Difference between revisions of "Part:BBa K302033"
PhilipHall (Talk | contribs) |
Jasmine wang (Talk | contribs) |
||
| Line 15: | Line 15: | ||
===Functional Parameters=== | ===Functional Parameters=== | ||
<partinfo>BBa_K302033 parameters</partinfo> | <partinfo>BBa_K302033 parameters</partinfo> | ||
| + | <!-- --> | ||
| + | |||
| + | == Characterized by BNU-China 2019 == | ||
| + | |||
| + | We characterize mazF (BBa_K302033) by an induced suicide system, in which the downstream mazF (BBa_K302033) gene encoding toxin MazF is put under the control of L-arabinose induced promoter pBAD (BBa_K206000). MazF is an endoribonuclease that cleaves RNAs at ACA sites and causes the death of microbe as a reporter [1]. As a result, we can characterize mazF in a cell density-dependent manner in Escherichia coli K-12. | ||
| + | |||
| + | [[Image:2019_BNU-China_BBa_K206000_pic1.png]] | ||
| + | |||
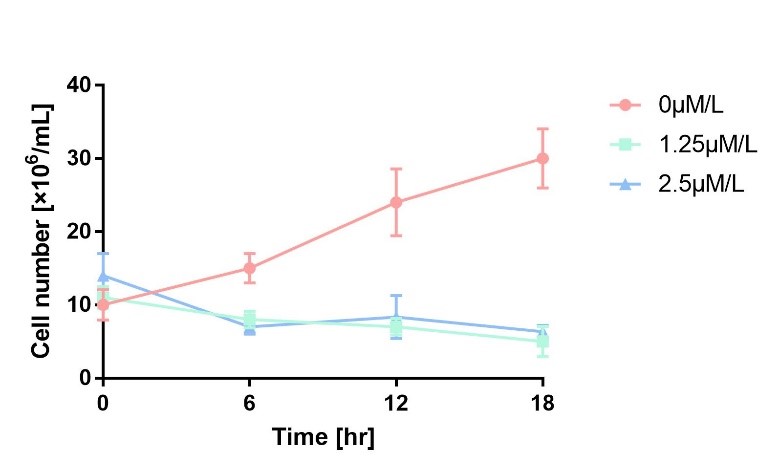
| + | In order to characterize the toxicity of protein encoded by mazF, we take engineered microbe without induction as control group. As is shown in Fig.1, the cell number of experimental groups show a significant decrease upon induction, which indicates the protein encoded by mazF is lethal to E. coli. | ||
| + | |||
| + | |||
| + | [[Image:2019_BNU-China_BBa_K206000_pic2.png]] | ||
| + | |||
| + | Figure 1 Cell number declines after induction. It proves that protein encoded by mazF is lethal to E. coli. | ||
| + | |||
| + | Furthermore, in order to prove that MazF kills the cell without lysing it, we measure OD600 of each sample to give an overall number of intact bacteria, dead and alive. As is shown in Fig.2, there is little difference between control and experimental groups, although it is validated numbers of alive cells differ. Hence, we reach a conclusion that protein encoded by mazF mediates suicide of cells without causing lysis of bacterial cells. | ||
| + | |||
| + | [[Image:2019_BNU-China_BBa_K206000_pic3.png]] | ||
| + | |||
| + | Figure 2 Absorbance at 600nm with time | ||
| + | |||
| + | <b>Experimental approach</b> | ||
| + | |||
| + | 1. Transform the plasmids into E. coli DH5α competent cells. | ||
| + | 2. The engineered bacteria are cultured in 200mL LB-ampicillin (50 ng/µl) medium overnight at 37℃, 200rpm; | ||
| + | 3. Equally divide the culture into 90 centrifuge tubes, which is 1mL respectively. Centrifuge them at 4000rpm for 5 minutes. Discard the liquid. | ||
| + | 4. Resuspend 30 tubes of collected bacteria with LB-ampicillin (50 ng/µl) containing 1.25μM/L and 2.5μM/L L-arabinose respectively as experimental groups. Resuspend 30 tubes of bacteria with pure LB-ampicillin (50 ng/µl) medium. | ||
| + | 5. Collect 3 tubes of all groups every 6 hours, dilute all of the samples to 107 times and then spread them on solid LB-ampicillin (50 ng/µl) medium separately. At the same time, refresh the medium to maintain the concentration of L-arabinose. | ||
| + | 6. Count the number of colonies in 5 cm2 per plate after cultured for 24 hours at 37℃ | ||
| + | 7. Three repicas are tested in each group. | ||
| + | |||
| + | <b>Reference</b> | ||
| + | |||
| + | [1] Diana Széliová, Ján Krahulec, Martin Šafránek, et al. Modulation of heterologous expression from PBAD promoter in Escherichia coli production strains[J]. Journal of Biotechnology, 2016, 236:1-9. | ||
| + | |||
| + | <!-- Add more about the biology of this part here | ||
| + | ===Usage and Biology=== | ||
| + | |||
| + | <!-- --> | ||
| + | <span class='h3bb'>Sequence and Features</span> | ||
| + | <partinfo>BBa_K654059 SequenceAndFeatures</partinfo> | ||
| + | |||
| + | |||
| + | <!-- Uncomment this to enable Functional Parameter display | ||
| + | ===Functional Parameters=== | ||
| + | <partinfo>BBa_K654059 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
Revision as of 04:09, 9 October 2019
mazF
Encodes a stable non-specific ribonuclease in Bacillus Subtilis. It is used in conjungtion with mazE in Bacillus Subtilis to provide a toxin-antitoxin kill switch in various stressful conditions. When expression of both genes is turned off (as both are under contol of same promoter) mazE will be degraded faster than mazF. There is then no inhibiton of mazF, killing the cell.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Characterized by BNU-China 2019
We characterize mazF (BBa_K302033) by an induced suicide system, in which the downstream mazF (BBa_K302033) gene encoding toxin MazF is put under the control of L-arabinose induced promoter pBAD (BBa_K206000). MazF is an endoribonuclease that cleaves RNAs at ACA sites and causes the death of microbe as a reporter [1]. As a result, we can characterize mazF in a cell density-dependent manner in Escherichia coli K-12.
In order to characterize the toxicity of protein encoded by mazF, we take engineered microbe without induction as control group. As is shown in Fig.1, the cell number of experimental groups show a significant decrease upon induction, which indicates the protein encoded by mazF is lethal to E. coli.
Figure 1 Cell number declines after induction. It proves that protein encoded by mazF is lethal to E. coli.
Furthermore, in order to prove that MazF kills the cell without lysing it, we measure OD600 of each sample to give an overall number of intact bacteria, dead and alive. As is shown in Fig.2, there is little difference between control and experimental groups, although it is validated numbers of alive cells differ. Hence, we reach a conclusion that protein encoded by mazF mediates suicide of cells without causing lysis of bacterial cells.
File:2019 BNU-China BBa K206000 pic3.png
Figure 2 Absorbance at 600nm with time
Experimental approach
1. Transform the plasmids into E. coli DH5α competent cells. 2. The engineered bacteria are cultured in 200mL LB-ampicillin (50 ng/µl) medium overnight at 37℃, 200rpm; 3. Equally divide the culture into 90 centrifuge tubes, which is 1mL respectively. Centrifuge them at 4000rpm for 5 minutes. Discard the liquid. 4. Resuspend 30 tubes of collected bacteria with LB-ampicillin (50 ng/µl) containing 1.25μM/L and 2.5μM/L L-arabinose respectively as experimental groups. Resuspend 30 tubes of bacteria with pure LB-ampicillin (50 ng/µl) medium. 5. Collect 3 tubes of all groups every 6 hours, dilute all of the samples to 107 times and then spread them on solid LB-ampicillin (50 ng/µl) medium separately. At the same time, refresh the medium to maintain the concentration of L-arabinose. 6. Count the number of colonies in 5 cm2 per plate after cultured for 24 hours at 37℃ 7. Three repicas are tested in each group.
Reference
[1] Diana Széliová, Ján Krahulec, Martin Šafránek, et al. Modulation of heterologous expression from PBAD promoter in Escherichia coli production strains[J]. Journal of Biotechnology, 2016, 236:1-9.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]