Difference between revisions of "Part:BBa K2259091"
(→About SynORI) |
|||
| Line 3: | Line 3: | ||
<partinfo>BBa_K2259091 short</partinfo> | <partinfo>BBa_K2259091 short</partinfo> | ||
| − | This device consists of replication initiator RNA II that is modulated by replication regulator RNA I under the control of | + | This device consists of replication initiator RNA II that is modulated by replication regulator RNA I under the control of Rhamnose promoter. |
| − | Increased rhamnose concentrations lead to increase in RNA I and consequently - decrease in plasmid copy number. | + | Increased rhamnose concentrations lead to increase in RNA I concentration and consequently - decrease in plasmid copy number. |
See how this part fits into the whole SynORI framework [[#About SynORI|by pressing here!]] | See how this part fits into the whole SynORI framework [[#About SynORI|by pressing here!]] | ||
| Line 27: | Line 27: | ||
===ColE1 plasmid replication overview=== | ===ColE1 plasmid replication overview=== | ||
| − | [[Image:Cole1 horizontal cropped.png|center| | + | [[Image:Cole1 horizontal cropped.png|300px| center|]] |
| − | + | <b><center>Figure 1. Main principles of ColE1 plasmid family replication</center> | |
| − | |||
| − | For RNA I to inhibit primer formation, it must bind before the nascent RNA II transcript extends to the replication origin. Consequently, the concentration of RNA I and the rate of binding of RNA I to RNA II is critical for regulation of primer formation and thus for plasmid replication. | + | ColE1-type plasmid replication begins with the synthesis of plasmid encoded RNA II</b> (also called primer transcript) by RNA polymerase which initiates transcription at a site 555bp upstream of origin of replication. The RNA transcript forms a RNA - DNA hybrid with template DNA near the origin of replication. Hybridized RNA is then cleaved at the replication origin by RNAse H and serves as a primer for DNA synthesis by DNA polymerase I (Figure 1. A).<ref>Itoh, T. and Tomizawa, J. (1980). Formation of an RNA primer for initiation of replication of ColE1 DNA by ribonuclease H. Proceedings of the National Academy of Sciences, 77(5), pp.2450-2454.</ref> |
| + | |||
| + | <b>Initiation of replication can be inhibited by plasmid encoded small RNA, called RNA I </b>. Synthesis of RNA I starts 445 bp upstream of the replication origin and proceeds in the direction opposite to that of RNA II synthesis and terminates near the RNA II transcription initiation site. <b>RNA I binds to RNA II</b> and thereby prevents the formation of a secondary structure of RNA II that is necessary for hybridization of RNA II to the template DNA (Figure 1. B).<ref>Tomizawa, J. (1984). Control of cole 1 plasmid replication: The process of binding of RNA I to the primer transcript. Cell, 38(3), pp.861-870.</ref> | ||
| + | |||
| + | For RNA I to inhibit primer formation, it must bind before the nascent RNA II transcript extends to the replication origin. Consequently, the concentration of RNA I and the rate of binding of RNA I to RNA II is critical for regulation of primer formation and thus for plasmid replication. <ref>Tomizawa, J. (1984). Control of cole 1 plasmid replication: The process of binding of RNA I to the primer transcript. Cell, 38(3), pp.861-870.</ref> | ||
The interaction between RNA I and RNA II can be amplified by Rop protein, see [[part:BBa_K2259010]]. | The interaction between RNA I and RNA II can be amplified by Rop protein, see [[part:BBa_K2259010]]. | ||
| Line 42: | Line 45: | ||
SynORI is a framework for multi-plasmid systems created by ''Vilnius-Lithuania 2017'' which enables quick and easy workflow with multiple plasmids, while also allowing to freely pick and modulate copy number for every unique plasmid group! Read more about [http://2017.igem.org/Team:Vilnius-Lithuania SynORI here]! | SynORI is a framework for multi-plasmid systems created by ''Vilnius-Lithuania 2017'' which enables quick and easy workflow with multiple plasmids, while also allowing to freely pick and modulate copy number for every unique plasmid group! Read more about [http://2017.igem.org/Team:Vilnius-Lithuania SynORI here]! | ||
| − | === | + | ===This device in SynORI=== |
This device demonstrates SynORI framework ability to create an inducible plasmid copy number device. | This device demonstrates SynORI framework ability to create an inducible plasmid copy number device. | ||
Revision as of 17:09, 1 November 2017
SynORI inducible plasmid copy number device
This device consists of replication initiator RNA II that is modulated by replication regulator RNA I under the control of Rhamnose promoter.
Increased rhamnose concentrations lead to increase in RNA I concentration and consequently - decrease in plasmid copy number.
See how this part fits into the whole SynORI framework by pressing here!
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Contents
Introduction
Biology
ColE1 plasmid replication overview
ColE1-type plasmid replication begins with the synthesis of plasmid encoded RNA II (also called primer transcript) by RNA polymerase which initiates transcription at a site 555bp upstream of origin of replication. The RNA transcript forms a RNA - DNA hybrid with template DNA near the origin of replication. Hybridized RNA is then cleaved at the replication origin by RNAse H and serves as a primer for DNA synthesis by DNA polymerase I (Figure 1. A).[1]
Initiation of replication can be inhibited by plasmid encoded small RNA, called RNA I . Synthesis of RNA I starts 445 bp upstream of the replication origin and proceeds in the direction opposite to that of RNA II synthesis and terminates near the RNA II transcription initiation site. RNA I binds to RNA II and thereby prevents the formation of a secondary structure of RNA II that is necessary for hybridization of RNA II to the template DNA (Figure 1. B).[2]
For RNA I to inhibit primer formation, it must bind before the nascent RNA II transcript extends to the replication origin. Consequently, the concentration of RNA I and the rate of binding of RNA I to RNA II is critical for regulation of primer formation and thus for plasmid replication. [3]
The interaction between RNA I and RNA II can be amplified by Rop protein, see part:BBa_K2259010.
Usage with SynORI (Framework for multi-plasmid systems)
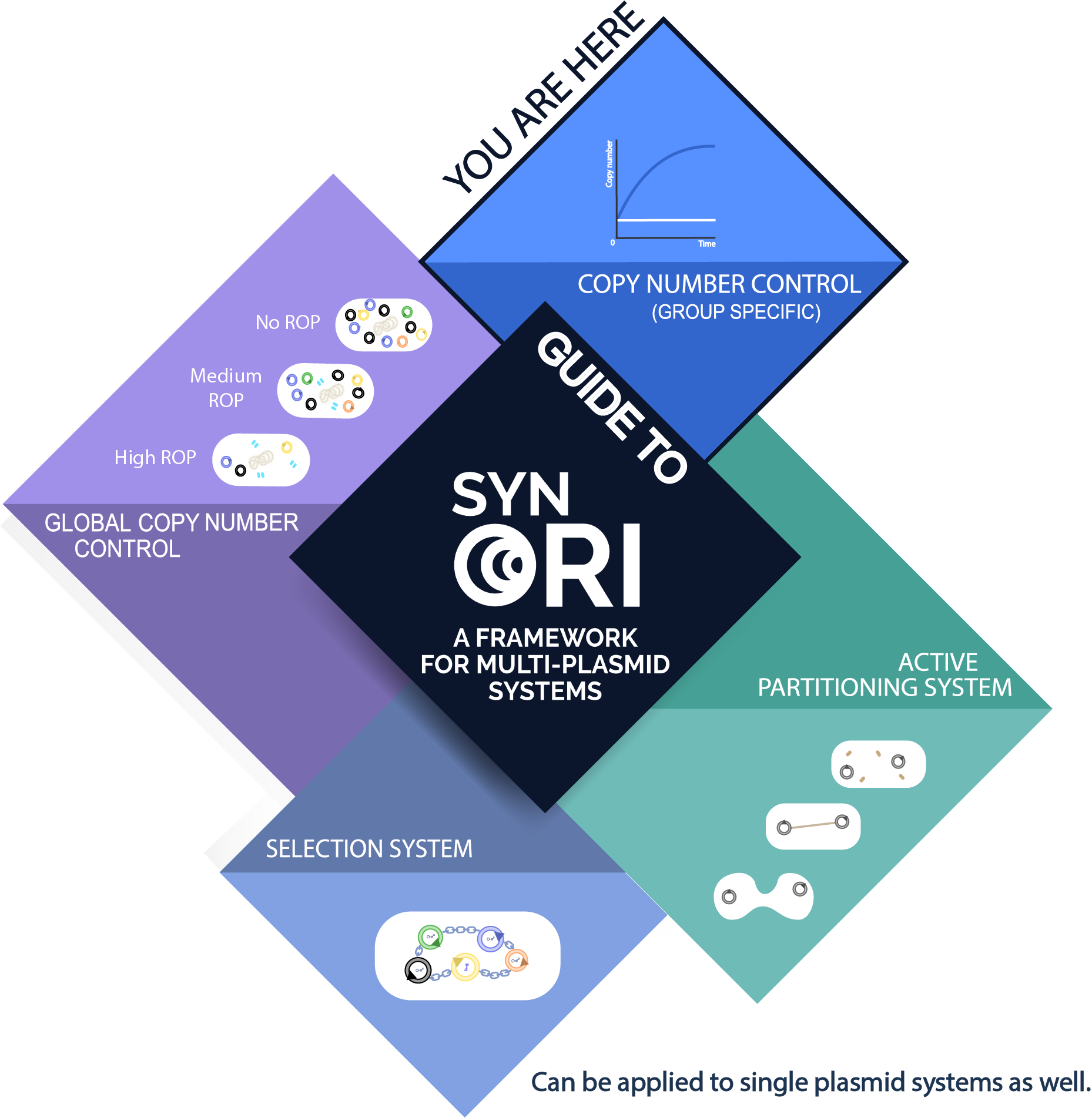
About SynORI
SynORI is a framework for multi-plasmid systems created by Vilnius-Lithuania 2017 which enables quick and easy workflow with multiple plasmids, while also allowing to freely pick and modulate copy number for every unique plasmid group! Read more about [http://2017.igem.org/Team:Vilnius-Lithuania SynORI here]!
This device in SynORI
This device demonstrates SynORI framework ability to create an inducible plasmid copy number device.
Characterization (Vilnius-Lithuania 2017)
RNA I and rhamnose
References
- ↑ Itoh, T. and Tomizawa, J. (1980). Formation of an RNA primer for initiation of replication of ColE1 DNA by ribonuclease H. Proceedings of the National Academy of Sciences, 77(5), pp.2450-2454.
- ↑ Tomizawa, J. (1984). Control of cole 1 plasmid replication: The process of binding of RNA I to the primer transcript. Cell, 38(3), pp.861-870.
- ↑ Tomizawa, J. (1984). Control of cole 1 plasmid replication: The process of binding of RNA I to the primer transcript. Cell, 38(3), pp.861-870.