not shown) | |||
| Line 3: | Line 3: | ||
<partinfo>BBa_K3453055 short</partinfo> | <partinfo>BBa_K3453055 short</partinfo> | ||
| − | + | This part is a toehold switch sensor for sequence-based detection of rosewood. It targets a fragment of the TrnL-UAA gene of ''Dalbergia maritima'' var. ''pubescens'' ([[Part:BBa_K3453060|BBa_K3453060]]). | |
| − | |||
===Usage and Biology=== | ===Usage and Biology=== | ||
| + | |||
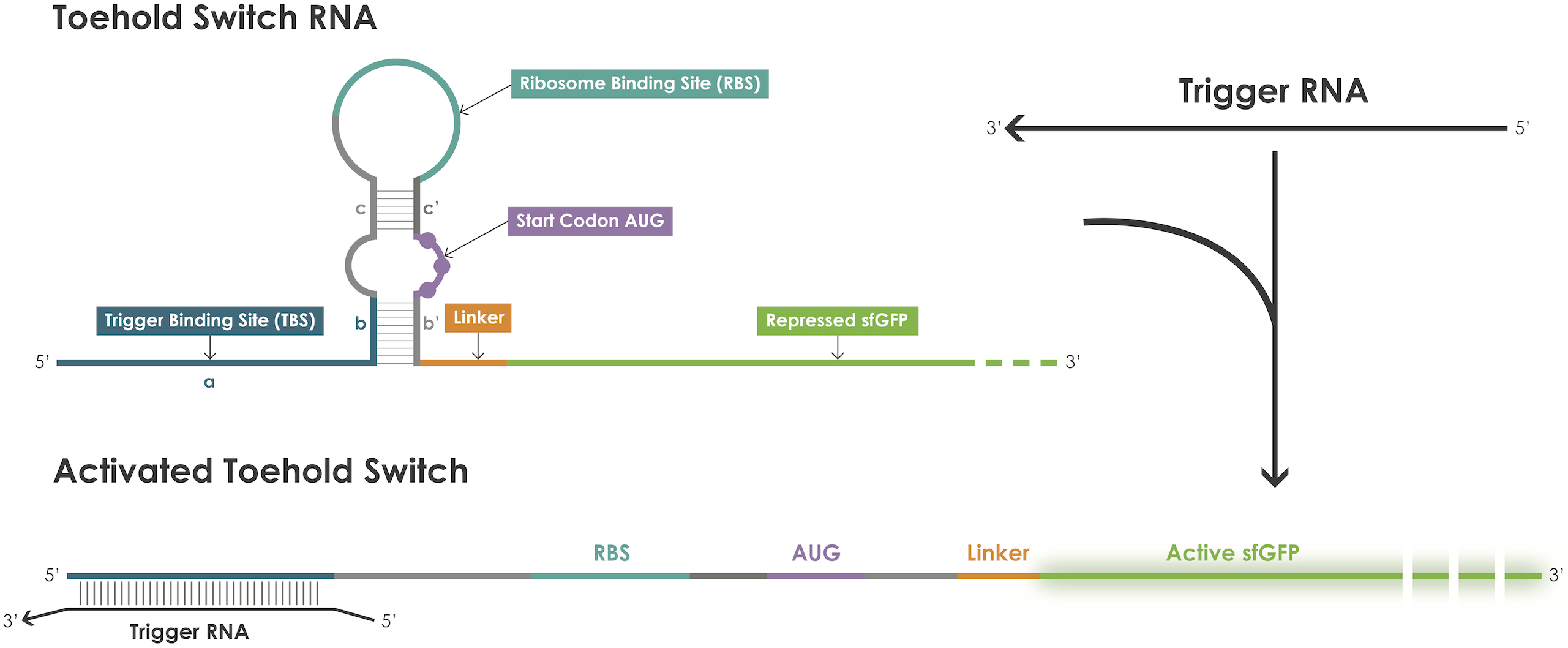
| + | A toehold switch is an RNA–based device containing a ribosome binding site (RBS) and an ATG start codon embedded in the middle of a hairpin structure that blocks translation initiation [1]. The hairpin can be unfolded upon binding of a trigger RNA thereby exposing the RBS and the ATG start codon and thus permitting translation of the reporter protein (Figure 1). | ||
| + | |||
| + | |||
| + | [[File: T--Evry_Paris-Saclay--toehold-switch.png|600px]] | ||
| + | |||
| + | Figure 1. Toehold switches principle. | ||
| + | |||
| + | This part is a toehold switch sensor that targets a fragment of the TrnL-UAA gene of ''Dalbergia maritima'' var. ''pubescens'' ([[Part:BBa_K3453060|BBa_K3453060]]). | ||
| + | It was designed using our internal toehold switch pipeline which calls NUPACK [2] and RBS Calculator for switch prediction and ranking [3]. Its secondary structure predicted by NUPACK web server [2] using default parameters is represented in Figure 2 and here-after in dot-bracket notation. | ||
| + | |||
| + | .....(((...))).((((((((((((((((((...............))))))))))))))))))..((....))...((((.....)))).... | ||
| + | |||
| + | [[File: T--Evry_Paris-Saclay--RNAstructure_p74_DmTrnL-UAA2-2.png|center]] | ||
| + | |||
| + | Figure 2. Secondary-structure prediction of this part with the ATG of the reporter gene. The prediction was realised using the NUPACK web server [2] with default parameters and graphically represented using the ''forna'' RNA secondary structure visualization tool [4]. Nucleotides were coloured to match the different segments in Figure 1. | ||
| + | |||
| + | |||
| + | The corresponding trigger sequence of this toehold switch is [[Part:BBa_K3453065|BBa_K3453065]]. | ||
| + | |||
| + | The functionality of this part was tested using sfGFP-LVAtag ([[Part:BBa_K2675006|BBa_K2675006]]) as a reporter. The expression was controlled by the T7 promoter ([[Part:BBa_K2150031|BBa_K2150031]]) and the strong SBa_000587 synthetic terminator ([[Part:BBa_K3453000|BBa_K3453000]]) in the composite part [[Part:BBa_K3453155|BBa_K3453155]]. | ||
| + | |||
| + | This part proved to be functional: a readable output was generated only in the presence of rosewood RNA. | ||
| + | |||
| + | Full results are available on the [[Part:BBa_K3453155|BBa_K3453155]] page in the registry. | ||
| + | |||
| + | ===References=== | ||
| + | [1] Green AA, Silver PA, Collins JJ, Yin P. Toehold switches: de-novo-designed regulators of gene expression. Cell (2014) 159, 925-939. | ||
| + | |||
| + | [2] Zadeh JN, Steenberg CD, Bois JS, Wolfe BR, Pierce MB, Khan AR, Dirks RM, Pierce NA. NUPACK: Analysis and design of nucleic acid systems. Journal of Computational Chemistry (2011) 32, 170–173. | ||
| + | |||
| + | [3] Salis HM. The ribosome binding site calculator. Methods in Enzymology (2011) 498: 19–42. | ||
| + | |||
| + | [4] Kerpedjiev P, Hammer S, Hofacker IL. Forna (force-directed RNA): Simple and effective online RNA secondary structure diagrams. Bioinformatics (Oxford, England) (2015) 31, 3377–3379. | ||
| + | |||
<!-- --> | <!-- --> | ||
Latest revision as of 21:20, 26 October 2020
Rosewood DmTrnL-UAA Toehold Switch 2.2
This part is a toehold switch sensor for sequence-based detection of rosewood. It targets a fragment of the TrnL-UAA gene of Dalbergia maritima var. pubescens (BBa_K3453060).
Usage and Biology
A toehold switch is an RNA–based device containing a ribosome binding site (RBS) and an ATG start codon embedded in the middle of a hairpin structure that blocks translation initiation [1]. The hairpin can be unfolded upon binding of a trigger RNA thereby exposing the RBS and the ATG start codon and thus permitting translation of the reporter protein (Figure 1).
Figure 1. Toehold switches principle.
This part is a toehold switch sensor that targets a fragment of the TrnL-UAA gene of Dalbergia maritima var. pubescens (BBa_K3453060). It was designed using our internal toehold switch pipeline which calls NUPACK [2] and RBS Calculator for switch prediction and ranking [3]. Its secondary structure predicted by NUPACK web server [2] using default parameters is represented in Figure 2 and here-after in dot-bracket notation.
.....(((...))).((((((((((((((((((...............))))))))))))))))))..((....))...((((.....))))....
Figure 2. Secondary-structure prediction of this part with the ATG of the reporter gene. The prediction was realised using the NUPACK web server [2] with default parameters and graphically represented using the forna RNA secondary structure visualization tool [4]. Nucleotides were coloured to match the different segments in Figure 1.
The corresponding trigger sequence of this toehold switch is BBa_K3453065.
The functionality of this part was tested using sfGFP-LVAtag (BBa_K2675006) as a reporter. The expression was controlled by the T7 promoter (BBa_K2150031) and the strong SBa_000587 synthetic terminator (BBa_K3453000) in the composite part BBa_K3453155.
This part proved to be functional: a readable output was generated only in the presence of rosewood RNA.
Full results are available on the BBa_K3453155 page in the registry.
References
[1] Green AA, Silver PA, Collins JJ, Yin P. Toehold switches: de-novo-designed regulators of gene expression. Cell (2014) 159, 925-939.
[2] Zadeh JN, Steenberg CD, Bois JS, Wolfe BR, Pierce MB, Khan AR, Dirks RM, Pierce NA. NUPACK: Analysis and design of nucleic acid systems. Journal of Computational Chemistry (2011) 32, 170–173.
[3] Salis HM. The ribosome binding site calculator. Methods in Enzymology (2011) 498: 19–42.
[4] Kerpedjiev P, Hammer S, Hofacker IL. Forna (force-directed RNA): Simple and effective online RNA secondary structure diagrams. Bioinformatics (Oxford, England) (2015) 31, 3377–3379.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]