Difference between revisions of "Part:BBa K1172904"
(→Barnase) |
(→Barnase) |
||
| Line 6: | Line 6: | ||
==='''Barnase'''=== | ==='''Barnase'''=== | ||
<br> | <br> | ||
| − | [[Image:IGEM Bielefeld 2013 biosafety RNase Ba test.png| | + | [[Image:IGEM Bielefeld 2013 biosafety RNase Ba test.png|3000px|left]] |
<p align="justify"> | <p align="justify"> | ||
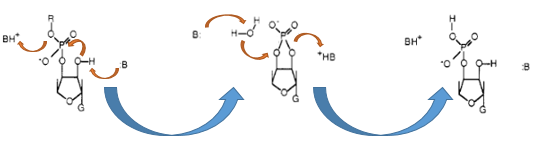
The Barnase (EC 3.1.27) is an 12 kDa extracellular microbial ribonuclease, which is naturally found in the gram-positive soil bacteria ''Bacillus amyloliquefaciens'' and consist a single chain of 110 amino acids. The Barnase (RNase Ba) catalyses the cleavage of single stranded RNA, where the hydrolysis of the dinucleotides has the highest affinity to the structure GpN. In the first step of the RNA-degradation a cyclic intermediate is formed by transesterification and afterwards this intermediate is hydrolysed yielding in a 3'-nucleotide ([http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Mossakowska ''et al.'', 1989]).</p> | The Barnase (EC 3.1.27) is an 12 kDa extracellular microbial ribonuclease, which is naturally found in the gram-positive soil bacteria ''Bacillus amyloliquefaciens'' and consist a single chain of 110 amino acids. The Barnase (RNase Ba) catalyses the cleavage of single stranded RNA, where the hydrolysis of the dinucleotides has the highest affinity to the structure GpN. In the first step of the RNA-degradation a cyclic intermediate is formed by transesterification and afterwards this intermediate is hydrolysed yielding in a 3'-nucleotide ([http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Mossakowska ''et al.'', 1989]).</p> | ||
Revision as of 05:00, 5 October 2013
Rnase Ba (Barnase) from Bacillus amyloliquefaciens
Usage and Biology
Barnase
The Barnase (EC 3.1.27) is an 12 kDa extracellular microbial ribonuclease, which is naturally found in the gram-positive soil bacteria Bacillus amyloliquefaciens and consist a single chain of 110 amino acids. The Barnase (RNase Ba) catalyses the cleavage of single stranded RNA, where the hydrolysis of the dinucleotides has the highest affinity to the structure GpN. In the first step of the RNA-degradation a cyclic intermediate is formed by transesterification and afterwards this intermediate is hydrolysed yielding in a 3'-nucleotide ([http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Mossakowska et al., 1989]).
In Bacillus amyloliquefaciens the activity ot the Barnase (RNase Ba) is inhibited intracelluar by the Inhibitor called barstar. Barstar consists only about 89 amino acids and binds with a high affinity to the toxic Barnase. This prevents the cleavage of the intracellular RNA in the host organism ([http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Paddon et al., 1989]). Therefore the Barnase acts naturally only outside the cell and is translocated under natural conditions. For the Biosafety-System we tried to modified this aspect by cloning only the sequence responsible for the cleavage of the RNA, but not the part of the native Barnase, which is essential for the extracellular transport.
As shown in the graphic below, the transcription of the DNA, which encodes the Barnase produces a 474 nt RNA. The translation of the RNA of this ribonuclease starts about 25 nucleotides downstream from the transcription start and can be divided into two parts. The first part (colored in orange) is translated into a signal peptide at the N-Terminus of the Barnase. This part is responsible for the extracellular translocation of the RNase Ba, while the peptide sequence for the active Barnase starts 142 nucleotides downstream from the transcription start (colored in red).
For the Biosafety-System we used only the coding sequence (BBa_K1172904) of the Barnase itself to prevent the extracellular translocation of the toxic gene product. This leads to a rapid cell death if the expression of the Barnase isn't repressed by the repressor of the Biosafety-System.
600px|thumb|center|Figure 9: Sequence of the Signalpeptide in front of the RNase Ba (Barnase). The Biobrick BBa_K1172904 does not consist the signal sequence for the extracellular translocation, but only the coding sequence for the mature enzyme.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]