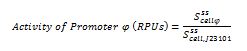Difference between revisions of "Part:BBa J23111:Experience"
Ao.patricia (Talk | contribs) (→User Reviews) |
|||
| Line 1: | Line 1: | ||
| − | |||
__NOTOC__ | __NOTOC__ | ||
This experience page is provided so that any user may enter their experience using this part.<BR>Please enter | This experience page is provided so that any user may enter their experience using this part.<BR>Please enter | ||
| Line 19: | Line 18: | ||
<!-- End of the user review template --> | <!-- End of the user review template --> | ||
<!-- DON'T DELETE --><partinfo>BBa_J23111 EndReviews</partinfo> | <!-- DON'T DELETE --><partinfo>BBa_J23111 EndReviews</partinfo> | ||
| + | |||
| + | <I>'''iGEM CINVESTAV_IPN_UNAM''' CHARACTERIZATION OF IGEM DISTRIBUTION BIOPARTS</I> | ||
| + | |||
| + | For contribute to the parts registry our team decided to make the characterization of constitutive promoters, in ''E. coli'', belonging to the family isolated from a small combinatorial library (J23101 , J23102, J23104, J23107, J23108, J2311, and J23115) which were attached to GFP, in psB1C3, to determine promoter activity, using the equipment Victor X3 Multilabel Plate Reader. | ||
| + | |||
| + | [[Image:Gfp1.jpg]] | ||
| + | |||
| + | '''Fig. 1 Construction of the promoter J23111 expressing GFP.''' | ||
| + | |||
| + | '''Methods''' | ||
| + | |||
| + | With the selected colonies, an overnight culture was made in M9 media(minimal media supplemented with 0.2% CAA). After 12 hours the culture was transferred to a 96 well plate at a 1:10 dilution (20 μl of culture and 180 μL of fresh M9 medium). | ||
| + | OD and fluorescence measurements of the selected colonies were performed at intervals of 30 minutes for 16 h. | ||
| + | From the results the PopS were calculated (polymerases per second). | ||
| + | |||
| + | '''Modeling''' | ||
| + | |||
| + | The ecuations used for calulated de promoter activity were based on (R. K. Jason et. al 2009). | ||
| + | |||
| + | [[Image:ecu4.jpg]] | ||
| + | |||
| + | [[Image:ecu5.jpg]] | ||
| + | |||
| + | '''Results''' | ||
| + | |||
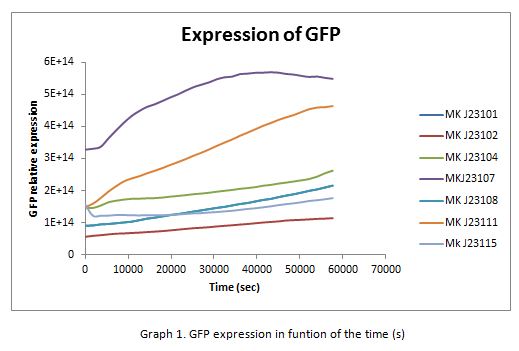
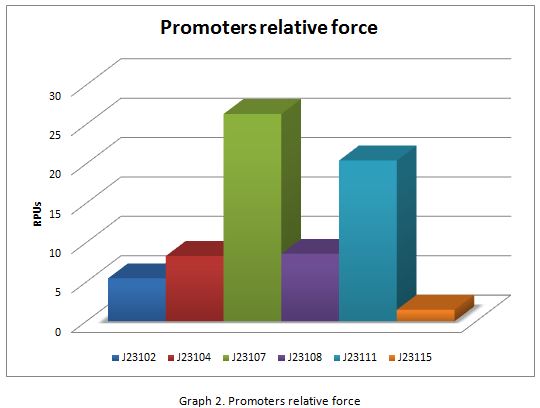
| + | In the following graphs there is shown the GFP expression in function of th time and the realtive promotor intensity. | ||
| + | |||
| + | |||
| + | [[Image:graf1.jpg]] | ||
| + | |||
| + | [[Image:graf2.jpg]] | ||
| + | |||
| + | |||
| + | With the previous results of the characterization of the promoters there is concluded that the promoter J23107, is the strongest because it produces more RPUs” | ||
Revision as of 18:47, 29 September 2012
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_J23111
User Reviews
UNIQ75c9879475c0842f-partinfo-00000000-QINU UNIQ75c9879475c0842f-partinfo-00000001-QINU
iGEM CINVESTAV_IPN_UNAM CHARACTERIZATION OF IGEM DISTRIBUTION BIOPARTS
For contribute to the parts registry our team decided to make the characterization of constitutive promoters, in E. coli, belonging to the family isolated from a small combinatorial library (J23101 , J23102, J23104, J23107, J23108, J2311, and J23115) which were attached to GFP, in psB1C3, to determine promoter activity, using the equipment Victor X3 Multilabel Plate Reader.
Fig. 1 Construction of the promoter J23111 expressing GFP.
Methods
With the selected colonies, an overnight culture was made in M9 media(minimal media supplemented with 0.2% CAA). After 12 hours the culture was transferred to a 96 well plate at a 1:10 dilution (20 μl of culture and 180 μL of fresh M9 medium). OD and fluorescence measurements of the selected colonies were performed at intervals of 30 minutes for 16 h. From the results the PopS were calculated (polymerases per second).
Modeling
The ecuations used for calulated de promoter activity were based on (R. K. Jason et. al 2009).
Results
In the following graphs there is shown the GFP expression in function of th time and the realtive promotor intensity.
With the previous results of the characterization of the promoters there is concluded that the promoter J23107, is the strongest because it produces more RPUs”