Difference between revisions of "Part:BBa K2065000"
Verakoomen (Talk | contribs) |
|||
| (79 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
| + | <html> | ||
<style> | <style> | ||
th, td { | th, td { | ||
| Line 12: | Line 13: | ||
} | } | ||
</style> | </style> | ||
| + | </html> | ||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K2065000 short</partinfo> | <partinfo>BBa_K2065000 short</partinfo> | ||
| − | + | <p> | |
| − | The | + | The CT52-SmallBiT BioBrick is part of a split luciferase system. This luciferase system consists of two parts, namely SmallBiT and LargeBiT. It is an ATP-independent luciferase, that produces high intensity luminescence. The total weight of LargeBiT-CT52 is 9,3 kDa. The NanoLuc system consists of 93 amino acids. When they dimerize, this means when they come into close proximity of each other, a substrate can create bioluminescence. The emission peak of this NanoBiT system is around 460 nanometers. The CT52 part of this BioBrick has affinity for the T14-3-3 scaffold protein. The binding between CT52 and T14-3-3 is mediated by the small molecule fusicoccin. Together with CT52-LargeBiT "https://parts.igem.org/Part:BBa_K2065007" this BioBrick is used in testing the quality of T14-3-3 heterodimers. |
| − | + | </p> | |
===Usage and Biology=== | ===Usage and Biology=== | ||
<p> | <p> | ||
| − | SmallBiT is linked to CT52 with a GGS10-linker, so it actually consists of three subparts. SmallBiT is 33 basepairs long and thus consists of 11 amino acids. The GGS10-linker that is used, consists of 30 amino acids | + | SmallBiT is linked to CT52 with a GGS10-linker, so it actually consists of three subparts. SmallBiT is 33 basepairs long and thus consists of 11 amino acids. The GGS10-linker that is used, consists of 30 amino acids: it is a ten-times repeated sequence of glycine-glycine-serine amino acids. This complex is linked to CT52. |
| − | This protein | + | This protein consists of the last 52 amino acids of the C-terminal region of the H+-ATPase. The N-terminus of CT52 is a free end, which can be used to link proteins to. The binding between CT52 and T14-3-3 is stabilized by the small molecule Fusicoccin. When fusicoccin is present, the affinity of CT52 for T14-3-3 is increased in 30 fold with respect to the situation when fusicoccin is not present[1]. The binding interaction with fusicoccin is reversibel, this means CT52 will dissociate from the scaffold. |
</p> | </p> | ||
<p> | <p> | ||
| − | This means that the CT52 -SmallBiT complex will move towards the scaffold protein, after | + | This means that the CT52-SmallBiT complex will move towards the scaffold protein, after the small molecule fusicoccin, is added. |
Three different variants of the CT52-protein were created with the help of the Rosetta software package. Each mutation in the CT52 protein has its complementary mutation in the T14-3-3 protein. Because these mutations were found with the help of the Rosetta software package, the quality of these mutations have to be verified. | Three different variants of the CT52-protein were created with the help of the Rosetta software package. Each mutation in the CT52 protein has its complementary mutation in the T14-3-3 protein. Because these mutations were found with the help of the Rosetta software package, the quality of these mutations have to be verified. | ||
| − | One variant of the | + | One variant of the CT52 protein was already known and tested. This mutation functioned as a positive control in the verification process of the other mutations. |
</p> | </p> | ||
| − | |||
===Sequence=== | ===Sequence=== | ||
| + | The sequence of our CT52-SmallBiT has been verified by StarSeq. It contains the prefix and suffix with the correct restriction sites (EcoRI, XbaI, SpeI and PstI). CT52-smallBiT is 279 basepairs long. | ||
| + | [[File:T--TU-Eindhoven--SmallBitSnap.png]] | ||
| + | <br> | ||
| + | ''Figure 1: Snapgene map BBa_K2065000'' | ||
| Line 35: | Line 40: | ||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
<partinfo>BBa_K2065000 SequenceAndFeatures</partinfo> | <partinfo>BBa_K2065000 SequenceAndFeatures</partinfo> | ||
| − | |||
| − | |||
| − | |||
===Bioluminescence confirmation=== | ===Bioluminescence confirmation=== | ||
<p> | <p> | ||
| − | To determine whether the split luciferase system is working, | + | To determine whether the split luciferase system is working, a bioluminescence signal should be visible when the two components come into close proximity of each other. A bioluminescence peak around 460nm must be visible when SmallBiT and LargeBiT dimerize. When the two NanoBiTs are into close proximity of each other, they dimerize. When Furimazine is added to the dimerized Nanoluc luciferase, the Furimazine is converted into Furimamide. When this reaction occurs, light with a wavelength of 460 nm will be emitted. The spectrum of the CT52-NanoBiT system is visualed below. |
| + | |||
| + | [[File:T--TU-Eindhoven--LumPeak.png]] | ||
| + | <br> | ||
| + | ''Figure 2: Bioluminescence spectrum with CT52(I947H)-LargeBiT and CT52-SmallBiT, assembled on T14-3-3(S71L) - T14-3-3'' | ||
</p> | </p> | ||
<p> | <p> | ||
| − | + | An example of a measurement which can be done on a heterodimer is a T14-3-3 heterodimer concentration gradient assay, in which the functionality can be verified. This measurement is shown below for T14-3-3(S71L) - T14-3-3. More measurements on T14-3-3 heterodimer scaffold with the CT52-NanoBiT system can be found on our wiki "http://2016.igem.org/Team:TU-Eindhoven/Results". | |
| − | + | ||
| + | [[File:T--TU-Eindhoven--SLGRAD.png]] | ||
| + | <br> | ||
| + | ''Figure 3: T14-3-3 functionality measurement with varying scaffold concentrations from 0 nM to 1 µM T14-3-3(S71L/I72V)-T14-3-3. 50 µM fusicoccin was used for all measurements. Mutant forms CT52 (I947F)-LargeBiT and CT52(wildtype)-SmallBiT were used with a concentration of 200 nM. The substrate used was 1250x diluted from stock.'' | ||
| + | </p> | ||
| + | |||
| + | ===NanoBiT calibration=== | ||
| + | In order to analyse the performance of our CT52 fused NanoBiT fragments we measured the luminescence they produce at varying concentrations from 10nM to 10mM. | ||
| + | After global optimization using metropolis Monte Carlo and Latin hypercube sampling the best fit resulted in LUmax = 3.5* 105 ±1.5 a.u. and KD = 996±163 μM (see figure 4), standard deviations are calculated with the Jacobian. | ||
| + | <br> | ||
| + | [[File:T--TU-Eindhoven--NANOactBB.png]] | ||
| + | <br> | ||
| + | ''Figure 4: The luminescence of NanoBiT as a function of the concentration of LargeBiT. The concentration of SmallBiT is kept constant at 100 nM. The circles represent the measured datapoints, the errorbars represent the standard deviation of the measures (n = 2). The red circle and errorbars represent the dissociation constant and its standard deviation respectively.'' | ||
| − | === | + | ===CT52-SmallBiT Statistics=== |
| − | <table> | + | <table style="border:black 1px solid;"> |
<tr> | <tr> | ||
| − | <th style="height:40px;" colspan="4"> Protein Specifications</th> | + | <th style="height:40px;" colspan="4"> Protein Specifications CT52-SmallBiT</th> |
</tr> | </tr> | ||
<tr> | <tr> | ||
| Line 199: | Line 216: | ||
| − | + | ==References== | |
| + | [1] - Milroy, L., Brunsveld, L., & Ottmann, C. (2013). Stabilization and Inhibition of Protein–Protein Interactions: The 14-3-3 Case Study. ACS Chem. Biol., 8(1),27-35. | ||
<!-- Uncomment this to enable Functional Parameter display | <!-- Uncomment this to enable Functional Parameter display | ||
Latest revision as of 19:19, 19 October 2016
CT52-SmallBiT
The CT52-SmallBiT BioBrick is part of a split luciferase system. This luciferase system consists of two parts, namely SmallBiT and LargeBiT. It is an ATP-independent luciferase, that produces high intensity luminescence. The total weight of LargeBiT-CT52 is 9,3 kDa. The NanoLuc system consists of 93 amino acids. When they dimerize, this means when they come into close proximity of each other, a substrate can create bioluminescence. The emission peak of this NanoBiT system is around 460 nanometers. The CT52 part of this BioBrick has affinity for the T14-3-3 scaffold protein. The binding between CT52 and T14-3-3 is mediated by the small molecule fusicoccin. Together with CT52-LargeBiT "https://parts.igem.org/Part:BBa_K2065007" this BioBrick is used in testing the quality of T14-3-3 heterodimers.
Usage and Biology
SmallBiT is linked to CT52 with a GGS10-linker, so it actually consists of three subparts. SmallBiT is 33 basepairs long and thus consists of 11 amino acids. The GGS10-linker that is used, consists of 30 amino acids: it is a ten-times repeated sequence of glycine-glycine-serine amino acids. This complex is linked to CT52. This protein consists of the last 52 amino acids of the C-terminal region of the H+-ATPase. The N-terminus of CT52 is a free end, which can be used to link proteins to. The binding between CT52 and T14-3-3 is stabilized by the small molecule Fusicoccin. When fusicoccin is present, the affinity of CT52 for T14-3-3 is increased in 30 fold with respect to the situation when fusicoccin is not present[1]. The binding interaction with fusicoccin is reversibel, this means CT52 will dissociate from the scaffold.
This means that the CT52-SmallBiT complex will move towards the scaffold protein, after the small molecule fusicoccin, is added. Three different variants of the CT52-protein were created with the help of the Rosetta software package. Each mutation in the CT52 protein has its complementary mutation in the T14-3-3 protein. Because these mutations were found with the help of the Rosetta software package, the quality of these mutations have to be verified. One variant of the CT52 protein was already known and tested. This mutation functioned as a positive control in the verification process of the other mutations.
Sequence
The sequence of our CT52-SmallBiT has been verified by StarSeq. It contains the prefix and suffix with the correct restriction sites (EcoRI, XbaI, SpeI and PstI). CT52-smallBiT is 279 basepairs long.
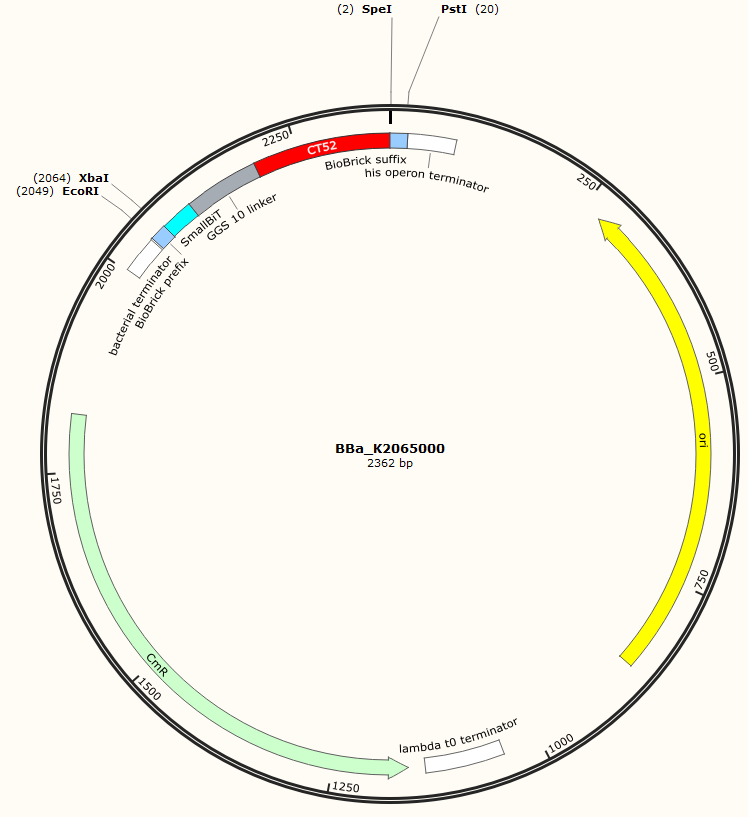
Figure 1: Snapgene map BBa_K2065000
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 289
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Bioluminescence confirmation
To determine whether the split luciferase system is working, a bioluminescence signal should be visible when the two components come into close proximity of each other. A bioluminescence peak around 460nm must be visible when SmallBiT and LargeBiT dimerize. When the two NanoBiTs are into close proximity of each other, they dimerize. When Furimazine is added to the dimerized Nanoluc luciferase, the Furimazine is converted into Furimamide. When this reaction occurs, light with a wavelength of 460 nm will be emitted. The spectrum of the CT52-NanoBiT system is visualed below.
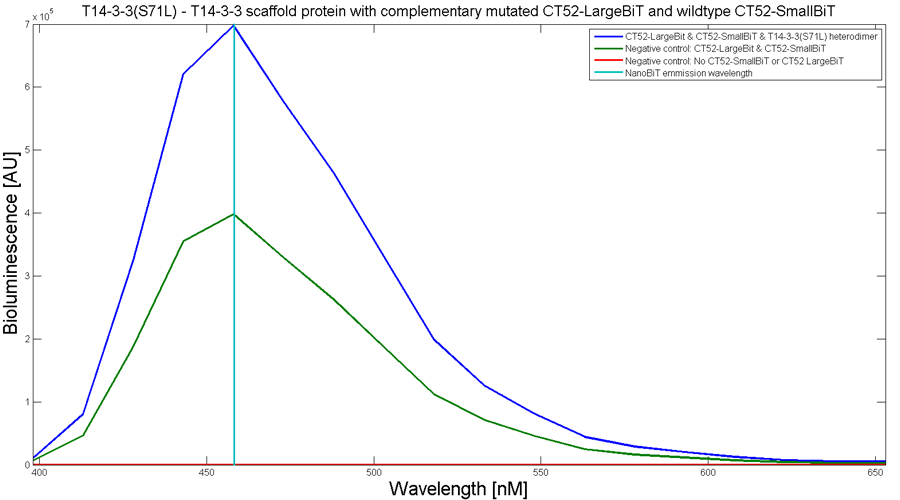
Figure 2: Bioluminescence spectrum with CT52(I947H)-LargeBiT and CT52-SmallBiT, assembled on T14-3-3(S71L) - T14-3-3
An example of a measurement which can be done on a heterodimer is a T14-3-3 heterodimer concentration gradient assay, in which the functionality can be verified. This measurement is shown below for T14-3-3(S71L) - T14-3-3. More measurements on T14-3-3 heterodimer scaffold with the CT52-NanoBiT system can be found on our wiki "http://2016.igem.org/Team:TU-Eindhoven/Results".
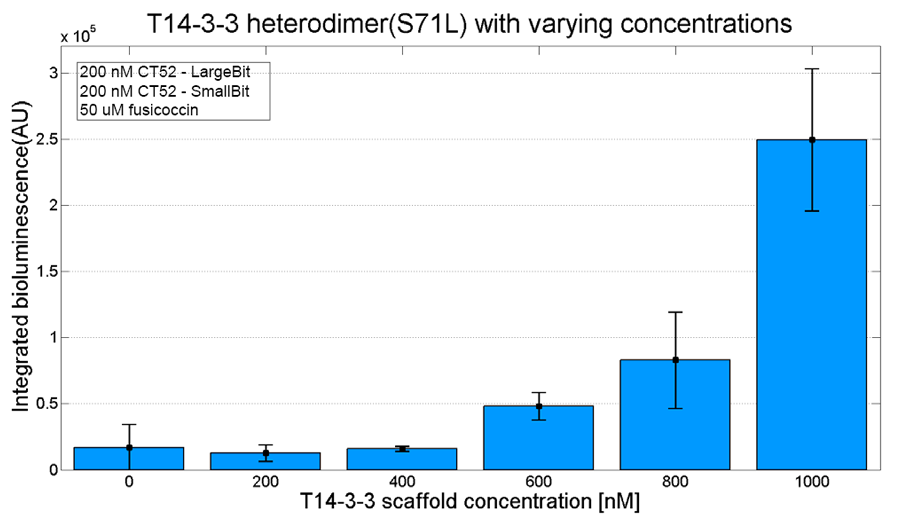
Figure 3: T14-3-3 functionality measurement with varying scaffold concentrations from 0 nM to 1 µM T14-3-3(S71L/I72V)-T14-3-3. 50 µM fusicoccin was used for all measurements. Mutant forms CT52 (I947F)-LargeBiT and CT52(wildtype)-SmallBiT were used with a concentration of 200 nM. The substrate used was 1250x diluted from stock.
NanoBiT calibration
In order to analyse the performance of our CT52 fused NanoBiT fragments we measured the luminescence they produce at varying concentrations from 10nM to 10mM.
After global optimization using metropolis Monte Carlo and Latin hypercube sampling the best fit resulted in LUmax = 3.5* 105 ±1.5 a.u. and KD = 996±163 μM (see figure 4), standard deviations are calculated with the Jacobian.
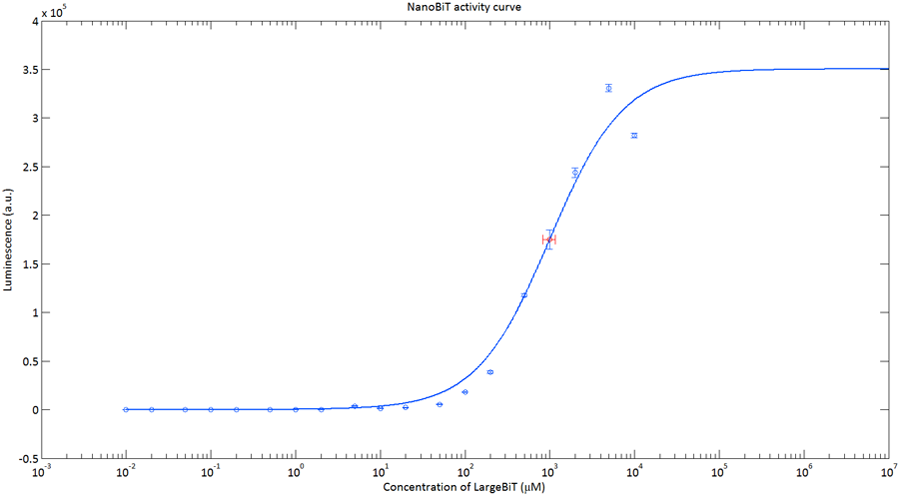
Figure 4: The luminescence of NanoBiT as a function of the concentration of LargeBiT. The concentration of SmallBiT is kept constant at 100 nM. The circles represent the measured datapoints, the errorbars represent the standard deviation of the measures (n = 2). The red circle and errorbars represent the dissociation constant and its standard deviation respectively.
CT52-SmallBiT Statistics
| Protein Specifications CT52-SmallBiT | |||
|---|---|---|---|
| General Information | Number of amino acids | 93 | |
| Molecular weight | 9367.169 | ||
| Theoretical pi | 5.62 | ||
| Extinction coefficient | 2980 | ||
| Formula | C395H639N123O141 | ||
| Total numbers of atons | 1298 | ||
| Amino Acid Composition | Amino Acid | Frequency | Percentage(%) |
| Ala(A) | 4 | 4.3 | |
| Arg(R | 5 | 5.4 | |
| Asn(N) | 3 | 3.2 | |
| Asp(D) | 2 | 2.2 | |
| Cys(C) | 0 | 0 | |
| Gln(Q) | 4 | 4.3 | |
| Glu(E) | 9 | 9.7 | |
| Gly(G) | 23 | 24.7 | |
| His(H) | 2 | 2.2 | |
| Ile(I) | 5 | 5.4 | |
| Leu(L) | 8 | 8.6 | |
| Lys(K) | 4 | 4.3 | |
| Met(M) | 0 | 0 | |
| Phe(F) | 2 | 2.2 | |
| Pro(P) | 0 | 0 | |
| Ser(S) | 12 | 12.9 | |
| Thr(T) | 4 | 4.3 | |
| Trp(W) | 0 | 0 | |
| Tyr(T) | 2 | 2.2 | |
| Val(V) | 4 | 4.3 | |
| Pyl(O) | 0 | 0.0 | |
| Sec(U) | 0 | 0.0 | |
References
[1] - Milroy, L., Brunsveld, L., & Ottmann, C. (2013). Stabilization and Inhibition of Protein–Protein Interactions: The 14-3-3 Case Study. ACS Chem. Biol., 8(1),27-35.
