Difference between revisions of "Part:BBa K771404"
AleAlejandro (Talk | contribs) |
|||
| Line 1: | Line 1: | ||
| − | |||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K771404 short</partinfo> | <partinfo>BBa_K771404 short</partinfo> | ||
| − | + | ||
| + | |||
| + | Membrane anchor 4, which consists of interacting PDZ domain([https://parts.igem.org/wiki/index.php?title=Part:BBa_K771103 BBa_K771103]), GBD domain([https://parts.igem.org/wiki/index.php?title=Part:BBa_K771105 BBa_K771105]) and membrane protein Lgt([https://parts.igem.org/wiki/index.php?title=Part:BBa_K771102 BBa_K771102]). Membrane Anchor 4 is proved to constitutively aggregate with Membrane Anchor 3 and 5(with and without VVD). | ||
| + | |||
| + | |||
| + | To testify the constitutive aggregation of Membrane Anchor 3, 4 and 5, we conducted Fluorescence Complementation Assay. | ||
| + | |||
| + | In fluorescence complementation assay, proteins that are postulated to interact are fused to unfolded complementary fragments of EGFP and expressed in ''E.coli''. Interaction of these proteins will bring the fluorescent fragments within proximity, allowing the reporter protein to reform in its native three-dimensional structure and emit its fluorescent signal. EGFP was split into two halves, named 1EGFP([https://parts.igem.org/wiki/index.php?title=Part:BBa_K771113 BBa_K771113]) and 2EGFP([https://parts.igem.org/wiki/index.php?title=Part:BBa_K771114 BBa_K771114]) respectively. If there is interaction between two proteins which were fused with 1EGFP and 2EGFP, it is expected that fluorescence should be observed. Otherwise, no fluorescence could be observed. | ||
| + | |||
| + | ===Membrane Anchor 3 and Membrane Anchor 4=== | ||
| + | |||
| + | |||
| + | We fused split EGFP 1 and 2 to Membrane Anchor 4 and Membrane Anchor 3 to test whether Membrane Anchor 3 and Membrane Anchor 4 could constitutively aggregate. Proteins in control group are not expected to aggregate like in experimental group.We coexpressed proteins in experimental group and control group respectively in ''E.coli''. After induction for 6 hours, bacteria samples were taken for fluorescence observationThe result shows that Membrane Anchor 3 and Membrane Anchor 4 could constitutively aggregate through PDZ domain and ligand. | ||
| + | [[Image:12SJTU_splitegfp34.png|400px|center]] | ||
| + | [[Image:12SJTU Anchor GFP.jpg|450px|center]] | ||
| + | <br\><br\><br\> | ||
| + | |||
| + | ===Membrane Anchor 4 and Membrane Anchor 5 without VVD=== | ||
| + | [[Image:12SJTU_splitegfp45.png|400px|center]] | ||
| + | |||
| + | [[Image:12SJTU Standard GFP 2.jpg|400px|center]] | ||
| + | We fused split EGFP 1 and 2 to Membrane Anchor 4 and Membrane Anchor 5 without VVD to test whether Membrane Anchor 4 and Membrane Anchor 5 without VVD could constitutively aggregate. Proteins in control group are not expected to aggregate like in experimental group.We coexpressed proteins in experimental group and control group respectively in ''E.coli''. After induction for 6 hours, bacteria samples were taken for fluorescence observationThe result shows that Membrane Anchor 4 and Membrane Anchor 5 without VVD could constitutively aggregate through GBD domain and ligand. | ||
| + | |||
| + | |||
| + | |||
| + | |||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
Latest revision as of 16:25, 2 October 2012
split EGFP 1 with Membrane Anchor 4
Membrane anchor 4, which consists of interacting PDZ domain(BBa_K771103), GBD domain(BBa_K771105) and membrane protein Lgt(BBa_K771102). Membrane Anchor 4 is proved to constitutively aggregate with Membrane Anchor 3 and 5(with and without VVD).
To testify the constitutive aggregation of Membrane Anchor 3, 4 and 5, we conducted Fluorescence Complementation Assay.
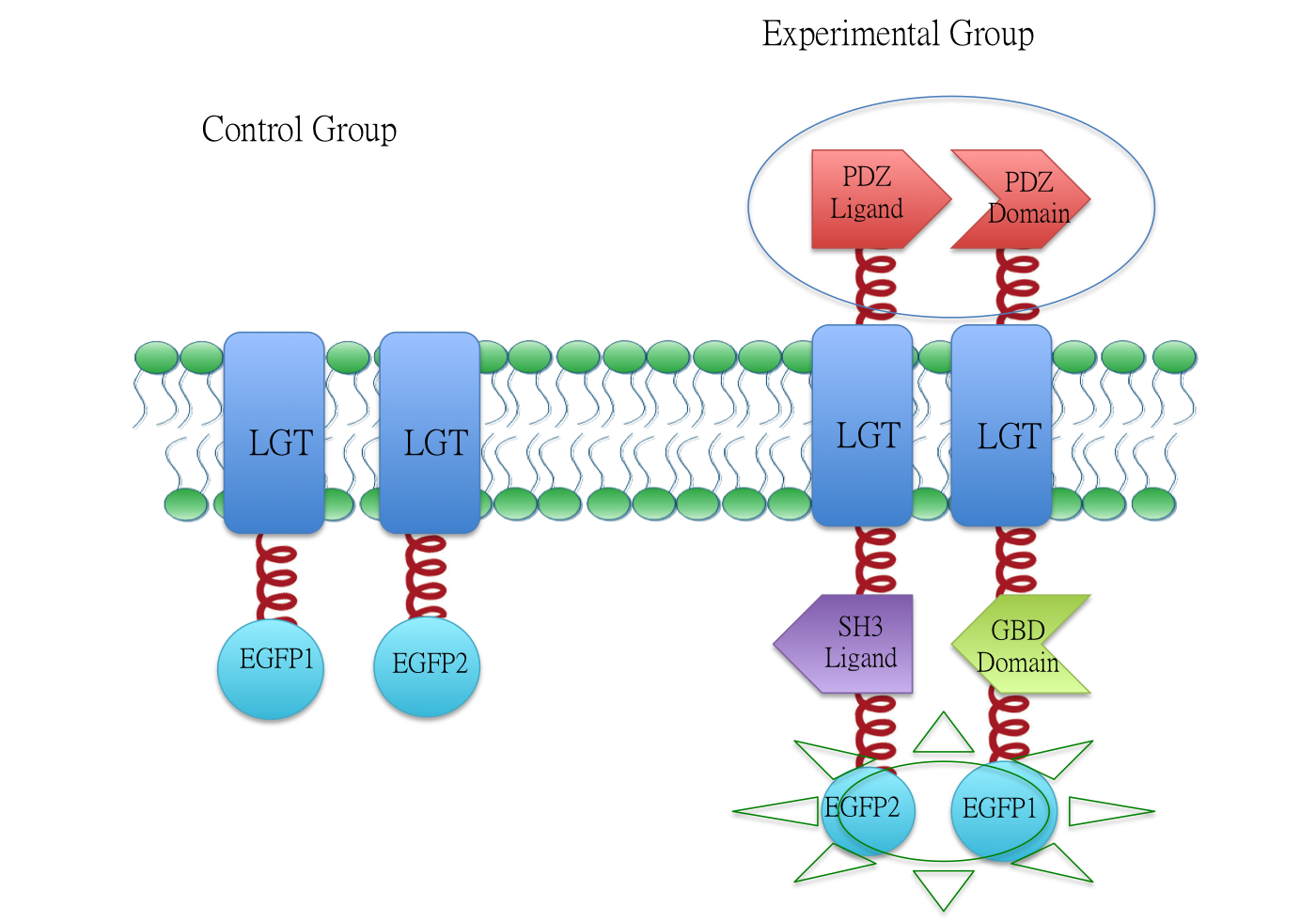
In fluorescence complementation assay, proteins that are postulated to interact are fused to unfolded complementary fragments of EGFP and expressed in E.coli. Interaction of these proteins will bring the fluorescent fragments within proximity, allowing the reporter protein to reform in its native three-dimensional structure and emit its fluorescent signal. EGFP was split into two halves, named 1EGFP(BBa_K771113) and 2EGFP(BBa_K771114) respectively. If there is interaction between two proteins which were fused with 1EGFP and 2EGFP, it is expected that fluorescence should be observed. Otherwise, no fluorescence could be observed.
Membrane Anchor 3 and Membrane Anchor 4
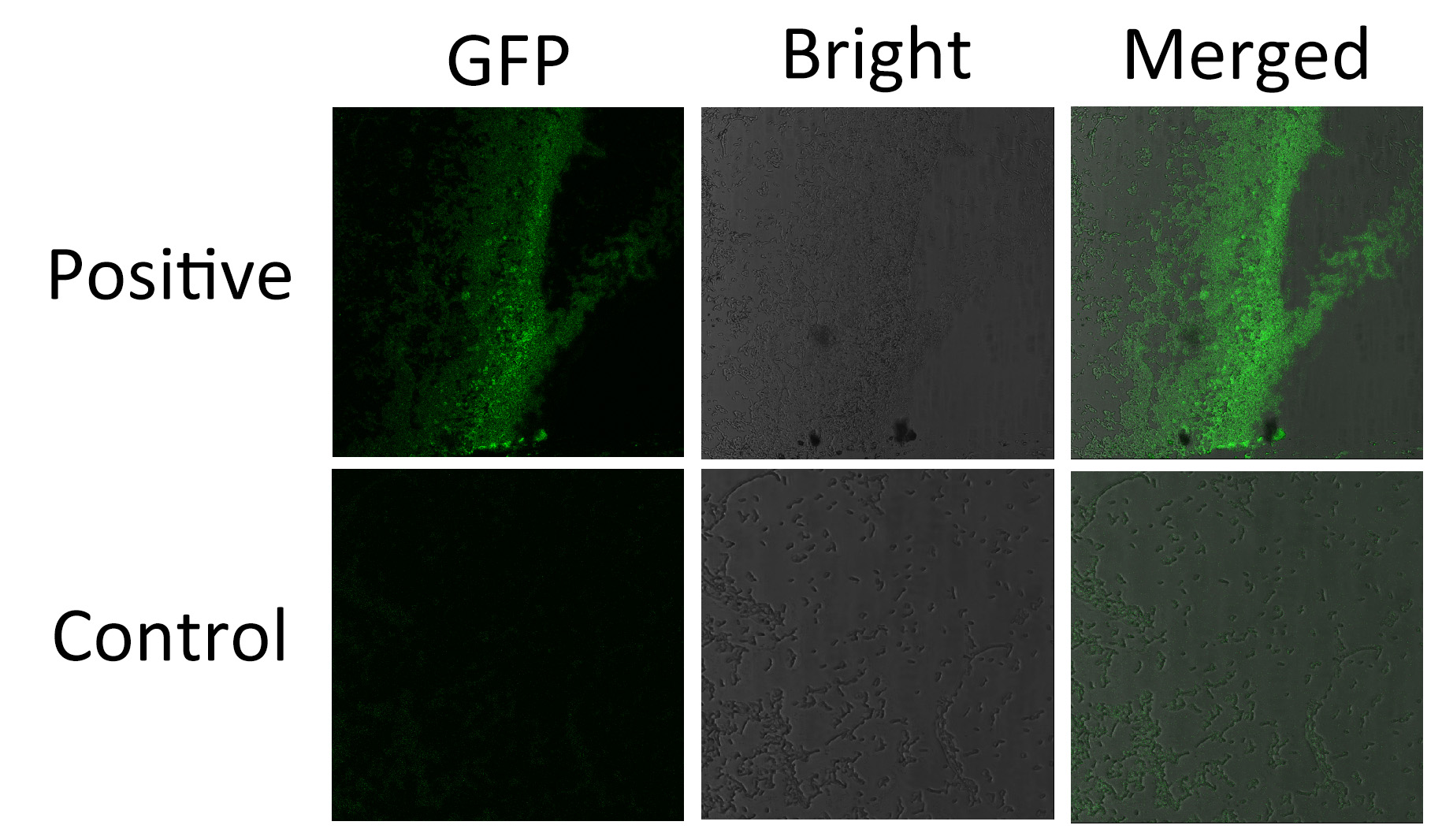
We fused split EGFP 1 and 2 to Membrane Anchor 4 and Membrane Anchor 3 to test whether Membrane Anchor 3 and Membrane Anchor 4 could constitutively aggregate. Proteins in control group are not expected to aggregate like in experimental group.We coexpressed proteins in experimental group and control group respectively in E.coli. After induction for 6 hours, bacteria samples were taken for fluorescence observationThe result shows that Membrane Anchor 3 and Membrane Anchor 4 could constitutively aggregate through PDZ domain and ligand.
<br\><br\><br\>
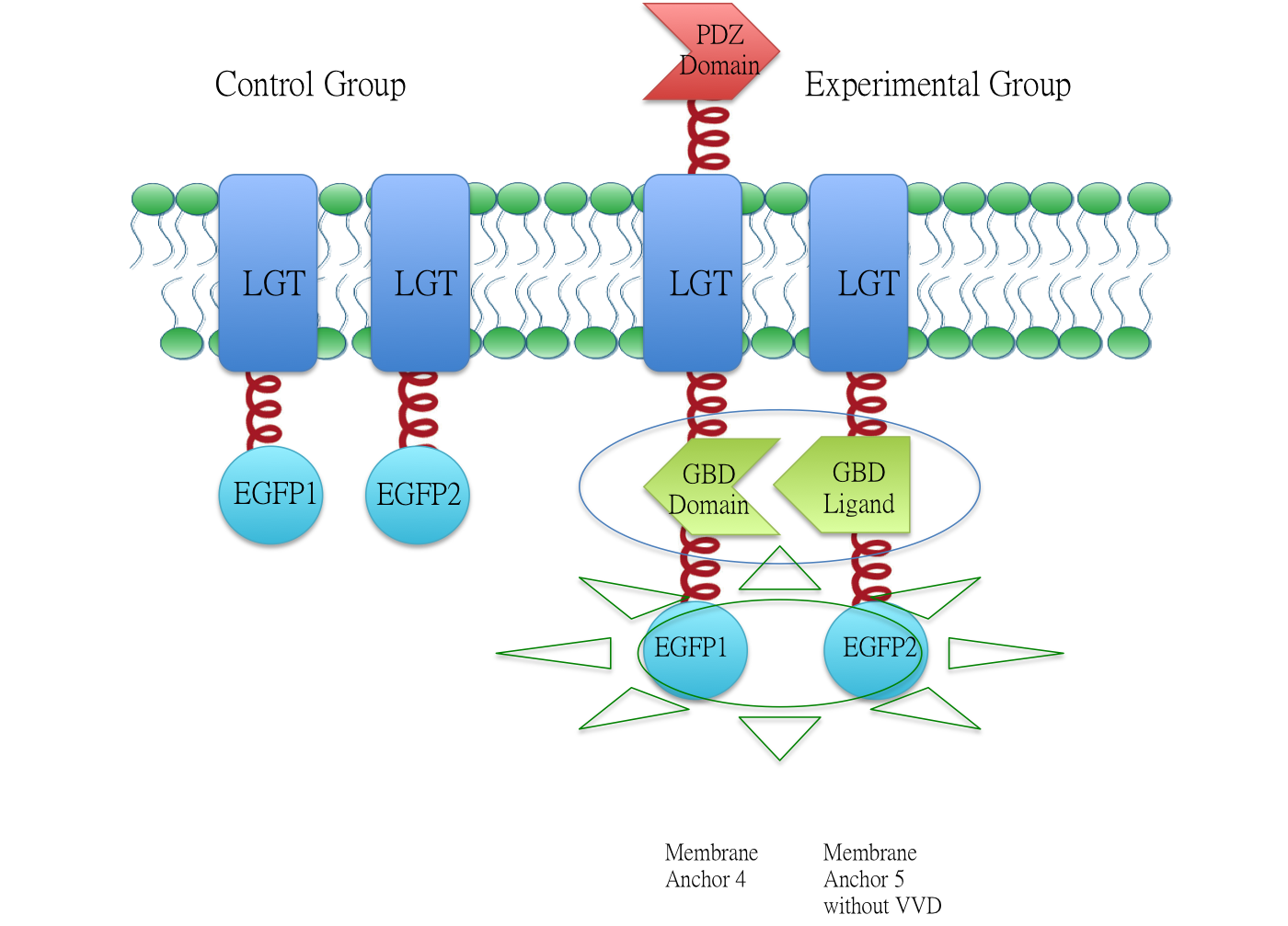
Membrane Anchor 4 and Membrane Anchor 5 without VVD
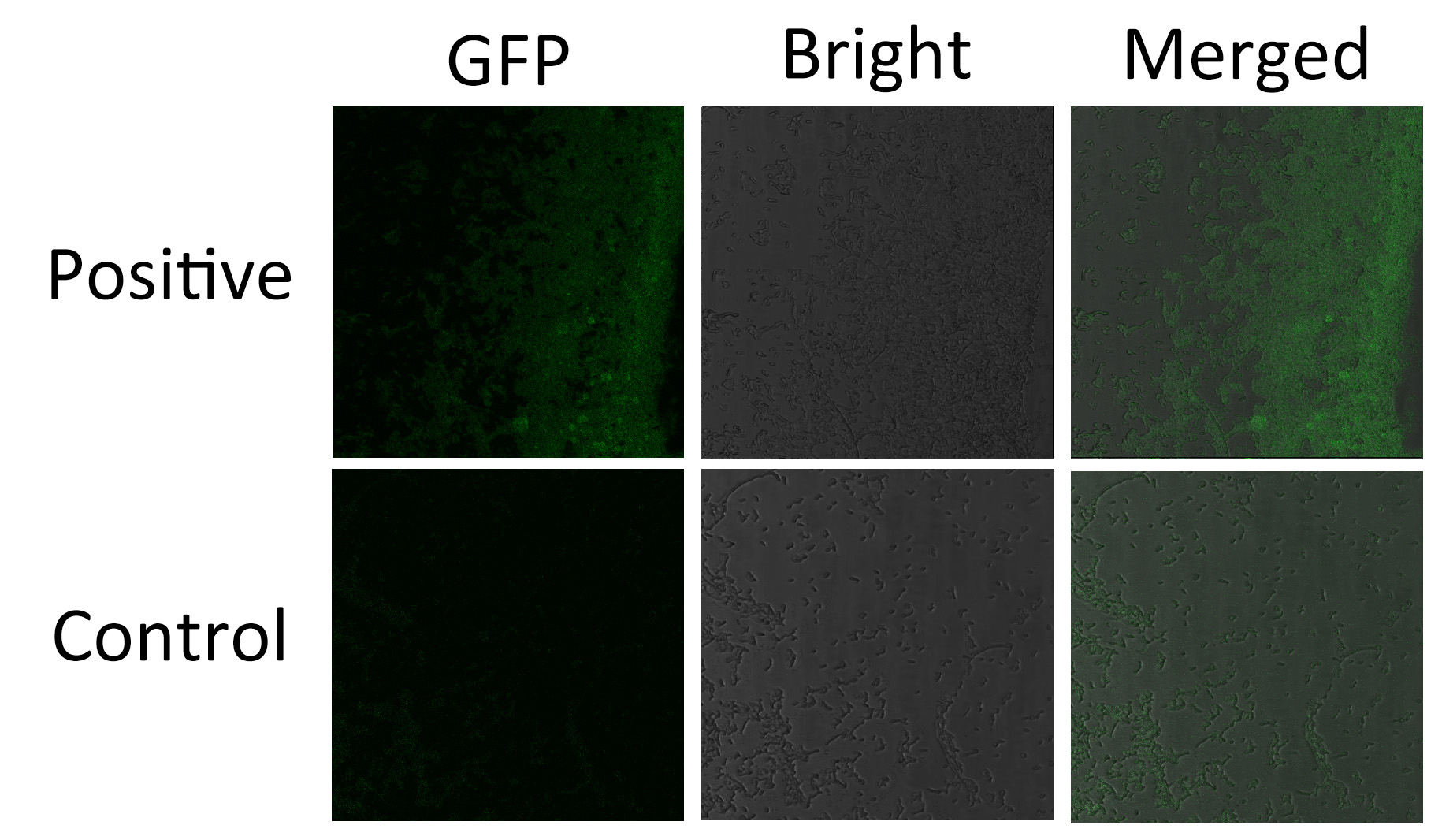
We fused split EGFP 1 and 2 to Membrane Anchor 4 and Membrane Anchor 5 without VVD to test whether Membrane Anchor 4 and Membrane Anchor 5 without VVD could constitutively aggregate. Proteins in control group are not expected to aggregate like in experimental group.We coexpressed proteins in experimental group and control group respectively in E.coli. After induction for 6 hours, bacteria samples were taken for fluorescence observationThe result shows that Membrane Anchor 4 and Membrane Anchor 5 without VVD could constitutively aggregate through GBD domain and ligand.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 135
Illegal AgeI site found at 1194 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 365
Illegal BsaI site found at 815