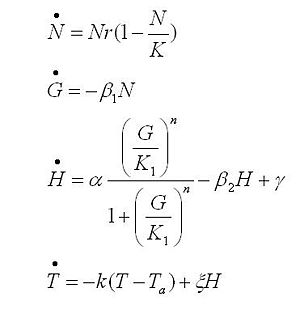Difference between revisions of "Part:BBa K141000"
| Line 64: | Line 64: | ||
| − | '''Common parameters values''' | + | '''Common parameters values:''' |
| − | [[Image:commonparametershar.jpg]] | + | [[Image:commonparametershar.jpg|450px]] |
Revision as of 20:31, 29 October 2008
Ucp1
Ucp1 is a gene which encodes for UCP1 protein.
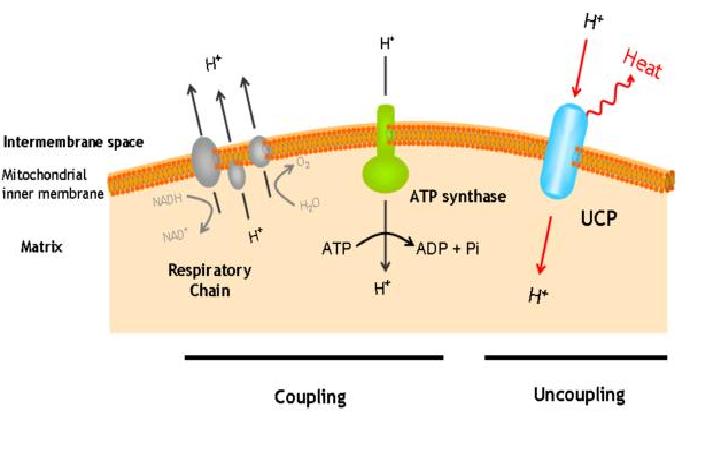
The uncoupling protein UCP1 is a proton carrier characteristic of brown adipose tissue. UCP1 uncouples the respiratory chain of ATP production, converting the metabolic energy in heat.
|
UCP1 is a 33kd protein which is exclusively located in the brown adipocites.
It is an integral protein present in inner mitocondrial membrane.
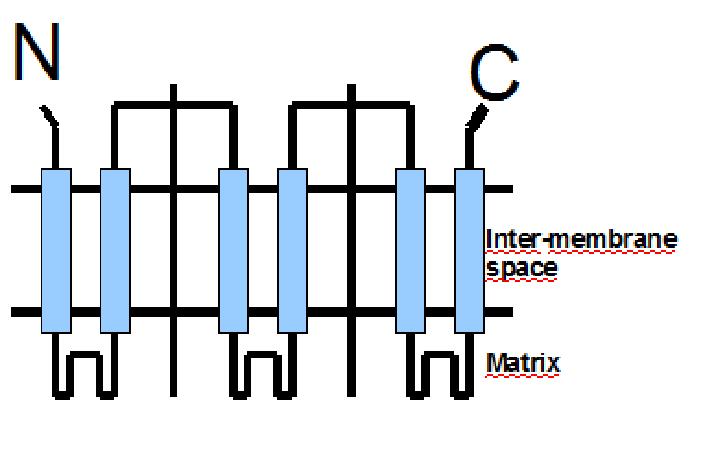
The protein has a tripartite structure. The structure displays an around 100 residues region which is three times repeated. Each part encodes for two transmembrane segments and one long hydrophilic loop.The functional carrier unit is an homodimer.
|
The main difference between UCP1 and most of the proteins with a nuclear codification is the lack of the importation targeting to the mitochondria in UCP+ proteins.
The condition that determines the mitochondria as the protein target lays in the first loop which protudes in the mitochondrial matrix.
The second loop of the matrix is essential for the insertion of the protein in the inner mitochondrial matrix. Purine nucleotides act as inhibitors of protein activity and esterificated fatty acids act as inductors.
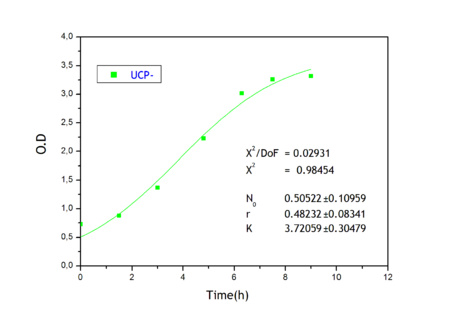
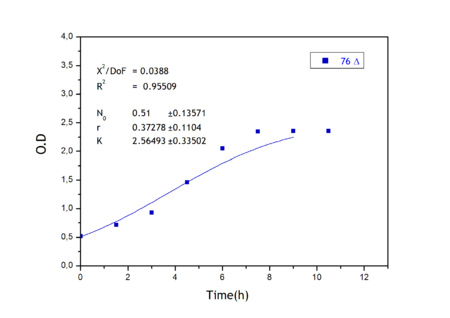
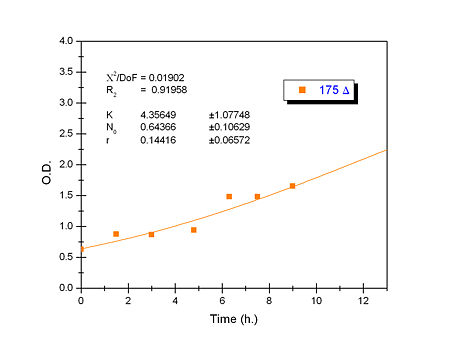
Apart from monitoring temperature evolution, we also characterized O.D. variations of each of our strains. We took O.D. measures every one a half hours for nine hours. We carried out this experiment both in Erlenmeyer flasks in the 30ºC shaking stove and in our LCCs. This measurements were useful in order to determine some parameters for our Black Box Model. Besides, we were able to prove that the UCP was indeed being produced even though we could not see the temperature increase. Since our mutant strains Gly175Δ and Gly76Δ do not have the same growing rate as UCP+ when they are expressing the protein, the difference between the results in the strains showed that the reason for our lack of temperature increase was that we had not found the optimum conditions yet. This results made us keep on working until we obtained successful results.
Strains growth equations:
The black box model simulates the temperature evolution of the system as a function of the growth rate, galactose concentration and thermogenin expresion. We assume that our system can be reproduced by the following system of equations:
where:
- first equation: Growth of the culture: logistic growth of the different mutants used in our experiments.
- second equation: Galactose evolution: Galactose is the metabolite which induces the thermogenin expresion.
- third equation: Thermogenin concentration level.
- Fourth equation: Temperature evolution. the first term of the equation represents the losses to the ambient of the calorimeter and the second one the temperature increase as a consequence of the thermogenin expresion.
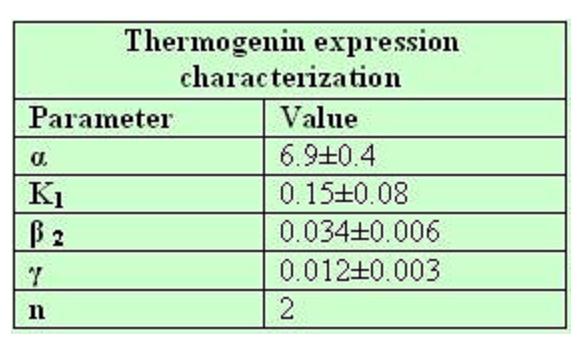
From a sample of 10 experiments in which the evolution of the temperature of the four strain was measured we perform a fit using the software simulink.
Common parameters values:
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 47
Illegal BglII site found at 347
Illegal BamHI site found at 835 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 58
- 1000COMPATIBLE WITH RFC[1000]