Difference between revisions of "Part:BBa K1937002:Design"
| Line 13: | Line 13: | ||
<b>Creation:</b> | <b>Creation:</b> | ||
The BBa_K1937002 part contains the <i>repU</i> origin of replication of <i>B. subtilis</i> and the kanamycin resistance gene for <i>B. subtilis</i> (Figure 1).<br> | The BBa_K1937002 part contains the <i>repU</i> origin of replication of <i>B. subtilis</i> and the kanamycin resistance gene for <i>B. subtilis</i> (Figure 1).<br> | ||
| − | [[file:BBa_K1937002-map.jpg|thumb|center|800px|Figure 1: scheme of the Orikan part with <i>Bacillus subtilis</i> origin <i>repU</i> and a kanamycine resistance cassette.]] | + | [[file:BBa_K1937002-map.jpg|thumb|center|800px|<b>Figure 1:</b> scheme of the Orikan part with <i>Bacillus subtilis</i> origin <i>repU</i> and a kanamycine resistance cassette.]] |
<br>It was obtained by amplifying the repU-Kan region of pSB<sub>Bs</sub>0K-P (BBa_K1351040 or BBa_K823026) using primers OriKan forward (5’ cacagaatcaggggataacgcaggaaagaaACATGTAGTTATAAGTGACTAAACAAATAACTAAATAGATGGG) and OriKan reverse (5’ gttcctggccttttgctggccttttgctcaACATGTCGCAAAATGGCCCGATTTAAG). The resulting fragment was sub-cloned in the pSB<sub>1</sub>C<sub>3</sub> between the EcoRI and PstI restriction sites (Figure 2). | <br>It was obtained by amplifying the repU-Kan region of pSB<sub>Bs</sub>0K-P (BBa_K1351040 or BBa_K823026) using primers OriKan forward (5’ cacagaatcaggggataacgcaggaaagaaACATGTAGTTATAAGTGACTAAACAAATAACTAAATAGATGGG) and OriKan reverse (5’ gttcctggccttttgctggccttttgctcaACATGTCGCAAAATGGCCCGATTTAAG). The resulting fragment was sub-cloned in the pSB<sub>1</sub>C<sub>3</sub> between the EcoRI and PstI restriction sites (Figure 2). | ||
<br> | <br> | ||
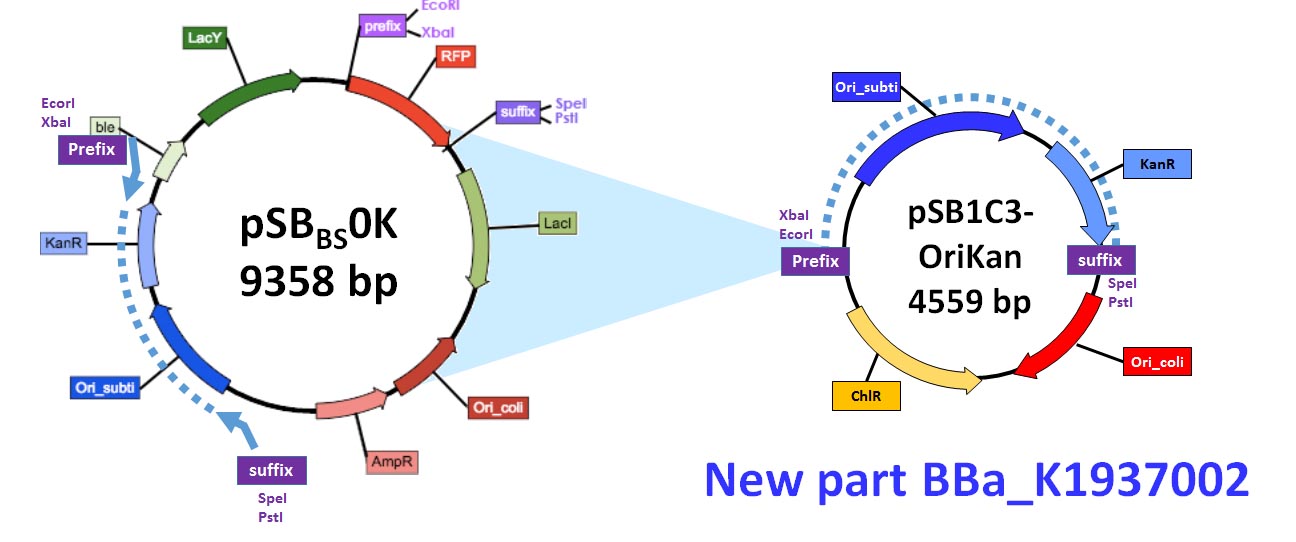
| − | [[Image:Toulouse_France_backbone2.jpg|thumb|center|800px|Figure 2: creation of the OriKan cassette. Position of the primers are indicated by the blue arrow on pSB<sub>Bs</sub>0K-P (left part of the figure). The resulting PCR fragment is the blue pointed line. After digestion by EcoRI and PstI and ligation in the pSB1C3 plasmid, the resulting pSB1C3-OriKan plasmid was obtained (right part of the figure)]] | + | [[Image:Toulouse_France_backbone2.jpg|thumb|center|800px|<b>Figure 2:</b> creation of the OriKan cassette. Position of the primers are indicated by the blue arrow on pSB<sub>Bs</sub>0K-P (left part of the figure). The resulting PCR fragment is the blue pointed line. After digestion by EcoRI and PstI and ligation in the pSB1C3 plasmid, the resulting pSB1C3-OriKan plasmid was obtained (right part of the figure)]] |
<br> | <br> | ||
| Line 22: | Line 22: | ||
We successfully transformed <i>E. coli</i> and selected clones on chloramphenicol. Likewise, we successfully transformed <i>B. subtilis</i> and selected clones on kanamycine. Presence of the pSB1C3-OriKan plasmid was demonstrated in <i>B. subtilis</i> by PCR checking (Figure 3). | We successfully transformed <i>E. coli</i> and selected clones on chloramphenicol. Likewise, we successfully transformed <i>B. subtilis</i> and selected clones on kanamycine. Presence of the pSB1C3-OriKan plasmid was demonstrated in <i>B. subtilis</i> by PCR checking (Figure 3). | ||
<br> | <br> | ||
| − | [[Image:Toulouse_France_backbone3.jpg|thumb|center|800px|Figure 3: validation of the pSB1C3-Orikan presence in | + | [[Image:Toulouse_France_backbone3.jpg|thumb|center|800px|<b>Figure 3:</b> validation of the pSB1C3-Orikan presence in <i>B. subtilis.</i> PCR on colonies was performed using primers hybridizing in the kanamycine resistance gene and in the suffix. The colonies were issued from the transformation of B. subtilis by pSB1C3-Orikan (assays), by the pSB<sub>Bs</sub>0K-P plasmid (negative control), or from the transformation of E. coli by pSB1C3-Orikan (positive control).]] |
<br>The part sequence has been verified by sequencing. | <br>The part sequence has been verified by sequencing. | ||
| Line 29: | Line 29: | ||
<b>Interest:</b> | <b>Interest:</b> | ||
| − | This part can be sub-cloned in any | + | This part can be sub-cloned in any pSB1C3 plasmid to make it compatible with <i>B. subtilis</i>. It will therefore greatly simplified the use of <i>Bacillus subtilis</i> as a chassis and the registration of new <i>Bacillus</i>-aimed parts. |
Revision as of 19:44, 29 October 2016
Part: BBa_K1937002 (pSB1C3-OriKan)
(Chassis E. coli/B. subtilis, carrier plasmid pSB1C3) Length: 2510 bp
Background: While Bacillus subtilis is of huge interest for a growing number of iGEM projects, it is not easy to develop new parts as it is required that they registered after sub-cloning in the E. coli plasmid pSB1C3. During the iGEM Toulouse 2016 project, we had the idea to create a part that could turn any pSB1C3-based plasmid in a shuttle vector (E. coli/B. subtilis). This BioBrick is a part developed by the Toulouse 2016 iGEM team (http://2016.igem.org/Team:Toulouse_France) from the pSBBs0K-P (BBa_K823026) previously developed by the 2012 Munich iGEM team.
More information available at http://2016.igem.org/Team:Toulouse_France/Description
Creation:
The BBa_K1937002 part contains the repU origin of replication of B. subtilis and the kanamycin resistance gene for B. subtilis (Figure 1).
It was obtained by amplifying the repU-Kan region of pSBBs0K-P (BBa_K1351040 or BBa_K823026) using primers OriKan forward (5’ cacagaatcaggggataacgcaggaaagaaACATGTAGTTATAAGTGACTAAACAAATAACTAAATAGATGGG) and OriKan reverse (5’ gttcctggccttttgctggccttttgctcaACATGTCGCAAAATGGCCCGATTTAAG). The resulting fragment was sub-cloned in the pSB1C3 between the EcoRI and PstI restriction sites (Figure 2).
Validation:
We successfully transformed E. coli and selected clones on chloramphenicol. Likewise, we successfully transformed B. subtilis and selected clones on kanamycine. Presence of the pSB1C3-OriKan plasmid was demonstrated in B. subtilis by PCR checking (Figure 3).

The part sequence has been verified by sequencing.
More information available at http://2016.igem.org/Team:Toulouse_France/Description
Interest:
This part can be sub-cloned in any pSB1C3 plasmid to make it compatible with B. subtilis. It will therefore greatly simplified the use of Bacillus subtilis as a chassis and the registration of new Bacillus-aimed parts.
Sequence:
Annotation:
repU : 431-1435
Kan : 1595-2365

