Difference between revisions of "Part:BBa C0071"
| Line 15: | Line 15: | ||
Author: Yoshio Takata | Author: Yoshio Takata | ||
| − | + | summary of characterization: I. Improved Prhl by iGEM 2014 Tokyo_Tech team and characterize RhlR assay | |
| − | + | <span style="margin-left: 10px;">We Tokyo Tech 2016 simulated our final genetic circuits and found that the circuits did not work, because Prhl strength was too weak. (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). We therefore considered using the improved Prhl (<partinfo>BBa_K1529310</partinfo>,<partinfo>BBa_K1529310</partinfo>) established by Tokyo_Tech 2014, but we noticed that they were inappropriate for two reasons (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). Then, we decided to improve Prhl Promoter and obtain our original improved Prhl (included in <partinfo>BBa_K1949060</partinfo>) that suited our goal. | |
| − | <span style="margin-left: 10px;">We found that Prhl(RL) (<partinfo>BBa_K1529300</partinfo>) activity was weak and the expression level depended on LVA tag (Fig.1); LVA-tagged proteins are prone to be degraded by cellular proteases. Prhl(LR) (<partinfo>BBa_K1529310</partinfo>) activity was strong and unexpectedly reacted with C12 (crosstalk | + | <span style="margin-left: 10px;">Our purpose is to create strong Prhl for our final genetic circuits. |
| + | |||
| + | I. Improved Prhl by iGEM 2014 Tokyo_Tech team and characterize rhlR assay | ||
| + | |||
| + | ====I. Improved Prhl by iGEM 2014 Tokyo_Tech item and characterize RhlR assay==== | ||
| + | |||
| + | <span style="margin-left: 10px;">We found that Prhl(RL) (<partinfo>BBa_K1529300</partinfo>) activity was weak and the expression level depended on LVA tag (Fig.1); LVA-tagged proteins are prone to be degraded by cellular proteases. Prhl(LR) (<partinfo>BBa_K1529310</partinfo>) activity was strong and unexpectedly reacted with C12 (crosstalk). | ||
[[Image:prhl1.png|thumb|center|400px|Fig.1 Comparison of the Past improved Prhl and RhlR ]]<br> | [[Image:prhl1.png|thumb|center|400px|Fig.1 Comparison of the Past improved Prhl and RhlR ]]<br> | ||
| Line 27: | Line 33: | ||
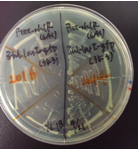
[[Image:prhl2.png|thumb|center|400px|Fig.2 The colonies of transformants with a rhlR (left) or a rhlR-LVA (right)]]<br> | [[Image:prhl2.png|thumb|center|400px|Fig.2 The colonies of transformants with a rhlR (left) or a rhlR-LVA (right)]]<br> | ||
| − | If you want more information, you see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki] | + | If you want more information, you see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]! |
| + | |||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
Revision as of 20:10, 19 October 2016
rhlR repressor/activator from P. aeruginosa PA3477 (+LVA)
Transcriptional regulator, in complex with N-butyryl-HSL, RhlR binds to the Rhl promoter
Usage and Biology
Transcriptional regulator, binds with N-butyryl-HSL
Characterization
Group: Tokyo Tech 2016
Author: Yoshio Takata
summary of characterization: I. Improved Prhl by iGEM 2014 Tokyo_Tech team and characterize RhlR assay
We Tokyo Tech 2016 simulated our final genetic circuits and found that the circuits did not work, because Prhl strength was too weak. (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). We therefore considered using the improved Prhl (BBa_K1529310,BBa_K1529310) established by Tokyo_Tech 2014, but we noticed that they were inappropriate for two reasons (see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]). Then, we decided to improve Prhl Promoter and obtain our original improved Prhl (included in BBa_K1949060) that suited our goal.
Our purpose is to create strong Prhl for our final genetic circuits.
I. Improved Prhl by iGEM 2014 Tokyo_Tech team and characterize rhlR assay
I. Improved Prhl by iGEM 2014 Tokyo_Tech item and characterize RhlR assay
We found that Prhl(RL) (BBa_K1529300) activity was weak and the expression level depended on LVA tag (Fig.1); LVA-tagged proteins are prone to be degraded by cellular proteases. Prhl(LR) (BBa_K1529310) activity was strong and unexpectedly reacted with C12 (crosstalk).
The colonies of transformants with a rhlR (BBa_C0171) plasmid looked rough and the growth rate was low(Fig.2-left), while the colonies of transformants with a rhlR-LVA (BBa_C0071) plasmid looked smooth and the growth rate was normal(Fig.2-right). However, the reason for this result is unclear.
If you want more information, you see [http://2016.igem.org/Team:Tokyo_Tech our work in Tokyo_Tech 2016 wiki]!
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 240
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 715