Difference between revisions of "Part:BBa I742106"
(→2019 NCKU-Tainan Part Improvement) |
(→NCKU_Tainan 2019 Improvement) |
||
| (13 intermediate revisions by 2 users not shown) | |||
| Line 17: | Line 17: | ||
<partinfo>BBa_I742106 parameters</partinfo> | <partinfo>BBa_I742106 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| − | = | + | =NCKU_Tainan 2019 Improvement= |
| − | + | ||
| + | Click [[Part:BBa_K2997011 | BBa_K2997011]] to view the part, which we improved and used for the expression. | ||
===Background=== | ===Background=== | ||
| − | Tyrosine ammonia-lyase ( | + | Tyrosine ammonia-lyase (TAL) is an enzyme that converts tyrosine into <i>p</i>-Coumaric acid. We engineered <i>E. coli</i> Nissle to express TAL to turn the excess tyrosine inside the gut into <i>p</i>-Coumaric acid, and compare the different constructs for improvement, [[Part:BBa_K2997009 | BBa_K2997009]] and [[Part:BBa_K2997010 | BBa_K2997010]]. |
| − | ===Expression in <i>E.coli</i>=== | + | ===Expression in <i>E. coli</i>=== |
| − | We constructed Tyrosine Ammonia Lyase with a native RBS , and transformed the plasmid into E. coli Nissle 1917 and confirmed it by double digestion. The results are as follows: | + | We constructed Tyrosine Ammonia Lyase with a native RBS , and transformed the plasmid into <i>E. coli</i> Nissle 1917 and confirmed it by double digestion. The results are as follows: |
<html> | <html> | ||
| Line 33: | Line 34: | ||
<br> | <br> | ||
</html> | </html> | ||
| − | Fig. 1. Confirmation of BBa_K2997009 by double digestion, arrow indicates TAL | + | Fig. 1. Confirmation of [[Part:BBa_K2997009 | BBa_K2997009]] by double digestion, arrow indicates TAL with NRBS (~1600bp).M: Marker; Lane 1: pSB1C3-[[Part:BBa_K2997009 | BBa_K2997009]]; Lane 2: [[Part:BBa_K2997009 | BBa_K2997009]] |
Then, we carried out SDS-PAGE to check the protein expression of TAL, and the expected protein size of TAL is 54kDa. As seen in the results below, however, there’s no distinguishable band around both sizes. | Then, we carried out SDS-PAGE to check the protein expression of TAL, and the expected protein size of TAL is 54kDa. As seen in the results below, however, there’s no distinguishable band around both sizes. | ||
| Line 44: | Line 45: | ||
<br> | <br> | ||
</html> | </html> | ||
| − | Fig. 2. 12% SDS PAGE of E. coli Nissle 1917 with different plasmids. M: Marker; Lane 1: Wild Type; Lane 2: pSB1C3; Lane 3: BBa_K2997009 ; Lane 4: BBa_K2997010; Lane 5: Dual plasmid containing BBa_K2997009 and BBa_K2997000; Lane 6: Dual plasmid containing BBa_K2997010 and BBa_K2997000; Lane 7: Positive control (c.d. 3392) | + | Fig. 2. 12% SDS PAGE of <i>E. coli</i> Nissle 1917 with different plasmids. M: Marker; Lane 1: Wild Type; Lane 2: pSB1C3; Lane 3: [[Part:BBa_K2997009 | BBa_K2997009]] ; Lane 4: [[Part:BBa_K2997010 | BBa_K2997010]]; Lane 5: Dual plasmid containing [[Part:BBa_K2997009 | BBa_K2997009]] and [[Part:BBa_K2997000 | BBa_K2997000]]; Lane 6: Dual plasmid containing [[Part:BBa_K2997010 | BBa_K2997010]] and [[Part:BBa_K2997000 | BBa_K2997000]]; Lane 7: Positive control (c.d. 3392) |
===RT-PCR=== | ===RT-PCR=== | ||
| − | RT-PCR experiment was used to confirm that the constructed TAL Biobrick is being transcribed. As seen in Fig.3, cDNA for both bacteria carrying TAL constructs are being detected by PCR. Confirming that the TAL genes is actually being transcribed in E. coli Nissle. | + | RT-PCR experiment was used to confirm that the constructed TAL Biobrick is being transcribed. As seen in Fig. 3, cDNA for both bacteria carrying TAL constructs are being detected by PCR. Confirming that the TAL genes is actually being transcribed in <i>E. coli</i> Nissle. |
<html> | <html> | ||
<br> | <br> | ||
| Line 69: | Line 70: | ||
===TyrP and TAL Assay=== | ===TyrP and TAL Assay=== | ||
| − | To confirm the protein activity of TAL and TyrP, we performed a functional test using n-octanol extraction method | + | |
| + | To confirm the protein activity of TAL and TyrP, we performed a functional test using <i>n</i>-octanol extraction method, [https://2019.igem.org/Team:NCKU_Tainan/Protocols TyrP and TAL Assay], which was previously proposed by iGEM Uppsala 2013 and has been verified by HPLC<sup>[1]</sup>. The <i>p</i>-Coumaric acid concentration was measured through the absorbance value at 310nm wavelength under Nanodrop UV-Vis wavelength. The standard curve of <i>p</i>-Coumaric acid was drawn in Fig.4 to determine the relationship between <i>p</i>-Coumaric acid concentrations and its 310nm arbitrary unit (a.u). Our samples with TAL constructs were then mapped onto the standard curve, to know how much <i>p</i>-Coumaric acid is being produced. | ||
| + | |||
<html> | <html> | ||
<br> | <br> | ||
| Line 77: | Line 80: | ||
<br> | <br> | ||
</html> | </html> | ||
| − | Fig. 4. The standard curve of p-Coumaric acid concentration in correlation with absorbance at 310nm, which is provided by genetic E. coli Nissle in LB broth after 48 hours. | + | Fig. 4. The standard curve of <i>p</i>-Coumaric acid concentration in correlation with absorbance at 310nm, which is provided by genetic <i>E. coli</i> Nissle in LB broth after 48 hours. |
| − | We compared the TAL constructs containing the native and B0034 ribosome binding sites (BBa_K2997010), to determine if p-Coumaric Acid production is improved by changing the ribosome binding sites. From the results seen in Fig. 5, BBa_K2997010 is able to produce a higher amount of p-Coumaric acid. Hence, we are able to prove that by changing the RBS (from Native to B0034), the conversion of tyrosine into p-Coumaric acid can increase by 1.73-fold. | + | We compared the TAL constructs containing the native and B0034 ribosome binding sites ([[Part:BBa_K2997010 | BBa_K2997010]] ), to determine if <i>p</i>-Coumaric Acid production is improved by changing the ribosome binding sites. From the results seen in Fig. 5, [[Part:BBa_K2997010 | BBa_K2997010]] is able to produce a higher amount of <i>p</i>-Coumaric acid. Hence, we are able to prove that by changing the RBS (from Native to B0034), the conversion of tyrosine into <i>p</i>-Coumaric acid can increase by 1.73-fold. |
<html> | <html> | ||
<br> | <br> | ||
| Line 88: | Line 91: | ||
</html> | </html> | ||
| − | Fig. 5. p-Coumaric acid/O.D.600 levels of E. coli Nissle with TAL and tyrP in LB with 1mM tyrosine | + | Fig. 5. <i>p</i>-Coumaric acid/O.D.600 levels of <i>E. coli</i> Nissle with TAL and <i>tyrP</i> in LB with 1mM tyrosine |
| + | |||
| + | ===Refence=== | ||
| + | [1] http://2013.igem.org/Team:Uppsala | ||
Latest revision as of 19:03, 21 October 2019
Sam8 gene with ribosome binding site
This part includes the sam8 gene of the bacterium Saccharothrix espanaensis DSM 4429 along with its native ribosome binding site. The coding sequence alone, with no ribosome binding site, as also been submitted as BBa_I742141. Sam8 encodes a tyrosine ammonia lyase which converts tyrosine to p-coumaric acid. This gene is naturally involved in caffeic acid biosynthesis (Reference: Berner, M., Krug, D., Bihlmaier, C., Vente, A., Muller, R. and Bechthold, A. 'Genes and Enzymes Involved in Caffeic Acid Biosynthesis in the Actinomycete Saccharothrix espanaensis' J. Bacteriol. 188 (7), 2666-2673 (2006)) and is part of the artifical vanillin biosynthesis pathway devised by the Edinburgh iGEM2007 team.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 808
Illegal BamHI site found at 1210 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 551
Illegal AgeI site found at 498
Illegal AgeI site found at 667 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 102
Illegal BsaI site found at 368
NCKU_Tainan 2019 Improvement
Click BBa_K2997011 to view the part, which we improved and used for the expression.
Background
Tyrosine ammonia-lyase (TAL) is an enzyme that converts tyrosine into p-Coumaric acid. We engineered E. coli Nissle to express TAL to turn the excess tyrosine inside the gut into p-Coumaric acid, and compare the different constructs for improvement, BBa_K2997009 and BBa_K2997010.
Expression in E. coli
We constructed Tyrosine Ammonia Lyase with a native RBS , and transformed the plasmid into E. coli Nissle 1917 and confirmed it by double digestion. The results are as follows:
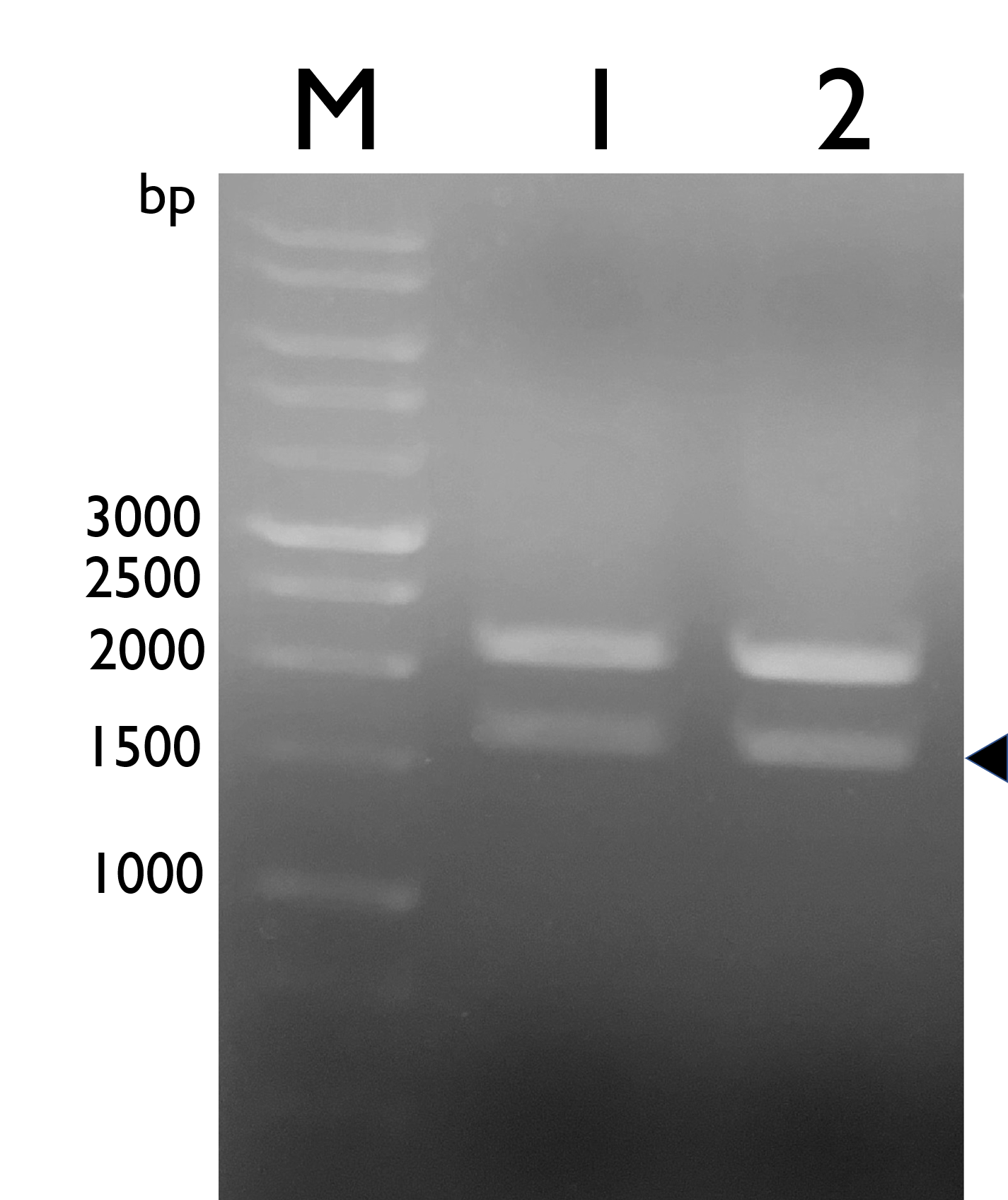
Fig. 1. Confirmation of BBa_K2997009 by double digestion, arrow indicates TAL with NRBS (~1600bp).M: Marker; Lane 1: pSB1C3- BBa_K2997009; Lane 2: BBa_K2997009
Then, we carried out SDS-PAGE to check the protein expression of TAL, and the expected protein size of TAL is 54kDa. As seen in the results below, however, there’s no distinguishable band around both sizes.
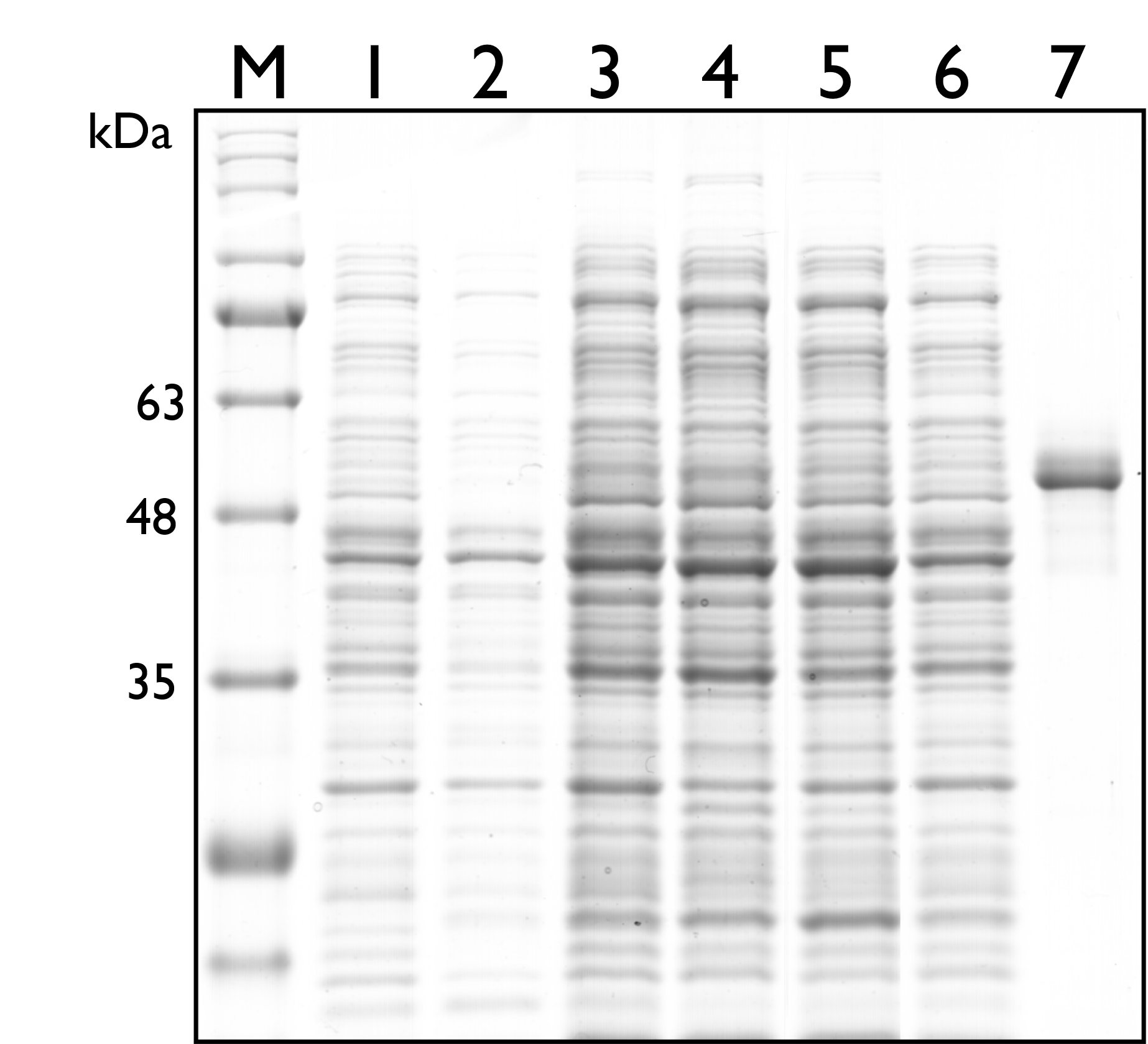
Fig. 2. 12% SDS PAGE of E. coli Nissle 1917 with different plasmids. M: Marker; Lane 1: Wild Type; Lane 2: pSB1C3; Lane 3: BBa_K2997009 ; Lane 4: BBa_K2997010; Lane 5: Dual plasmid containing BBa_K2997009 and BBa_K2997000; Lane 6: Dual plasmid containing BBa_K2997010 and BBa_K2997000; Lane 7: Positive control (c.d. 3392)
RT-PCR
RT-PCR experiment was used to confirm that the constructed TAL Biobrick is being transcribed. As seen in Fig. 3, cDNA for both bacteria carrying TAL constructs are being detected by PCR. Confirming that the TAL genes is actually being transcribed in E. coli Nissle.

Fig. 3. Reverse Transcription(RT)-PCR Results to confirm that our construct is being transcribed. (a) Schematics show location of amplified regions and primers. (b) 1.5% Agarose gel shows PCR results. All products have expected size of 250bp as shown in (a). (cDNA: Total cDNA; RNA: Total RNA; plasmid: pSB1C3 containing respective Biobrick.)

Table 1. Description of templates and primers being used in each lane.
TyrP and TAL Assay
To confirm the protein activity of TAL and TyrP, we performed a functional test using n-octanol extraction method, TyrP and TAL Assay, which was previously proposed by iGEM Uppsala 2013 and has been verified by HPLC[1]. The p-Coumaric acid concentration was measured through the absorbance value at 310nm wavelength under Nanodrop UV-Vis wavelength. The standard curve of p-Coumaric acid was drawn in Fig.4 to determine the relationship between p-Coumaric acid concentrations and its 310nm arbitrary unit (a.u). Our samples with TAL constructs were then mapped onto the standard curve, to know how much p-Coumaric acid is being produced.
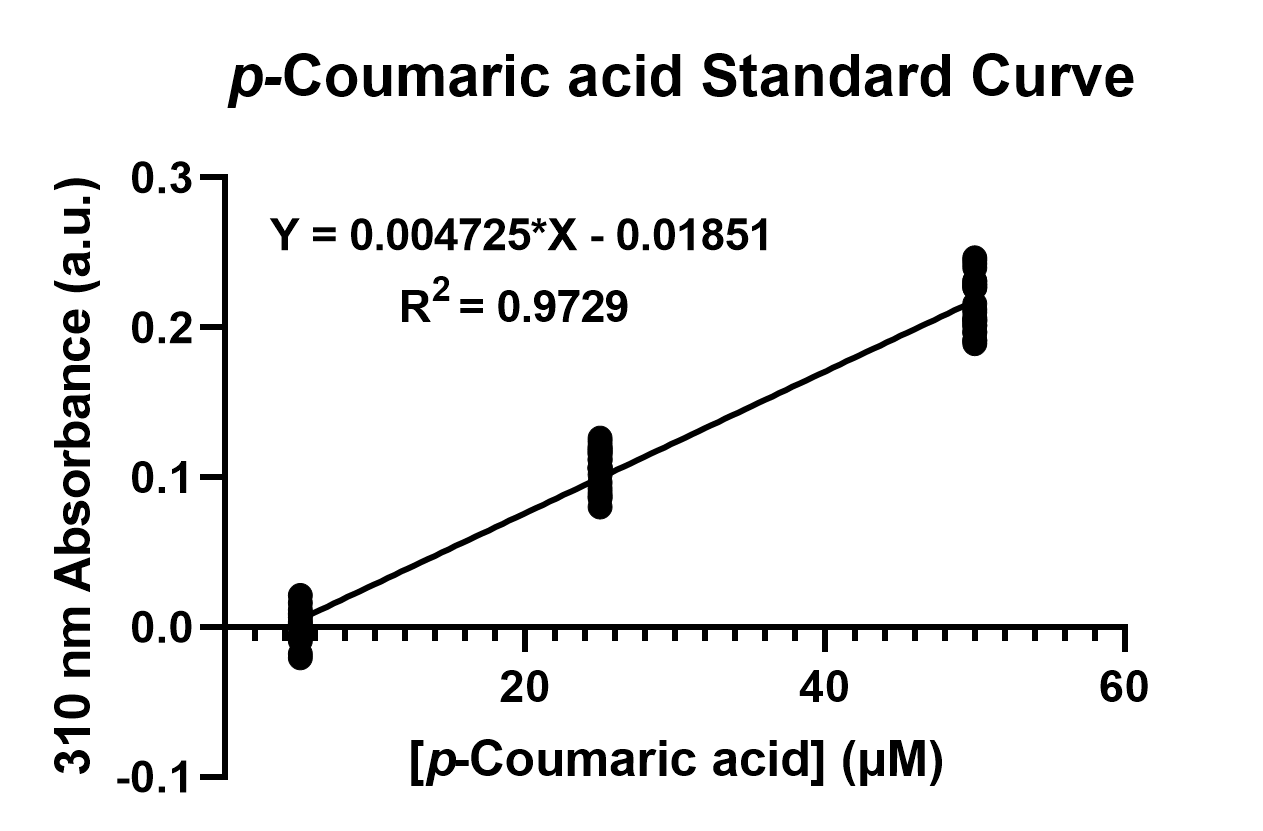
Fig. 4. The standard curve of p-Coumaric acid concentration in correlation with absorbance at 310nm, which is provided by genetic E. coli Nissle in LB broth after 48 hours.
We compared the TAL constructs containing the native and B0034 ribosome binding sites ( BBa_K2997010 ), to determine if p-Coumaric Acid production is improved by changing the ribosome binding sites. From the results seen in Fig. 5, BBa_K2997010 is able to produce a higher amount of p-Coumaric acid. Hence, we are able to prove that by changing the RBS (from Native to B0034), the conversion of tyrosine into p-Coumaric acid can increase by 1.73-fold.
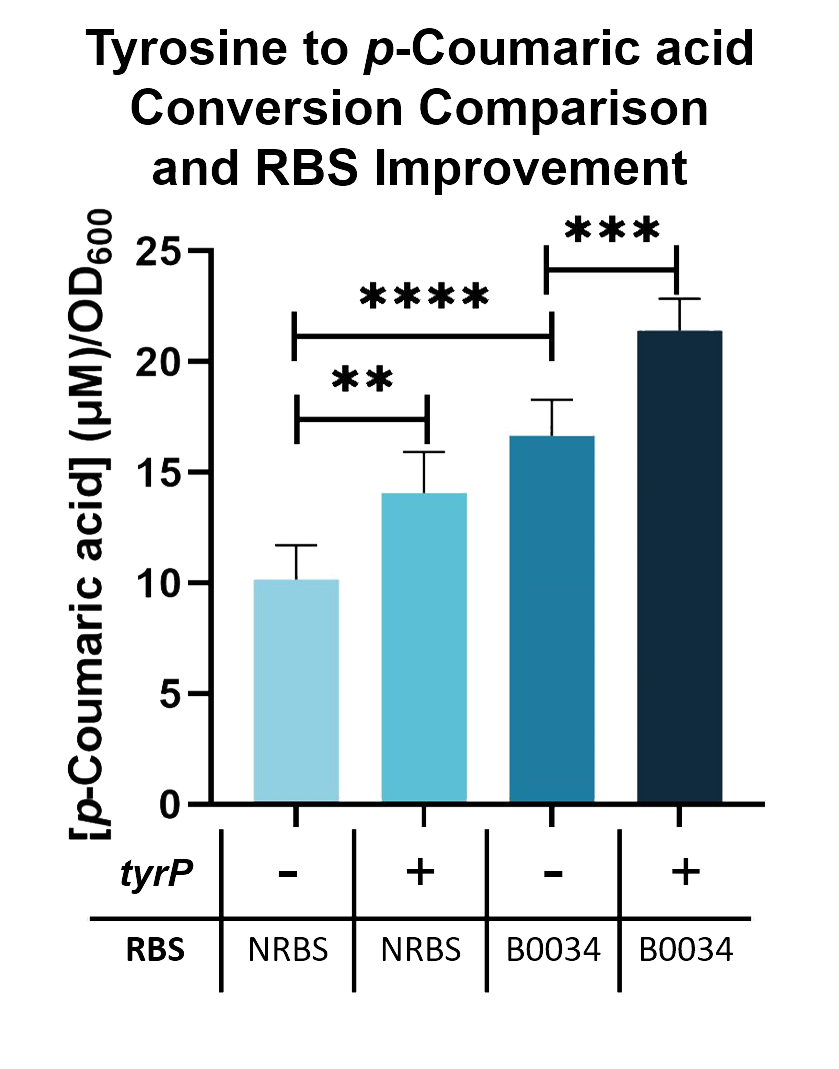
Fig. 5. p-Coumaric acid/O.D.600 levels of E. coli Nissle with TAL and tyrP in LB with 1mM tyrosine
Refence
[1] http://2013.igem.org/Team:Uppsala
