Difference between revisions of "Part:BBa K3147003"
(→I : parts BBa_K3147003 (mRFP1-TEVcs-SSRA) function) |
(→II. Proof of function) |
||
| Line 15: | Line 15: | ||
===II. Proof of function=== | ===II. Proof of function=== | ||
| − | The experimental approach to test the activity of these reporters was to compare the basal fluorescence rate of mRFP1 with a TEV cutting site "cleaved" to this construction. For this purpose we made a control construction: mRFP1-TEVcs (BBa_K3147004) similar to this one by removing the proteolysis tag and simulating a cut by the TEV. The construction was cloned by Gibson Assembly in a pBbB8k-GFP (https://www.addgene.org/35363/ ) backbone under the control of a | + | The experimental approach to test the activity of these reporters was to compare the basal fluorescence rate of mRFP1 with a TEV cutting site "cleaved" to this construction. For this purpose we made a control construction: mRFP1-TEVcs (BBa_K3147004) similar to this one by removing the proteolysis tag and simulating a cut by the TEV. The construction was cloned by Gibson Assembly in a pBbB8k-GFP (https://www.addgene.org/35363/ ) backbone under the control of a pBAD promoter. |
| − | + | ||
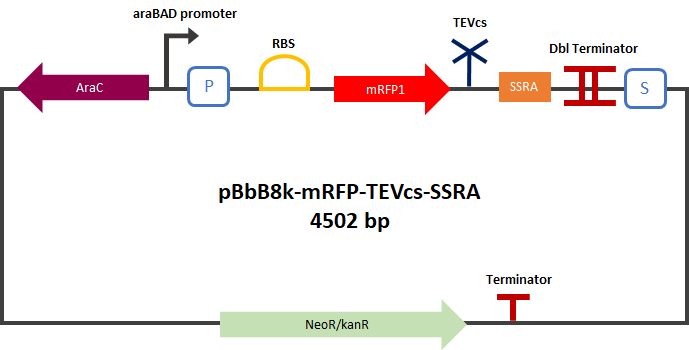
<div align="center">[[File:PlasmideK3147003.png|400px]]</div> | <div align="center">[[File:PlasmideK3147003.png|400px]]</div> | ||
<div align="center"><b>Figure 2</b>: mRFP1-TEVcs-ssrA reporter gene in its pBbB8k-GFP backbone.</div> | <div align="center"><b>Figure 2</b>: mRFP1-TEVcs-ssrA reporter gene in its pBbB8k-GFP backbone.</div> | ||
| − | We compared the basal fluorescence of strain E. coli NEB10β transformed with the mRFP1-TEVcs construction to an E. coli NEB10β strain transformed with the mRFP1-TEVcs-ssrA construction. Fluorescence was quantified after induction with | + | We compared the basal fluorescence of strain E. coli NEB10β transformed with the mRFP1-TEVcs construction to an E. coli NEB10β strain transformed with the mRFP1-TEVcs-ssrA construction. Fluorescence was quantified after overnight induction with 1% arabinose on a plate reader. |
| + | |||
| + | Below are the fluorescence measurements of the mRFP1-TEVcs-ssrA and of the mRFP1-TEVcss at 30 and 37°C. | ||
Revision as of 09:45, 18 October 2019
mRFP1 fused to a TEV-cleavable ssrA tag
I : parts BBa_K3147003 (mRFP1-TEVcs-SSRA) function
This construction produces an mRFP1 (BBA_E1010)[1] fused in C-ter with a fast degradation tag called ssrA [2] (BBA_M0050). The TEV cutting site (BBa_J18918) was added between the mRFP1 and the ssrA tag. This is a sister construction to our sfGFP TEV reporter (Ba_K3147000). This construction was used to test the specificity of KARMA using a VHH targeting GFP. In the presence of TEV the ssrA is cleaved and mRFP1 is not degraded anymore.
II. Proof of function
The experimental approach to test the activity of these reporters was to compare the basal fluorescence rate of mRFP1 with a TEV cutting site "cleaved" to this construction. For this purpose we made a control construction: mRFP1-TEVcs (BBa_K3147004) similar to this one by removing the proteolysis tag and simulating a cut by the TEV. The construction was cloned by Gibson Assembly in a pBbB8k-GFP (https://www.addgene.org/35363/ ) backbone under the control of a pBAD promoter.
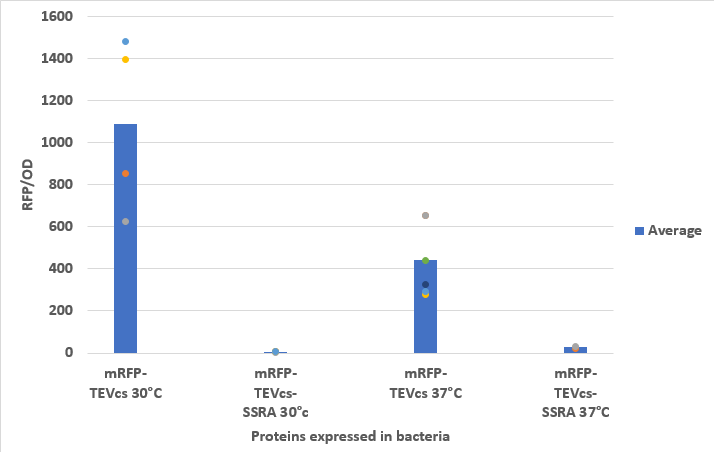
We compared the basal fluorescence of strain E. coli NEB10β transformed with the mRFP1-TEVcs construction to an E. coli NEB10β strain transformed with the mRFP1-TEVcs-ssrA construction. Fluorescence was quantified after overnight induction with 1% arabinose on a plate reader.
Below are the fluorescence measurements of the mRFP1-TEVcs-ssrA and of the mRFP1-TEVcss at 30 and 37°C.
Reference
[1] Levraud, Jean-Pierre et al. 2007. « Identification of the Zebrafish IFN Receptor: Implications for the Origin of the Vertebrate IFN System ». The Journal of Immunology 178(7): 4385‑94. doi:10.4049/jimmunol.178.7.4385
[2] Sunohara, T., Abo, T., Inada, T., & Aiba, H. (2002). The C-terminal amino acid sequence of nascent peptide is a major determinant of SsrA tagging at all three stop codons. RNA (New York, N.Y.), 8(11), 1416–1427. doi:10.1017/s1355838202020198
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 691
- 1000COMPATIBLE WITH RFC[1000]