Difference between revisions of "Part:BBa K1122676"
| Line 4: | Line 4: | ||
This parts enables expression of pyruvate decarboxylase (pdc) and alcohol dehydrogenase B (adhB) using IPTG induction. Those enzymes are involved in conversion of pyruvate to ethanol via an acetaldehyde intermediate. The pdc and adhB sequences come from ''Zymomonas mobilis'' however, they are codon optimised for ''E. coli''. | This parts enables expression of pyruvate decarboxylase (pdc) and alcohol dehydrogenase B (adhB) using IPTG induction. Those enzymes are involved in conversion of pyruvate to ethanol via an acetaldehyde intermediate. The pdc and adhB sequences come from ''Zymomonas mobilis'' however, they are codon optimised for ''E. coli''. | ||
| − | + | ||
===Usage and Biology=== | ===Usage and Biology=== | ||
| − | + | Microbial production of ethanol is of great importance due to its possible application as a biofuel. Increasing ethanol yields in bacteria is potentially beneficial as those are able to utilise a wider variety of renewable, biomass-derived carbon sources compared to standard ethanol producer: ''Sacharomyces cerevisiae''. | |
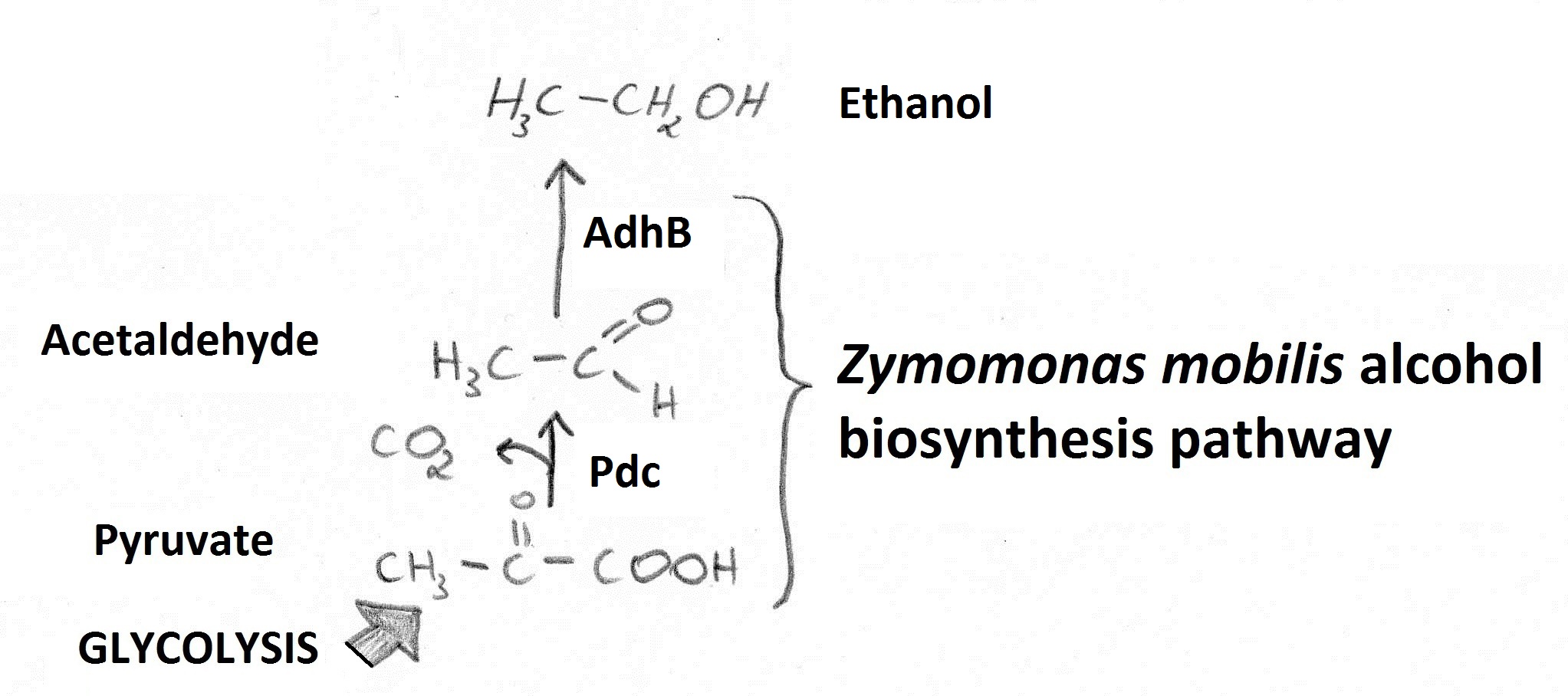
| + | Production of ethanol in ''E. coli'' was achieved by expression of ''Zymomonas mobilis'' pyruvate decarboxylase (Pdc) and alcohol dehydrogenase B (AdhB)(Ingram et al.,.1987) . Reactions catalysed by those enzymes (Figure A) enable conversion of pyruvate to ethanol. | ||
| + | |||
| + | [[File:bioethanol_introduction_1.jpg|700px]] | ||
| + | |||
| + | '''Figure A.''' Ethanol is generated from pyruvate (fed into the pathway from glycolysis) via an Acetaldehyde intermediate. | ||
| + | |||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
<partinfo>BBa_K1122676 SequenceAndFeatures</partinfo> | <partinfo>BBa_K1122676 SequenceAndFeatures</partinfo> | ||
Revision as of 19:22, 4 October 2013
IPTG inducible pdc and adhB
This parts enables expression of pyruvate decarboxylase (pdc) and alcohol dehydrogenase B (adhB) using IPTG induction. Those enzymes are involved in conversion of pyruvate to ethanol via an acetaldehyde intermediate. The pdc and adhB sequences come from Zymomonas mobilis however, they are codon optimised for E. coli.
Usage and Biology
Microbial production of ethanol is of great importance due to its possible application as a biofuel. Increasing ethanol yields in bacteria is potentially beneficial as those are able to utilise a wider variety of renewable, biomass-derived carbon sources compared to standard ethanol producer: Sacharomyces cerevisiae. Production of ethanol in E. coli was achieved by expression of Zymomonas mobilis pyruvate decarboxylase (Pdc) and alcohol dehydrogenase B (AdhB)(Ingram et al.,.1987) . Reactions catalysed by those enzymes (Figure A) enable conversion of pyruvate to ethanol.
Figure A. Ethanol is generated from pyruvate (fed into the pathway from glycolysis) via an Acetaldehyde intermediate.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 1109
Illegal AgeI site found at 2315 - 1000COMPATIBLE WITH RFC[1000]
References
INGRAM, L. O., CONWAY, T., CLARK, D. P., SEWELL, G. W. & PRESTON, J. F. 1987. GENETIC-ENGINEERING OF ETHANOL-PRODUCTION IN ESCHERICHIA-COLI. Applied and Environmental Microbiology, 53, 2420-2425.