Difference between revisions of "Part:BBa K1141000"
PeterGockel (Talk | contribs) |
PeterGockel (Talk | contribs) |
||
| Line 9: | Line 9: | ||
<p align="center" id="legend">Figure 1: Excitation (left peak) and Emission (right peak) spectra for mCherry.</p> | <p align="center" id="legend">Figure 1: Excitation (left peak) and Emission (right peak) spectra for mCherry.</p> | ||
| − | <h1> Translation Enhancing 5'-UTR (Improvement by iGEM | + | <h1> Translation Enhancing 5'-UTR (Improvement by iGEM TU_Darmstadt 2019) </h1> |
| + | |||
| + | <html> | ||
| + | <script id="MathJax-script" async | ||
| + | src="https://2019.igem.org/wiki/index.php?title=Template:TU_Darmstadt/MathjaxJS&action=raw&ctype=text/javascript"></script> | ||
| + | |||
| + | <div class="container"> | ||
| + | <div class="row"> | ||
| + | <div class="col mx-2"> | ||
| + | <h3>Usage and Biology</h3> | ||
| + | <hr class="head"> | ||
| + | <p> | ||
| + | Improved version: <a | ||
| + | href="https://parts.igem.org/Part:BBa_K1758100" | ||
| + | target="_blank"> BBa_K1758100</a> | ||
| + | <br></br> | ||
| + | The part was improved in terms of its expression level, based on the insertion of a 5’-untranslated region (5’-UTR) upstream of the coding sequence. This 5'-UTR was adapted from iGEM Bielefeld 2015 ( <a | ||
| + | href="https://parts.igem.org/Part:BBa_K1758100" | ||
| + | target="_blank"> BBa_K1758100</a>) and is based on the research of Olins <i>et</i> al | ||
| + | <sup id="cite_ref-1" class="reference"> | ||
| + | <a href="#cite_note-1">[1] </a> | ||
| + | </sup> | ||
| + | and Takahashi <i>et</i> al. | ||
| + | <sup id="cite_ref-2" class="reference"> | ||
| + | <a href="#cite_note-2">[2] </a> | ||
| + | </sup>. | ||
| + | It contains the strong ribosomal binding site (RBS) g10-L from the T7 bacteriophage and a sequence that plays a role in the regulation of mRNA binding to and release from the 30S ribosomal subunit. The 5'-UTR therefore enhances the translation efficiency of the following coding sequence (CDS) (Fig. 1). | ||
| + | |||
| + | </p> | ||
| + | |||
| + | <img | ||
| + | class="img-fluid center" | ||
| + | src="https://2019.igem.org/wiki/images/6/62/T--TU_Darmstadt--RibosomLiberation.png" | ||
| + | style="max-width:80%" | ||
| + | /> | ||
| + | <div class="caption"> | ||
| + | <p> | ||
| + | <b> | ||
| + | <center> Figure 1: | ||
| + | </b> Schematic depiction of the composition and interaction of the enhancer sequence with the 30S ribosomal subunit described by Takahashi <i>et.</i> al. <sup id="cite_ref-2" class="reference"> | ||
| + | <a href="#cite_note-2">[2] </a> | ||
| + | </sup>. | ||
| + | </center> | ||
| + | </p> | ||
| + | </div> | ||
| + | |||
| + | <p> | ||
| + | The sequence of the translation enhancing 5’-UTR can be divided into the four main features listed below: | ||
| + | </p> | ||
| + | |||
| + | |||
| + | |||
| + | <img | ||
| + | class="img-fluid center" | ||
| + | src="https://2019.igem.org/wiki/images/f/f3/T--TU_Darmstadt--EnhancerInColor.png" | ||
| + | style="text= align; max-width:85%" | ||
| + | /> | ||
| + | |||
| + | |||
| + | <br></br> | ||
| + | <div class="container-noborders"> | ||
| + | <div class="table-responsive-sm"> | ||
| + | <table class="table table-light"> | ||
| + | <tr> | ||
| + | <th scope="col">Sequence</th> | ||
| + | <th scope="col">Function</th> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="color:#d59b9b; word-wrap:break-word;"> | ||
| + | AATAATTTTGTT<br />TTAACTTTAA | ||
| + | </td> | ||
| + | <td> | ||
| + | The T7 g10 leader sequence (first described by Olins <i>et</i> al<sup id="cite_ref-1" class="reference"> | ||
| + | <a href="#cite_note-1">[1] </a> | ||
| + | </sup>)increases the efficiency of translation initiation. This sequence contains the epsilon motif TTAACTTTA which enhances the binding of the mRNA to the 16S rRNA. | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="color:#a5c8c8;">poly-A</td> | ||
| + | <td> | ||
| + | Referring to Takahashi et al.<sup id="cite_ref-2" class="reference"> | ||
| + | <a href="#cite_note-2">[2] </a> | ||
| + | </sup> a spacer between the epsilon motive and the RBS improves the translation rate. | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="color:#698888;">GAAGGAG</td> | ||
| + | <td> | ||
| + | According to Karig <i>et</i> al.<sup id="cite_ref-3" class="reference"> | ||
| + | <a href="#cite_note-3">[3] </a> | ||
| + | </sup> and Lentini et. al<sup id="cite_ref-4" class="reference"> | ||
| + | <a href="#cite_note-4">[4] </a> </sup> a distance of 4-9 bases between RBS and start codon increases the translation efficiency. | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="color:#cc9966;">AATAATCT</td> | ||
| + | <td> | ||
| + | According to Lentini et. al<sup id="cite_ref-4" class="reference"> | ||
| + | <a href="#cite_note-4">[4] </a> </sup> an AT-rich composition between the RBS and the start codon results in the best expression results. | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div> | ||
</html> | </html> | ||
Revision as of 22:20, 18 October 2019
Plac-RBS-mCherry-double terminator (IPTG-inducible)
This part contains all the necessary DNA to produce the red fluorescent protein mCherry with the PLac promoter. PLac repression is low in most E. coli strains and the promoter will be leaky. Efficient repression can be implemented by additionally transforming a strain with the pREP4 plasmid from the M15 E. coli strain of QIAGEN, or transforming the part into this strain. The figure below shows the excitation and emission spectra as can be obtained from CHROMA's spectra viewer:

Figure 1: Excitation (left peak) and Emission (right peak) spectra for mCherry.
Translation Enhancing 5'-UTR (Improvement by iGEM TU_Darmstadt 2019)
Usage and Biology
Improved version: BBa_K1758100
The part was improved in terms of its expression level, based on the insertion of a 5’-untranslated region (5’-UTR) upstream of the coding sequence. This 5'-UTR was adapted from iGEM Bielefeld 2015 ( BBa_K1758100) and is based on the research of Olins et al
[1]
and Takahashi et al.
[2]
.
It contains the strong ribosomal binding site (RBS) g10-L from the T7 bacteriophage and a sequence that plays a role in the regulation of mRNA binding to and release from the 30S ribosomal subunit. The 5'-UTR therefore enhances the translation efficiency of the following coding sequence (CDS) (Fig. 1).
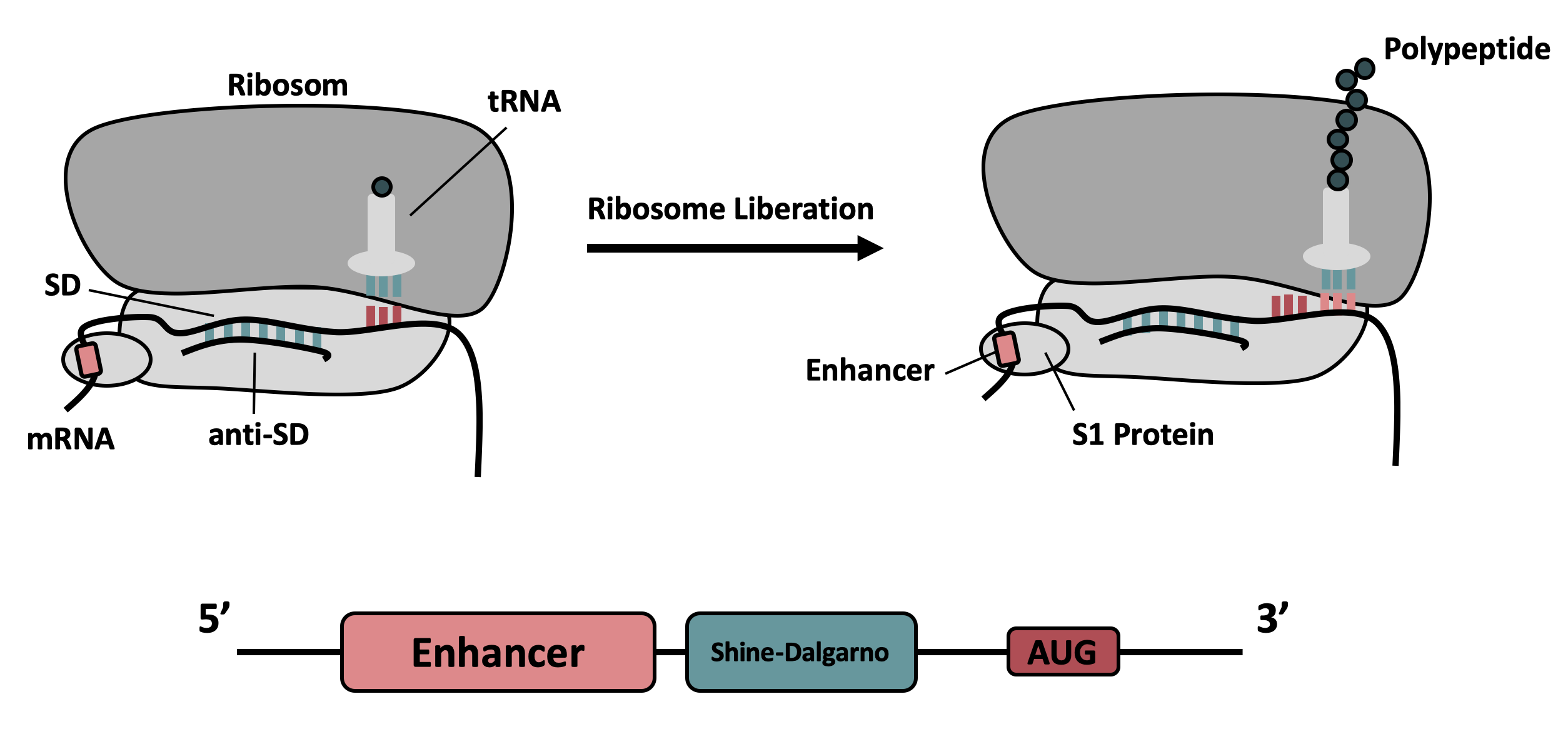
The sequence of the translation enhancing 5’-UTR can be divided into the four main features listed below:

| Sequence | Function |
|---|---|
|
AATAATTTTGTT TTAACTTTAA |
The T7 g10 leader sequence (first described by Olins et al [1] )increases the efficiency of translation initiation. This sequence contains the epsilon motif TTAACTTTA which enhances the binding of the mRNA to the 16S rRNA. |
| poly-A | Referring to Takahashi et al. [2] a spacer between the epsilon motive and the RBS improves the translation rate. |
| GAAGGAG | According to Karig et al. [3] and Lentini et. al [4] a distance of 4-9 bases between RBS and start codon increases the translation efficiency. |
| AATAATCT | According to Lentini et. al [4] an AT-rich composition between the RBS and the start codon results in the best expression results. |
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
