Difference between revisions of "Part:BBa K292006:Experience"
(→User Reviews) |
(→User Reviews) |
||
| Line 89: | Line 89: | ||
{|border="0" | {|border="0" | ||
|[[File:Testkutt_BBa_K292006.png|x300px]] | |[[File:Testkutt_BBa_K292006.png|x300px]] | ||
| − | |[[File:RBS+LacI+term-gel. | + | |[[File:RBS+LacI+term-gel.PNG|x300px]] |
|- | |- | ||
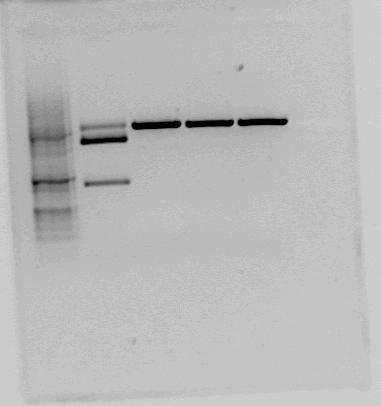
|This is the test cut of BBa_K292006 that the NTNU iGEM team 2011 performed. The testcut was performed with EcoRI+PstI (expected fragments: 1303 bp + 2053 bp), BglI+BclI (expected fragments: 1324 bp + 2032 bp), BglI+EcoRV (expected fragments: 1596 bp + 1760 bp) and BglI+BanII (expected fragments: 1521 bp + 1835 bp). The test cut shows that none of the expected fragments are present. | |This is the test cut of BBa_K292006 that the NTNU iGEM team 2011 performed. The testcut was performed with EcoRI+PstI (expected fragments: 1303 bp + 2053 bp), BglI+BclI (expected fragments: 1324 bp + 2032 bp), BglI+EcoRV (expected fragments: 1596 bp + 1760 bp) and BglI+BanII (expected fragments: 1521 bp + 1835 bp). The test cut shows that none of the expected fragments are present. | ||
Revision as of 20:54, 26 September 2012
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_K292006
Reliable to use when we want to control a gene expression by using pLac promoter. In fact when LacI is not expressed, pLac is activated, and when there is an expression of LacI, pLac is inhibited. We can easily design a system to induce the transcription of LacI by using another operon as the tetracycline operon and creates a failed feedback mechanism for pLac inhibition and so, for gene expression inhibition.
User Reviews
UNIQ0a1041578d3b90bc-partinfo-00000000-QINU
|
•
NTNU_Trondheim 2011 |
We found and orderd this part from the parts registry. Since it had RBS and double terminator it would save us from some cloning work. When testcutting the constructs containing BBa_K292006 we got some unexpected results. Therefore we decided to digest the brick with three differnt enzymes with restriction sites in the given biobrick-sequence in addition to EcoR1 and PstI (for looking at the biobrick length) Except from the sample cut with EcorR1 and PstI, the samples were also cut with BglI wich has a recognition site in the pSB1A2 backbone. Overwiev of restrctions and expected fragmentlengths:
As we can see on the resulting gel none of the restriction sites inn the LacI2 fragment appear to be present. In additon the LacI2 insert looks smaller than it's supposed to be. This indicates that the biobrick sequence does not match the sequence given in the parts registry.
|
UNIQ0a1041578d3b90bc-partinfo-00000002-QINU
UNIQ0a1041578d3b90bc-partinfo-00000003-QINU